Team:Brown/Parts
From 2009.igem.org
Biobricks...saving one sneeze at a time!
rEV131 ()
Paraphrased description from “Tick Histamine-Binding Proteins: Isolation, Cloning, and Three-Dimensional Structure” by Paesen et al. (1999)
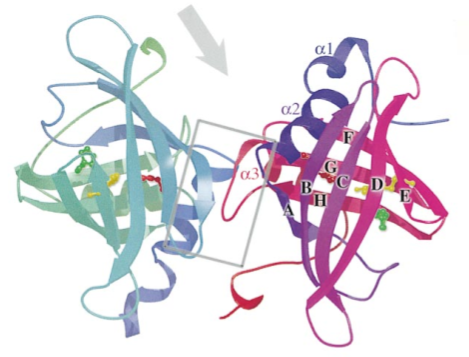
EV131 (also known as rRa-HBP2) is a histamine binding protein, and one of three discovered in the salivary gland extracts of Rhipicephalus appendiculatus ticks. Two major proteins (Ra-HBP1 and Ra-HBP2) were found in the saliva of female ticks, and a third protein (Ra-HBP3) was retained from male salivary gland extracts. On SDS gels, the proteins have apparent molecular masses between 20 and 25 kDa.
The crystallographic structure and biological activity of these HBPs (Histamine binding proteins) indicate that they sequester histamine at wound site, outcompeting histamine receptors for the ligand, thereby overcoming their hosts’ inflammatory (and related immune) responses and feeding successfully. Acting independently of the membrane-bound H1, H2, and H3 receptors, HBPs offer a new approach to the control of histamine-based diseases, such as allergic rhinitis.
Binding of histamine to the three rHBPs appears to be saturable. Scatchard plots show high affinities for rRa-HBP3 (equilibrium dissociation constant [KD] 1.2X10^-9 M; SD=0.4; three measurements) and for rRa-HBP2 (KD 1.7x10^-9 M; SD=0.9), but a lower affinity for rRa-HBP1 (KD 1.8x10^-8 M; SD=1.2).
A series of histamine-like compounds were tested for their ability to compete with 3H-histamine for binding to the proteins. Depending on the protein tested, 100–240 times more 1-methylhistamine than cold histamine, and 600–1000 times more 3-methylhistamine, were needed for a 50% reduction of bound radioactivity. No significant competition was observed with other related compounds (histidine, imidazole, serotonin, dopamine, the H1 receptor agonist betahistine, the H1 antagonists chlorpheniramine and pyrilamine, the H2 agonist dimaprit, the H2 antagonists ranitidine and cimetidine, and the polyamines putrescine, spermine, and spermidine). This indicates highly specific histamine binding, different from that of the mammalian H1 and H2 receptors (Gantz et al., 1992).
Paraphrased description from “Arthropod-Derived Histamine-Binding Protein Prevents Murine Allergic Asthma” by Couillin et al. (2004)
EV131 has a distinctive feature because it presents a second specific binding site for histamine with lower affinity than the high affinity binding site, as revealed by its crystal structure.
Allergen challenge enhances histamine release upon OVA-specific IgE cross-linking on mast cell and subsequent degranulation with release of histamine in sensitized mice, leading to bronchoconstriction, eosinophilia, and mucus hypersecretion. Administration of EV131 to BP2 strain of mice reduced significantly the peribronchial eosinophilia, mucus hypersecretion, and hyperplasia of bronchial smooth muscles. Therefore, complete in vivo neutralization of histamine with the high affinity histamine-binding protein EV131 inhibited the inflammatory cell recruitment and suppressed the characteristic allergic inflammation of the airways.
Tar-EnvZ ()
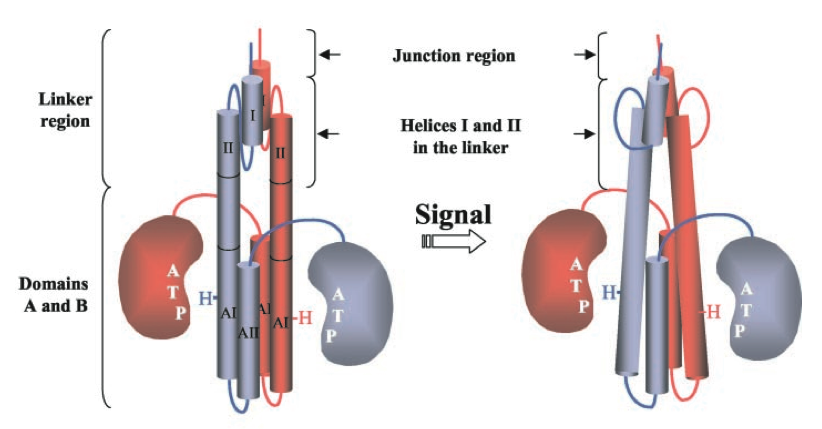
In our system, the Tar-EnvZ (or Taz) chimera protein is used to indicate and signal the presence of a chemical ligand. Endogenous to E. coli cells, Tar has three domains: a periplasmic ligand binding receptor domain, a transmembrane domain, and an intracellular kinase domain. When aspartate binds to the Tar receptor domain, the kinase domain subsequently propagates a message by modifying intracellular components, ultimately resulting in regulation of flagella rotation.
