Team:Groningen/Project/Vesicle
From 2009.igem.org
[http://2009.igem.org/Team:Groningen http://2009.igem.org/wiki/images/f/f1/Igemhomelogo.png]
|
|---|
- Transport
- Accumulation
- Metal-sensitive Promoters
- Gas Vesicles
Gas vesiclesOur goal in this project is to make cells bouyant in the presence of certain concentrations of metals like copper, zinc and arsenic. Metal induced gas vesicle production can provide our cells with this bouyancy. Gas vesicles are bacterial organelles consisting entirely of proteins that envelope a gas filled space. We made, and send to the registry, parts in which the metal sensitive promoters for copper, zinc and arsenic were cloned in front of the gvp (Gas Vesicle Protein) gene cluster. For further characterization of the gvp gene cluster inducible and constitutive promoters were also cloned in front of this cluster. Buoyancy tests showed that our constructs were able to increase cell buoyancy and electron micrographs showed the presence of gas vesicles. A model was made to predict what volume fraction of a cell would have to be gas vesicle for this cell to have a density equal to that of water.
|
Background
Gas vesicles are organelles made entirely out of proteins that envelope a gas filled space. Because only gas can penetrate the gas vesicles the total density of the cell is lowered. This lower cell density leads in turn to a buoyancy phenotype. Outside of the laboratory this buoyancy is used by microorganisms to vertically position themselves in the water column or simply to reduce their sinking rates. Organisms can regulate buoyancy by reducing gas vesicle production or by accumulation of denser compounds like carbohydrates. For certain cyanobacteria this regulation depends on light intensities.
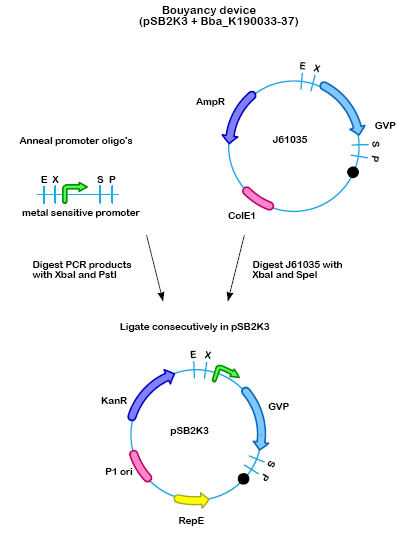
For a number of organisms it has been shown that all proteins important for the expression of gas vesicle are part of a single gene cluster. The gvp gene cluster used in this project was cloned from Bacillus megaterium into E. coli (Li & Cannon 1998). This gene cluster, now containing 11 genes was turned into a biobrick by Melbourne 2007 (). Figure 1 shows the gene cluster as it was send in by Melbourne in 2007.
For more info see: Walsby 1994.
Goal
The goal of our project is to engineer an organism that can remove heavy metals from water. To facilitate easy separation the cells that have taken up metals should float so they can be removed from the water. The introduction of gvp gene cluster provides the cell with the required buoyancy.
To make the buoyancy level of the cell responsive to the metal concentration inside the cell, metal sensitive promoters would have to be cloned in front of the gvp gene cluster. In this project these metal sensitive promoters were responsive to zinc, copper and arsenic. We also wanted to make a construct with constitutive and inducible promoters in front of the GVP cluster to show a proof of principle and as a back-up if our metal induced construct would fail.
Finally we wanted to improve the Melbourne 2007 biobrick () by removing a repeat that was accidentally introduced during the removal of forbidden restriction sites.
Cloning strategy
For our msGVP (metal sensitive GVP) constructs we ordered oligo's containing the promoter region and the necessary restriction sites. When annealed these pieces of DNA have EcoRI and SpeI sticky ends. The vector containing GVP () was cut with EcoRI and XbaI and was ligated to the promoter. (Protocol)
The metal sensitive GVP constructs are: , and .
Then because of compatibility issues when our entire system has to be assembled into one cell the whole metal sensitive promoter and GVP part were transfered to a different vector.
- Figure 2: The floating device will be built up of an inducible promoter which can be induced by a certain intracellular concentration of metal-ions, and a gas vesicle cluster.
