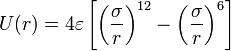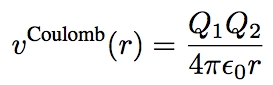Team:EPF-Lausanne/Modeling
From 2009.igem.org
(→Modeling) |
(→Molecular dynamics theory) |
||
| Line 51: | Line 51: | ||
==Molecular dynamics theory== | ==Molecular dynamics theory== | ||
| - | + | ||
| - | + | Molecular dynamics simulation consists of the numerical, step-by-step, solution of the classical equations of motion. For this purpose we need to be able to calculate the forces acting on the atoms,and these are usually derived from a potential energy. This potential energy can be divided into: | |
| + | * the non-bonded interactions: The Lennard-Jones potential is the most commonly used form. It takes into account the Van der Waals forces. It represents the non-bonded forces and the total potential energy can be calculated from the sum of energy contributions between pairs of atoms. | ||
[[Image:lennard_jones_vdw_forces.jpg|frame|center|Lennard Jones potential ]] | [[Image:lennard_jones_vdw_forces.jpg|frame|center|Lennard Jones potential ]] | ||
| + | |||
| + | * the bonded interactions | ||
| + | |||
| + | |||
| + | The protein moves thanks to different forces, which can be separated in bonded forces and non-bonded forces. The bonded forces correspond to | ||
| + | * the ''Lennard-Jones potential'', | ||
* another force is the well-known ''Coulomb force'' | * another force is the well-known ''Coulomb force'' | ||
[[Image:Coulomb.jpg|frame|center|Coulomb force ]] | [[Image:Coulomb.jpg|frame|center|Coulomb force ]] | ||
| - | |||
==To envisage == | ==To envisage == | ||
Revision as of 09:13, 23 July 2009
Contents |
Modeling
To do
- - Model allosteric interactions between LOVTAP & TrpR
- What will be done:
- - Model of LOVTAP in dark phase
- - Model of LOVTAP in light phase
- - Characterize how the J-alpha helix changes
- - Model sturctural changes that enhance the switch feature of LOVTAP e.g. in dark phase: really weak interaction between LOVTAP and the corresponding DNA sequence, in light phase: strong binding of LOVTAP on DNA.
Modeling reference
LOVTAP simulation
We will follow the following article protocol:
Freddolino, P.L., Dittrich M., Schulten K., Dynamic Switching Mechanisms in LOV1 and LOV2 Domains of Plant Phototropins. Biophysical Journal, 91, 3630-3639, 2006 (Pubmed)
VMD informations
VMD is used to visualize molecules. It is quite user friendly.
- A tutorial for VMD can be found here.
NAMD informations
NAMD performs minimization and equilibration.
- A tutorial is on the same page as for VMD, here.
- NAMD 2.7b1 User's Guide
Run a simulation
A simulation is composed of different steps. Here are a few links that deal with heating and stabilization.
- Building Gramicidin A: Equilibration: protocol uses a single .conf file, heating process is too fast.
- NAMD notes from Robinson Lab: a really nice heating process, but involves different .conf files, which is really painful.
Implementation of the simulation
LOV domains are the light-sensitive portion of phototropins. They absorb light through a flavin cofactor, photo-chemicaly form a covalent bond between the chromophore and a cysteine residue in the protein, and proceed to mediate activation of an attached kinase domain.
Generating input files
First we need a compatible .pdb in addition to parameter and topology files. Steps to generate all the input files are explained in detail on this page How to generate input files. This is a kind of summary of the tuto.
.conf parameters
We should explain here what are the keywords we use in the .conf.
Run a complete simulation
We start from .pdb, .psf, .rtf generated in the previous section. Complete process is on a separate page How to run a simulation.
Molecular dynamics theory
Molecular dynamics simulation consists of the numerical, step-by-step, solution of the classical equations of motion. For this purpose we need to be able to calculate the forces acting on the atoms,and these are usually derived from a potential energy. This potential energy can be divided into:
- the non-bonded interactions: The Lennard-Jones potential is the most commonly used form. It takes into account the Van der Waals forces. It represents the non-bonded forces and the total potential energy can be calculated from the sum of energy contributions between pairs of atoms.
- the bonded interactions
The protein moves thanks to different forces, which can be separated in bonded forces and non-bonded forces. The bonded forces correspond to
- the Lennard-Jones potential,
- another force is the well-known Coulomb force
To envisage
- Molecular mutationnal assay
Already done
Here is our first movie from the modeling, showing the behavior of the protein in the dark state condition: Dark State
After having modified some parameters in the parameter files, here is our second movie, concerning the light state of the protein this time, with the FMN: Light State with FMN without water
 "
"