Also endogenous to E. Coli is the EnvZ protein, an inner membrane kinase which responds to changes in osmolarity. When activated, the EnvZ kinase phosphorylates transcription factor OmpR, which subsequently activates transcription of the OmpC gene.
Tar-EnvZ (Taz) is a chimera protein, manufactured by Inouye, et al. We obtained the gene from his lab and Biobricked it. Taz comprises of the aspartate chemoreceptor region of Tar, the transmembrane region of Tar, and the intracellular kinase region of EnvZ. The genes were fused by digesting both with NdeI and ligating the overlapping ends together. The cut site is between amino acids H256 and M257.
OmpC promoter-RFP
()
This construct is a reporter for EnvZ activation. Once activated, EnvZ phosphorylates transcription factor OmpR, which in turn activates transcription of the OmpC gene. This cassette contains the promoter region of OmpC (BBa_R0082), placed over a red fluorescence protein gene (BBa_E1010). The ribosome binding site is . There is also a double terminator sequence.
[DNA Sequence]
pAgr ()
This part is the promoter region + RBS of the Agr operon in S. epidermidis. The agr operon transduces the signal that a quorum of S. epidermidis has been reached, based on the extracellular concentration of signalling oligopeptides. When the bacteria reaches a quorum, the genes under the agr operon are usually turned on (this includes virulence factors that can create a biofilm).
NOTE: This part only works in S. epidermidis.
secretion signal peptide + GFPmut3b ()
This is a composite signalling part. The secretion signal peptide for beta-lactamase in S. epidermidis is ligated N-terminally to registry part BBa_E0040 (GFPmut3b). The signal peptide motif is used by the bacteria to target the protein for secretion through the Sec pathway. Any part may be put in place of GFP, but in its current state, the amount of secretion of any protein may be studied using fluorescent microscopy.
NOTE: This part only works for S. epidermidis.
TrgEnvZ ()
Transduces ribose-binding protein signal to OmpC promoter. This chimeric protein is composed of a trg domain joined to an EnvZ histidine kinase domain.
mRBP (modified ribose binding protein - now binds histamine) ()
This is a mutated version of RBP, an E. coli periplasmic ribose-binding protein. Natively, ribose binds as a ligand, transducing a message to the trg domain of a transmembrane histidine kinase. Here we have computationally redesigned RBP's ligand binding pocket to bind histamine preferentially over ribose, yielding mRBP.
There are sixteen locations (S9K, N13D, F15E, F16H, N64H, D89G, R90D, S103T, I132D, A137T, R141L, F164D, N190H, F214V, D215G, Q235T) at which RBP is mutated to create mRBP. These mutations redesigned the protein's ligand-binding pocket to bind to histamine instead of ribose, while still allowing the protein to fold properly and maintain its shape. These mutations were introduced based on predictions made by the Rosetta 3 macromolecular modeling program, which probabilistically searches through mutations to RBP's ligand binding pocket, then simulating the interaction between the mutated protein and histamine (similar to the method in Looger et al.). The predicted design with the lowest energy was selected and the DNA that coded for it was synthesized by GeneArt (see Brown 2009 wiki>Projects>Histamine Binding for more details).
The mechanism of mRBP's binding may be thought of like that of a venus fly-trap. There are two domains that form the top and bottom of its "mouth", and the ligand binding pocket is between these two domains. The two domains are connected by a hinge that allows the protein to snap shut on a ligand when it goes into its bound conformation.
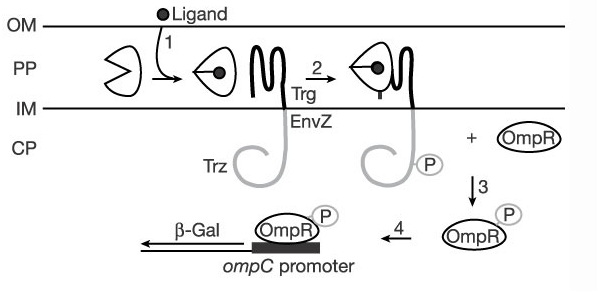
In its bound conformation, mRBP interacts with the Trg domain of the chimeric TrgEnvZ histidine kinase, which then phosphorylates OmpR, which causes expression of genes under the OmpC promoter. The more mRBPs exist in the periplasmic domain bound to histamine, the more it interacts with TrgEnvZ, and the more gene expression there is in a cell containing all three genes and the promoter.
 "
"