Buoyancy Tests
The buoyancy of GVP was tested by using the buoyancy test protocol. The cells were grown in medium and induced and were resuspended in a salt solution (0.15 mM NaCl) in a test tube and were left for a while in order to give the cells time to sink or float.
Different circumstances
Several different circumstances and small changes to the protocol were made in order to find the perfect circumstance for the buoyancy test. It appeared that with a low cell density the difference between floating and sinking could not be seen very well. The results were best visible with and cell density of OD600=1.5. Also we tried to do the buoyancy test in a longer tube since it was expected that the difference between floating and sinking would be more obvious. This, however, did not appear to be the case, unfortunately. Also doing the buoyancy test in a higher saline concentration did not have an enhanced floating effect. Another adaption we tried was the way of induction. In the standard protocol the cells were induced in the overnight culture. It was also tried if induction in the saline or at the exponential phase of growth or even induction on plate would make any difference. Unfortunately this did not make a huge difference.
Fermentor test
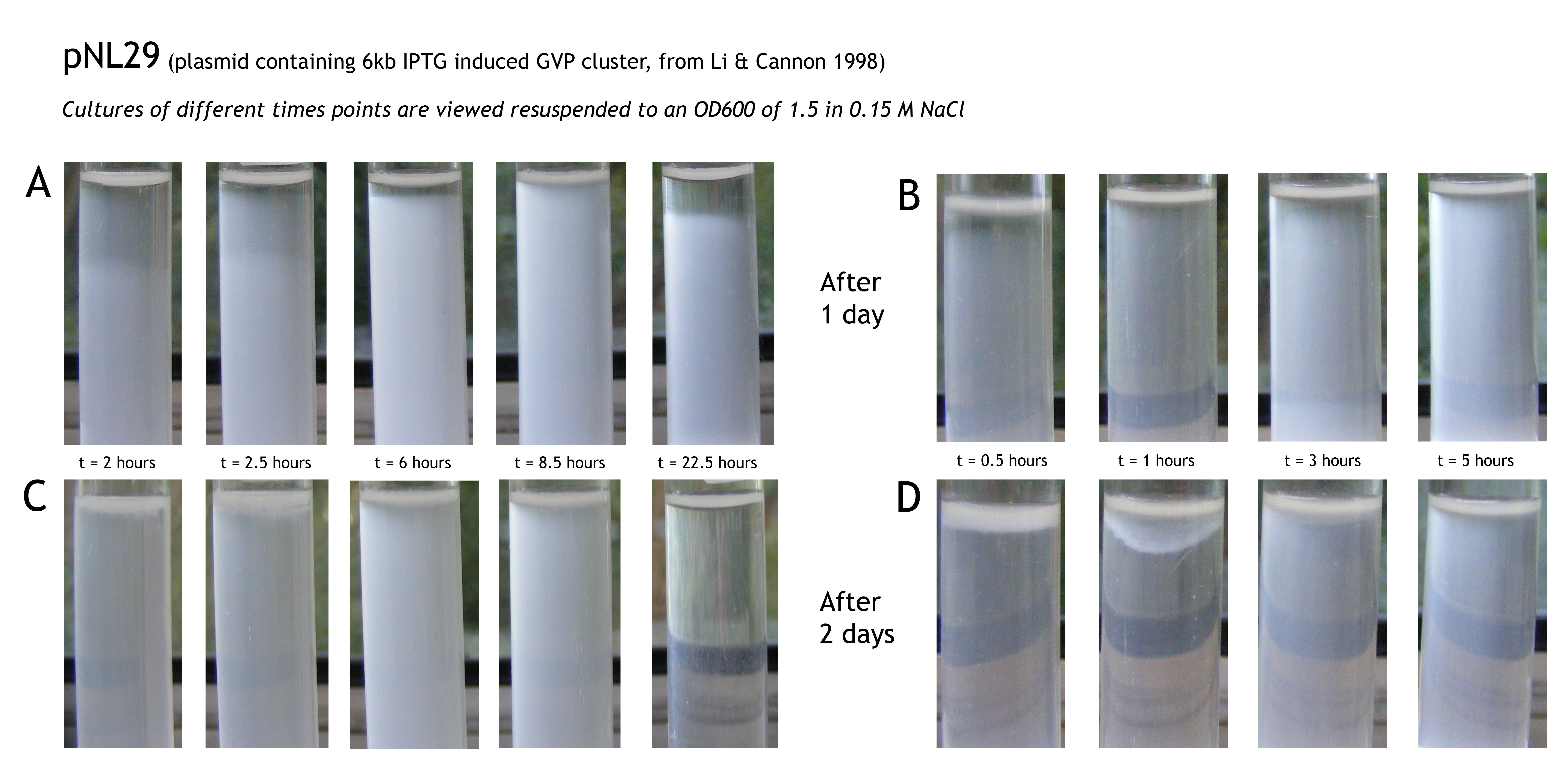

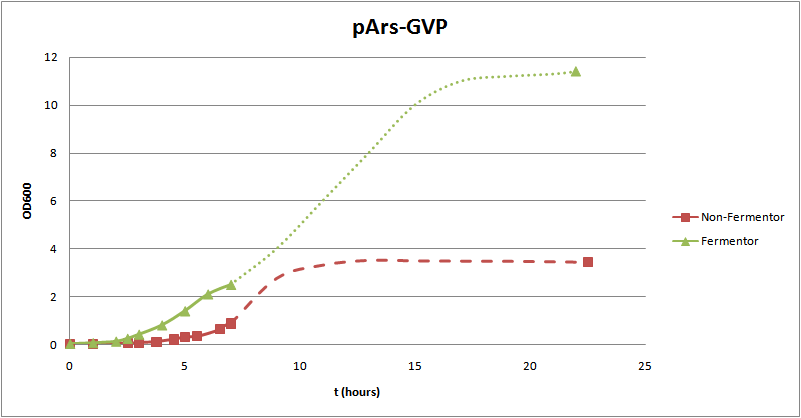
So the length of the tube, the saline concentration and the time of induction did work for the buoyancy test. It was also suggested that there was not enough gas in the surrounding of the cells and a better result could be achieved if this could be improve or another gas could be used. To test this we tried to grow the cells in a fermentor. It was also suggested that the fermentor test could be done with helium, however, modelling showed that it would not make a difference