Team:EPF-Lausanne/Results/ELS
From 2009.igem.org
Contents |




We run a a 62ns simulation on the 2v0w light state using 32 processors.
Here is a video.
Validation of the light state simulation
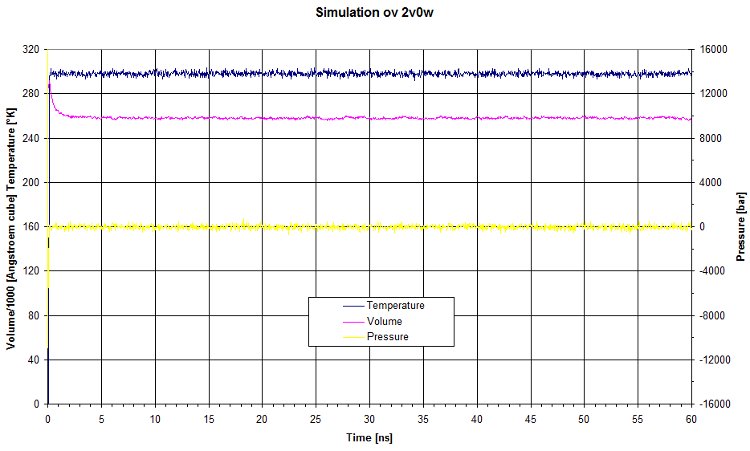
PVT
Here are the values of pressure, volume and temperature for this run.
RMSD
The rmsd along the run gives:
The RMSD is another test for the analysis of the evolution of the global structure of a protein. It compuites the mean quadratic distance between the structure troughout the simulation, and the original cristallographic structure.
Une deuxième analyse de l’évolution de la structure globale d’une protéine à laquelle on fait souvent appel est l’écart quadratique moyen entre la structure au cours de la simulation et la structure cristallo- graphique ou de départ (RMSD du terme anglais root mean square deviation). Le RMSD prend en compte la distance scalaire entre atomes du même type comparant deux structures, après que ces structures aient été superposées, le plus souvent en minimisant la valeur du RMSD. Souvent on se limite aux RM- SD des carbones α pour comparer les protéines.
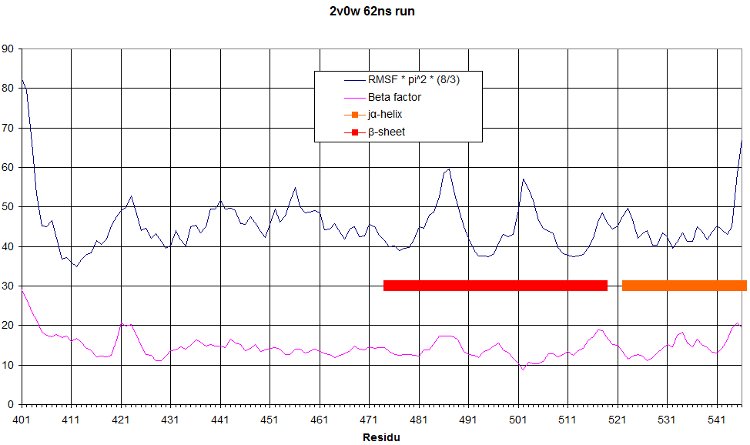
RMSF
And the rmsf is:
Analysis of the simulation
As for the dark state, we tried to find a change in the j-alpha helix then looked at the comportment of the residues.
Angle between beta sheet and alpha helix
We ploted the angle between the helix and a strand of the beta sheet. As in the light state we don't see a clear movement.
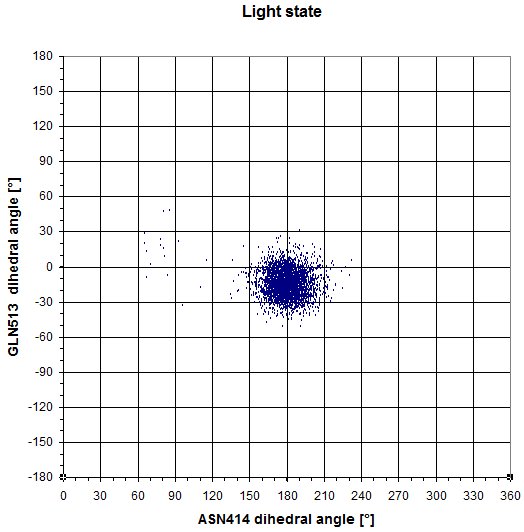
GLN513 - ASN414
We also redid the same experiments as in the dark state. Here is the plot of the dihedral angle of GLN513 and ASN414. In this case, both residues don't switch to another conformational state.
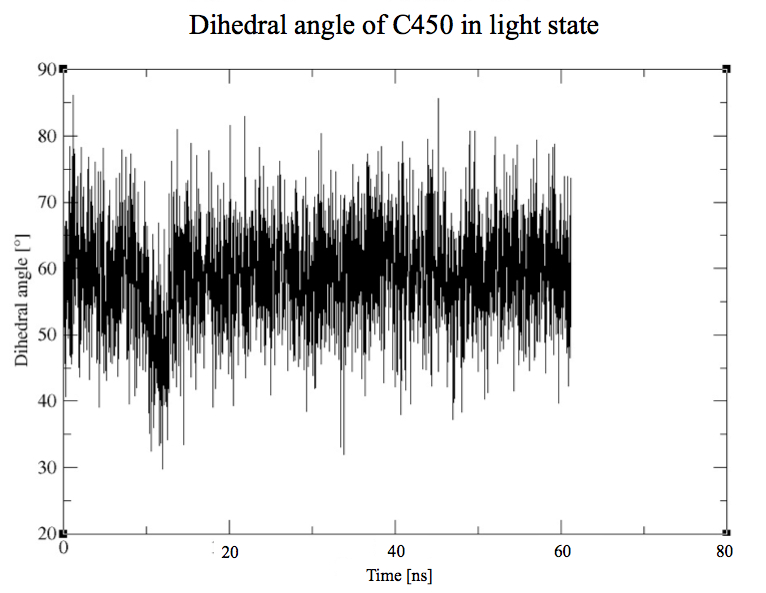
CYS450 - FMN
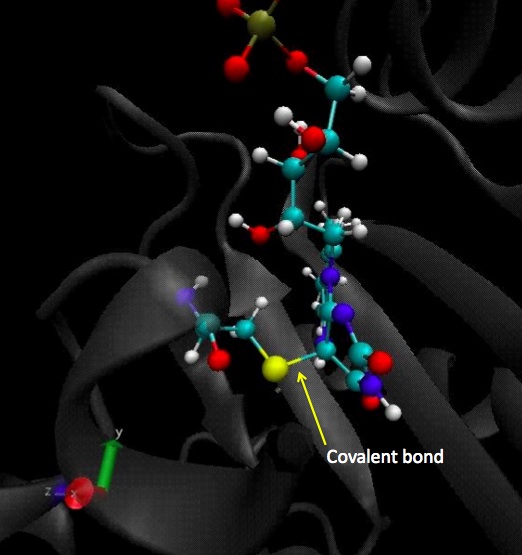
Same study than in the dark state for the residue n° 450, with an important difference to point out which is the covalent bond between the residue 450 and the cofactor.
Here is the covalent bond between the cystein 450 and the cofactor, FMN:
Here we plot the dihedral angle of this residue to see how it moves generally. As expected, the residue is now stable because of the covalent bond to the FMN.
 "
"