Team:Newcastle/Labwork/11 September 2009
From 2009.igem.org
(→Results) |
(→Results) |
||
| Line 38: | Line 38: | ||
[[Image:Team Newcastle 2009 iGEM Geldoc 2009-09-11 15hr 59min.jpg|500px|center]] | [[Image:Team Newcastle 2009 iGEM Geldoc 2009-09-11 15hr 59min.jpg|500px|center]] | ||
<br> | <br> | ||
| + | The lanes are as follows: | ||
| + | '''Row 1''' | ||
| + | * Lane 1 = blank | ||
| + | * Lane 2 = ''HindIII'' DNA Ladder | ||
| + | * Lane 3 = ''cotC'' midi-prep sample | ||
| + | * Lane 4 = ''kinA'' midi-prep sample | ||
| + | * Lane 5 = ''pGFP-rrnB'' midi-prep sample | ||
| + | * Lane 6 = ''pMUTIN4'' midi-prep sample | ||
| + | * Lane 7 = ''pSB1AT3'' midi-prep sample | ||
| + | * Lane 8 = blank | ||
| + | * Lane 9 = ''cotC'' PCR reaction 1 sample | ||
| + | * Lane 10 = ''cotC'' PCR reaction 2 sample | ||
| + | * Lane 11 = ''cotC'' PCR reaction 3 sample | ||
| + | * Lane 12 = ''cotC'' PCR reaction 4 sample | ||
| + | * Lane 13 = blank | ||
| + | * Lane 14 = ''pMUTIN4'' midi-prep sample (from second attempt) | ||
| + | * Lane 15 = leak-over of sample from lane 14 | ||
| + | * Lane 16 = ''pMUTIN4'' midi-prep sample (from second attempt) | ||
| + | * Lane 17 = ''HindIII'' DNA Ladder | ||
| + | <br> | ||
| + | '''Row 2''' | ||
| + | * Lane 1 = blank | ||
| + | * Lane 2 = ''HindIII'' DNA Ladder | ||
| + | * Lane 3 = ''sac'' midi-prep digest sample | ||
| + | * Lane 4 = ''ara'' midi-prep digest sample | ||
| + | * Lane 5 = ''sleB'' midi-prep sample | ||
| + | * Lane 6 = ''HindIII'' DNA ladder | ||
===Conclusions=== | ===Conclusions=== | ||
Revision as of 09:52, 15 September 2009
Lab Work - 11/09/09
Metal Sensing Team
Introduction and Outline
So far we have transformed DH5-alpha E. coli cells with two synthesized BioBricks, which are the cotC-GFP-smtA and the kinA BioBricks. The cotC-GFP-smtA BioBrick is involved within the metal sensing sub-project whereas the kinA BioBrick is involved with the Stochastic Switch sub-project. The Metal Sensing team have since taken these transformant colonies and used them to set up LB+kanamycin cultures for mini-preps and midi-preps; these were then left overnight in the orbital shaking incubator and are awaiting processing today. However there are a range of tasks which need to be accomplished today; these are listed below:
- Prepare Mini-preps of the kinA and cotC-GFP-smtA transformants
- Carry out the mini-prep processes
- Analyse the mini-prep DNA by treating with restriction enzymes.
- PCR the cotC-GFP-smtA BioBrick with pMK15 and pMK16
- Set up the PCR
- Run the PCR for the correct duration
- Analyse PCR products through DNA gel electrophoresis
- Clean up PCR products if PCR reaction proves to be successful
- Midi-prep cotC-GFP-smtA, kinA, pGFP-rrnB, pMUTIN4 and pSB1AT3
- Carry out the five midi-preps.
- Quantify the midi-prep samples
- Run the midi-prep samples on agarose gel through DNA gel electrophoresis
- Run products from a PCR reaction involving arsR and cadA DNA
- Run PCR products on gel
- If successful, clean up fragments, run on gel again, cut with restriction enzymes NheI and BamHI, clean and then ligate together fragments.
