Team:TUDelft/Modeling
From 2009.igem.org
Delay Device
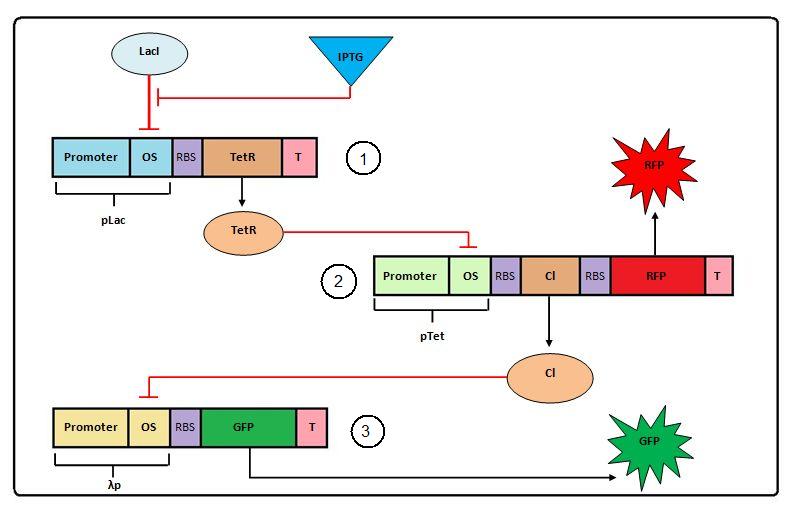
Figure 1: Negative Feedforward model
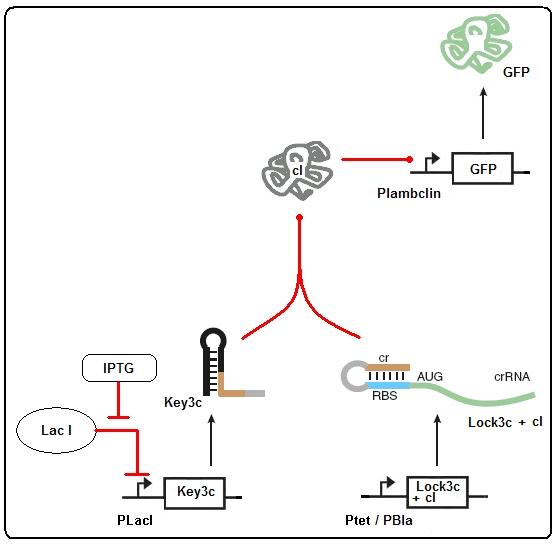
Figure 2: Riboregulator model
Mathematical Modeling
The Negative Feedforward model shown above can be expressed with the following set of ODEs (ordinary differential equations):
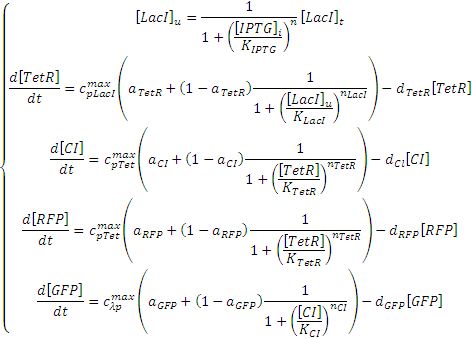
The Riboregulator model shown above can be expressed with the following set of ODEs:
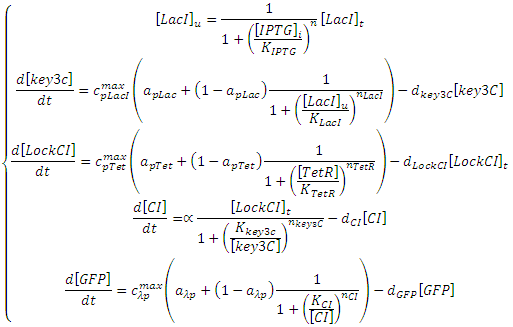
The symbols in this system of equations are found in the table below:
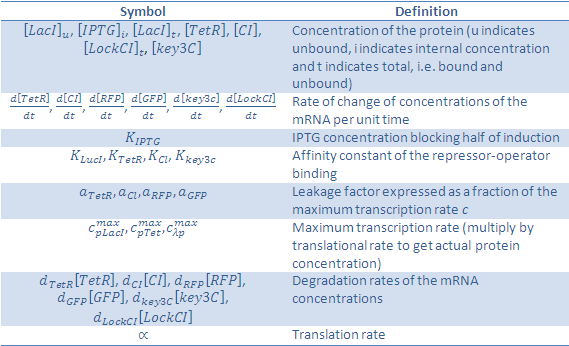
We would like to give credit to the Bologna '08 team and UNIPV-Pavia '08 team, who's project was useful in determining our own equations.
Conjugation
Using the paper "[http://www3.interscience.wiley.com/journal/118837812/abstract?CRETRY=1&SRETRY=0 A model for bacterial conjugal gene transfer on solid surfaces]", we established the assumptions and parameters that would be needed for the modeling of the conjugation.
Assumptions
- Cells are distributed randomly on the agar when introduced to it
- Number of cells grows until one or more of the medium components are exhausted, unless the initial nutrient concentration is lager than the saturation constant for growth (i.e. the ability of the colony to expand)
- Donors, recipients andf transconjugants have identical colony growth rates, specific growth rates and cell yields on solidified LB
- Conjugation occurs such that all recipients become transconjugants after a certain conjugation time
- Plasmid loss is negligable
- Cells only move on the surface through colony expansion
Parameters
- Surface area of media (A)
- Initial colony radius (r_0)
- Specific growth rate (g_n)
- Colony radial growth rate (g_r)
- Maximum numbers of cells sustained by system (N_max)
- Initial number of donors (N_d) and recipients (N_r)
All these parameters can be determined experimentally, or by literary research into previous conjugation experiments.
Spatial distribution of bacteria
Growth of colonies
Contact between colonies
Conjugation modeling
Final cell number
 "
"