in floating which gas is used as long as it is lighter than water (check this yourself by changing the density (ρ) of the gas here). Therefore the fermentor test was done by using O2. This resulted in better buoyancy results. As can be seen in figure 3A the positive control pNL29 showed better buoyancy over time. After 2 hours the cells were in exponential phase and were induced with IPTG. After 8,5 hours the buoyancy is best, after 22.5 hours the buoyancy the cell level is declining. This suggests that there is an optimum after 8,5 hours. The cells at t=22.5 are probably in stationary phase whereas the cells at t=8.5h could still be in exponential phase, this could explain the difference in buoyancy found. It suggests that in stationary phase less gas vesicles are produced. Figure 3C shows the same tubes 24 hours later. This still shows buoyancy for the t=6 and t=8 tubes and no buoyancy for the others. This suggest that the buoyancy last for at least 24 hours. Simultaneously a normal, non-fermentor, buoyancy test was also performed with the same construct. In figure 3B these results at day 1 can be seen, this shows nothing. After a day no buoyancy can be seen for t=1 and t=2 a more dense cell suspension can be seen for t=3 and t=4 (figure 3D), however still no confincing buoyancy can be seen. Figure 4 shows the results from one of our own constructs, pArsR-GvP () grown in a fermentor. This shows an increase in buoyancy in time, however, at t=22.5h no buoyant cells can be seen. A buoyancy test done at the same time without a fermentor shows the same increase in buoyancy but does show buoyancy at t=22.5h (figure 4B). This difference can be explained since the cells in the fermentor are probably already dead or dying. In a fermentor the cell density is large this causes the cells to die.