Procedure
The following procedures were carried out:
Freeze the kinA and cotC-GFP-smtA transformant cells
Instead of preparing mini-prep samples from the two groups of transformant E.coli cells (i.e. kinA transformed E.coli cells and cotC-GFP-smtA transformed cells), we instead took the decision to use the inoculated 5ml LB tubes in freezing the cells. Analysis of the midi-prep samples will be used to determine whether the BioBricks have been truly transformed - this decision will both save time and also provide a back-up stock of cells for future work. This procedure was carried out by James using the protocol
PCR the cotC-GFP-smtA BioBrick with pMK15 and pMK16
Midi-prep cotC-GFP-smtA, kinA, pGFP-rrnB, pMUTIN4 and pSB1AT3
Run products from a PCR reaction involving arsR and cadA DNA
Results
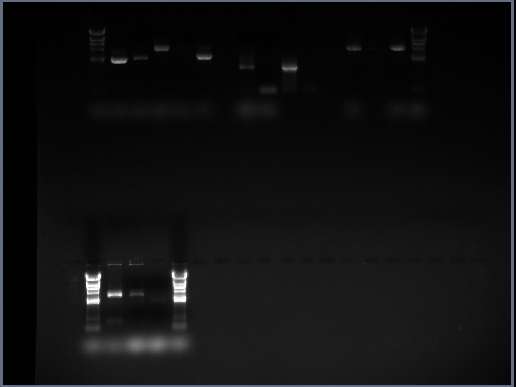
We took the decision to run 4 sets of DNA samples on 0.8% agarose gel - this includes the midi-prep samples (i.e. cotC, kinA, pGFP-rrnB, pMUTIN4 and pSB1AT3), products from a PCR reaction involving cotC, futher midi-preps of pMUTIN4 and digests from midi-prepped DNA the Stochastic Switch team have prepared (i.e. sac, ara and sleB) The resulting GelDoc photograph can be seen below:
The lanes are as follows:
Row 1
- Lane 1 = blank
- Lane 2 = HindIII DNA Ladder
- Lane 3 = cotC midi-prep sample
- Lane 4 = kinA midi-prep sample
- Lane 5 = pGFP-rrnB midi-prep sample
- Lane 6 = pMUTIN4 midi-prep sample
- Lane 7 = pSB1AT3 midi-prep sample
- Lane 8 = blank
- Lane 9 = cotC PCR reaction 1 sample
- Lane 10 = cotC PCR reaction 2 sample
- Lane 11 = cotC PCR reaction 3 sample
- Lane 12 = cotC PCR reaction 4 sample
- Lane 13 = blank
- Lane 14 = pMUTIN4 midi-prep sample (from second attempt)
- Lane 15 = leak-over of sample from lane 14
- Lane 16 = pMUTIN4 midi-prep sample (from second attempt)
- Lane 17 = HindIII DNA Ladder
Row 2
- Lane 1 = blank
- Lane 2 = HindIII DNA Ladder
- Lane 3 = sac midi-prep digest sample
- Lane 4 = ara midi-prep digest sample
- Lane 5 = sleB midi-prep sample
- Lane 6 = HindIII DNA ladder
Conclusions
News
Events
- 20 – 21 June 2009 - Europe workshop (London)
- 23 – 24 June 2009 - UK iGEM meetup (Edinburgh)
- 23 October Practice Presentation (Newcastle)
- 23 October T-shirts are ready
- 27 October Practice Presentation (Sunderland)
- 27 October Poster is ready
- 30 October – 2 November 2009 - Jamboree (Boston)
Social Net
- Newcastle iGEM Twitter
- [http://www.facebook.com/home.php#/group.php?gid=131709337641 Newcastle on Facebook]
- [http://www.youtube.com/user/newcastle2009igem Newcastle Youtube Channel]
 "
"