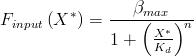Team:Cambridge/Modelling
From 2009.igem.org
Categories :
Project :
-
Overview
Sensitivity Tuner
--- Characterisation
--- Modelling
Colour Generators
--- Carotenoids (Orange/Red)
--- Melanin (Brown)
--- Violacein (Purple/Green)
The Future
Safety
Notebook :
Team Logistics :
Modelling
Introduction
This project focuses on novel outputs, for example, for environmental sensors. However, there is a need for an 'adaptor'; a middle section to the machine that takes an input and processes it. Our initial work was based around the development of an amplifier that permits a large output that is clearly visible. The next planned stage was the creation of a system that allows switching on of output at different calibrated input signal levels. Creating a model allows the feasibility of the proposed systems to be tested. A basic model of the original amplifier system was put forward, building on both our data and the Cambridge 2007 data.
Modelling the phage activator system
This is the basic 'amplifier' system that consists of an input sensitive promoter system,a protein activator and sensitive promoter. It can therefore be divided into two boxes, the approach taken in putting forward an initial model.
The pBAD promoter
An arabinose input acts as an inducer, permitting transcription, by binding the AraC transcription factor. This is a dual transcription factor; when unbound to arabinose a dimer restricts access of polymersase to reduce basal levels of transcription, upon binding arabinose the conformation changes and the dimer permits binding of polymerase. [1]
To model this situation, araC is first assumed to take the role of a repressor that reversibly binds and unbinds a site on the DNA. If it binds arabinose, it is sequestered and cannot bind the DNA. Here, an input function is created, after Alon [2]. This gives the rate of transcription from the promoter dependent on the concentration of arabinose. Since mRNA is then translated at a roughly constant rate, it is related with a multiplicative constant to the rate of protein production, in this case activator and RFP.
This gives the rate of transcription as a function of X* which represents the concentration of active repressor, unbound to arabinose. B is the maximum rate of transcription, here this rate is when induced by arabinose at highest concentration. K_d is the dissociation constant (see modelling derivations). Parameters must be found by a parameter scan for sensible values or by comparing to already gathered data.
The concentration of 'active' repressor is given as a function of arabinose concentration by:
where X^T is the total amount of araC available, bound or unbound. A is arabinose concentration. n is the number of arabinose molecules binding to each molecule of the repressor, and K is a binding constant. n was taken to be two by assuming that each araC dimer needs two molecules to be bound before it can permit transcription.
Combining these two gives the overall input function, which has leaky transcription included at A = 0, seen in the actual results.
The Activator and its Promoter
This is based on a similar idea. Activator is made by transcription from pBAD, the mRNA is then translated (the potential time delays will be taken into account). The activity of the phage promoter is dependent on activator concentration according to:
Assuming that translation rates remain constant, the rate of GFP and RFP production would be expected to be multiples of the above promoter activities/ input functions (which represent rate of transcription)
Making a Latch
A switch that remains on once stimulated would be useful if it was only necessary, say, to see if a hazardous contaminant had ever been present in a sample (it could still be there in low levels etc.). A method proposed is positive feedback; an activator placed downstream of its own promoter (as well as the reporter/ pigment) will, in theory, keep pigment production going. The rate of activator production from its own promoter is given in equation 3 above, which is dependent on activator concentration itself.
Modelling the proposed switching levels system
This idea was summarised in the diagram and discussion included in week 4 dry work: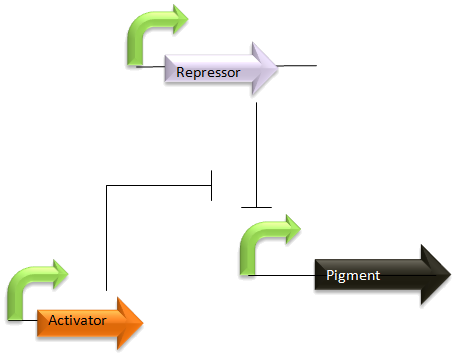
Most of these model systems will rely on the very basic equations outlined above, with extensions to add levels of complexity, such as the transport of arabinose into a cell.
Basic Phage Activators
The first step is to take a look at the preliminary data to try to find appropriate parameters. An overall aim for the modelling of this system is to refine the model to each component and attempt to predict how different combinations of components will behave. The data collection separates the promoter and activator by use of both RFP and GFP reporters.
Extended Systems
Modelling Basics
In order to model the kinetics of gene transcription and translation, the approach taken is similar to that in enzyme kinetics. The transcription factors bind reversibly to the relevant site on the DNA and depending upon the particular case, transcription is promoted or repressed.
The steps of working out the input function (which describes the rate of transcription as a function of the inducer concentration) of a gene are finding the concentration of active repressor (that unbound to the inducer, in this case, arabinose) and then finding the effect of that repressor on transcription.
We have taken araC to act as a repressor in these models, although as mentioned, it has dual activity, remaining on the DNA after arabinose binding to promote transcription.
Binding of repressor to site on DNA
First, working out the probability of the DNA site being occupied or free in the presence of a concentration of transcription factor. The equation describing the reversible DNA-Factor complex [DnX] formation is given below. Note that more than one molecule of repressor, the number here is given by n.
k_1 is the forward rate constant, k_2 is for the reverse reaction.
The probability of the DNA site being free of the repressor is given by [D]/[D_t] where D_t is the total amount of free DNA binding site available. Assuming a pseudo steady state where [Dnx] is stable, so that d[DnX]/dt = 0, and including the constraint that
allows the probability of finding a DNA site free of the repressor to be found by simple substitution and rearrangement of the rate equation below.
This gives the repressor dependent input function of the gene:
The constant K_d is effectively the nth root of k_2/k_1, as can be found by following through the derivations.
Sequestering of the repressor AraC by binding to arabinose
Again, a similar approach is taken as in enzyme kinetics. Effectively, all of the above derivation is followed but this time the amount of active repressor, X*, is found. This assumes that the amount of AraC repressor bound to arabinose is in pseudo steady state. The total effect of the two repressing systems becomes that of an activator and in effect the detail could be hidden to have a simple activated transcription model. This approach would be the best to pursue in characterising the activator systems.
The Phage Activator Systems
Including the pBAD promoter as described above, an additional level of complication needs to be taken account of with the phage activator systems. Here the activator promotes transcription, so the input function for the gene under the activator controlled promoter is found from the probability that the DNA binding site is occupied. This is [DnX]/[D_t] where [DnX] here represents the amount of activator-DNA binding site complex (note this is not the amount of free DNA sites this time). Otherwise, the derivation is as above, with the assumption again that [DnX] is in pseudo steady state.
References
1. [http://www.ncbi.nlm.nih.gov/pubmed/9600836 Apo-AraC actively seeks to loop. Schleif J.Mol. Biol (1988) 278, 529-538]
2. AN INTRODUCTION TO SYSTEMS BIOLOGY Design Principles Of Biological Circuits. Uri Alon
 "
"