Electron Microscopy
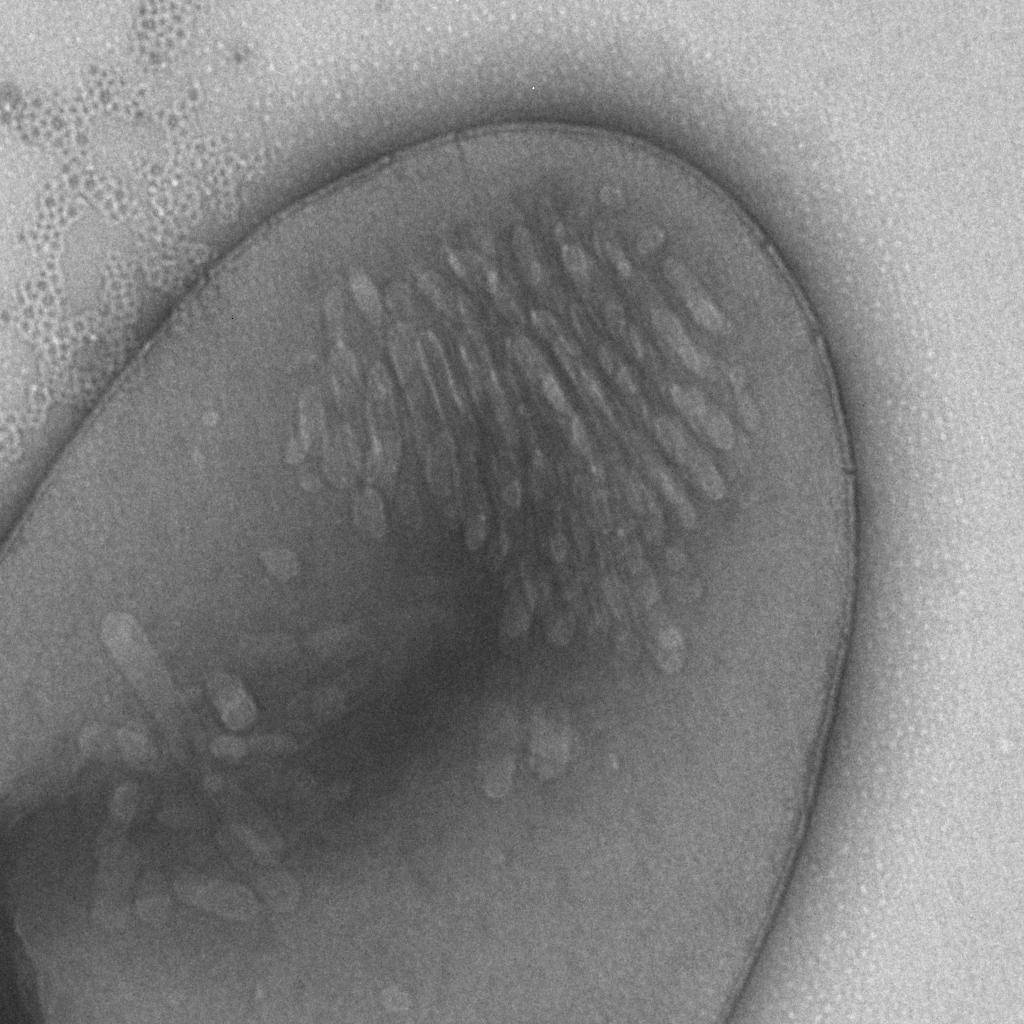
To check whether gas vesicles really were present in the cells we did some electron microscopy.
In Figure 5 a picture of gas vesicles in a protoplast can be seen. This protoplast comes from an E. coli cell that contained a plasmid with the gvp gene cluster behind an arsenic sensitive promoter ().
Modelling
Buoyancy
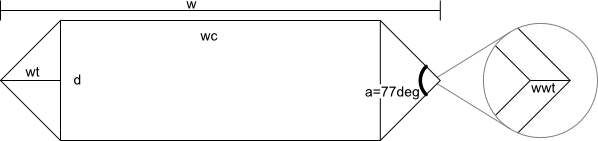
The gas vesicles are shaped roughly like a cylinder with a cone at each end, whose cross-section we model as (based mostly on Walsby 1994):
We assume the interior of the wall of the gas vesicle is similarly shaped to the exterior, just slightly smaller (the right-most part of the image above illustrates this situation for the left tip of the gas vesicle). This means the different dimensions are related through the equations below. To determine the total volume, just use them with the given width/diameter (at least for the dimensions given in Walsby 1994). To determine the gas volume, use them with wgas and dgas.
| w = total width tw = thickness of wall (1.8-1.95nm) d = diameter a = 77 degrees ρgas = density of gas in vesicle (kg/m^3 = yg/nm^3) ρwall = density of vesicle wall (kg/m^3) wwt = tw/sin(a/2) wt = (1/2)*d/tan(a/2) wc = w - 2*wt Vc = (1/4)*pi*d^2*wc Vt = (1/12)*pi*d^2*wt V = Vc+2*Vt M = ρ*V wgas = w-2*wwt = width of gas space dgas = d-2*tw = diameter of gas space V = Vgas + Vwall |
Now we can consider the buoyant density of E. coli with gas vesicles. We have chosen to approach this problem using densities and volume ratios. According to Baldwin 1995, Bylund 1991 and Poole 1977, the density of (wild-type) E. coli is 1100 kg/m3 ±3% under wildly varying conditions. This makes our method easier than trying to directly compute the density of a single cell, due to the fact that the volume can differ wildly (both during the life cycle and from strain to strain) and a lack of concrete data on the number of gas vesicles produced (in E. coli). Note that the computations below assume that the gas vesicles simply add to the existing structures.
Loading graph...
| Vc = volume of a cell without gas vesicles Vv = volume of gas vesicles in cell Vcv = volume of a cell with gas vesicles (assumed to be Vc+Vv) ρc = density of a cell without gas vesicles ρv = density of gas vesicles ρm = density of medium The following has to be true if the cell floats: Vc*ρc + Vv*ρv < Vcv*ρm (Vcv-Vv)*ρc + Vv*ρv < Vcv*ρm ρc + (Vv/Vcv)*(ρv-ρc) < ρm Assume (ρv - ρc)<0 Vv/Vcv > (ρm - ρc)/(ρv - ρc) Vv/Vcv > 1 - (ρm - ρv)/(ρc - ρv) Explanation of the graph Four curves are shown, corresponding to how many gas vesicles a cell needs with "our" gas vesicles (unless you changed the constants in the calculator above), the gas vesicles documented in Li 1998Using a width and diameter of 75nm and 50nm, respectively. Here we assume that their "width" should be interpreted as our diameter, as doing it the other way around would leave no room for a cylinder and they specifically mention that the vesicles appear to be shaped like cylinders with conical ends. i Using a width and diameter of 500nm and 84nm, respectively. i The X-axis depicts the cell density of the part of the cell not occupied by gas vesicles. The Y-axis depicts the minimum volume fraction of the cell that should consist of gas vesicles to make the cell float. |
Conclusion & Discussion
We have experimented with two different constructs containing the gvp gene cluster i.e. pNL29 containing the 6 kb gene cluster from Bacillus megaterium (Li & Cannon 1998) and [http://partsregistry.org/wiki/index.php/Part:BBa_I750016 BBa_I750016] from the [http://parts.mit.edu/igem07/index.php/Melbourne Melbourne 2007] iGEM team. We observed that it is best to have an OD600 of 1.5 when doing bouyancy tests, for withnessing differences with lower values is difficult. Furthermore buoyancy tests carried out in sea water or normal (LB) medium also give rise to difficult to interpret results.
For cells cultivated in a more aerobic environment, such as the ones carried out in a fermentor, an enhanced bouyancy phenotype is observed. The extra O2 added probably causes a higher concentration of intracellular oxygen, that can diffuse to the gas vesicles that are produced. The best buoyancy phenotype is withnessed at t=8.5 hours, however at t=22.5 hours no buoyancy can be seen. This suggests that there is an optimum after 8,5 hours.The cells at t=22.5 are probably in stationary phase whereas the cells at t=8.5h could still be in exponential phase, this could explain the difference in buoyancy found. It suggests that in stationary phase less gas vesicles are produced. Buoyancy is observed after 1 day, and also still after 2 days, which suggest that the buoyancy last for at least 24 hours. This is in accordance with experimental data from Walsby, 1994, who also still observed a buoyant phenotype after 2 days.
 "
"