Team:EPF-Lausanne/Strategy
From 2009.igem.org
(→Cloning strategy) |
|||
| (20 intermediate revisions not shown) | |||
| Line 3: | Line 3: | ||
<div CLASS="epfltrick">__TOC__ | <div CLASS="epfltrick">__TOC__ | ||
</div> | </div> | ||
| - | <div CLASS=" | + | <div CLASS="epfl09incub"> |
| + | |||
| + | <html><br> | ||
| + | <br> | ||
| + | <br><br><br> | ||
| + | <br><center> | ||
| + | |||
| + | <a href="https://2009.igem.org/Team:EPF-Lausanne/LOVTAP" | ||
| + | onMouseOver="document.MyImage7.src='https://static.igem.org/mediawiki/2009/thumb/d/d4/Theory_lov.jpg/100px-Theory_lov.jpg';" onMouseOut="document.MyImage7.src='https://static.igem.org/mediawiki/2009/thumb/5/52/Theory_lov_nb.jpg/100px-Theory_lov_nb.jpg';"> | ||
| + | <img src="https://static.igem.org/mediawiki/2009/thumb/5/52/Theory_lov_nb.jpg/100px-Theory_lov_nb.jpg" name="MyImage7"></a> | ||
| + | |||
| + | | ||
| + | |||
| + | <a href="https://2009.igem.org/Team:EPF-Lausanne/LOVTAP_Results" | ||
| + | onMouseOver="document.MyImage6.src='https://static.igem.org/mediawiki/2009/thumb/b/ba/Results_lab.jpg/100px-Results_lab.jpg';" onMouseOut="document.MyImage6.src='https://static.igem.org/mediawiki/2009/thumb/f/f1/Results_lab_nb.jpg/100px-Results_lab_nb.jpg';"> | ||
| + | <img src="https://static.igem.org/mediawiki/2009/thumb/f/f1/Results_lab_nb.jpg/100px-Results_lab_nb.jpg" name="MyImage6"></a> | ||
| + | |||
| + | | ||
| + | |||
| + | <a href="https://2009.igem.org/Team:EPF-Lausanne/Future_directions" onMouseOver="document.MyImage5.src='https://static.igem.org/mediawiki/2009/thumb/5/5b/Future.jpg/100px-Future.jpg';" onMouseOut="document.MyImage5.src='https://static.igem.org/mediawiki/2009/thumb/c/cd/Future_nb.jpg/100px-Future_nb.jpg';"> | ||
| + | <img src="https://static.igem.org/mediawiki/2009/thumb/c/cd/Future_nb.jpg/100px-Future_nb.jpg" name="MyImage5"></a> | ||
| + | |||
| + | |||
| + | | ||
| + | |||
| + | |||
| + | <a href="https://2009.igem.org/Team:EPF-Lausanne/References" onMouseOver="document.MyImage9.src='https://static.igem.org/mediawiki/2009/thumb/a/a2/Ref.jpg/100px-Ref.jpg';" onMouseOut="document.MyImage9.src='https://static.igem.org/mediawiki/2009/thumb/2/25/Ref_nb.jpg/100px-Ref_nb.jpg';"> | ||
| + | <img src="https://static.igem.org/mediawiki/2009/thumb/2/25/Ref_nb.jpg/100px-Ref_nb.jpg" name="MyImage9"></a> | ||
| + | |||
| + | </center> | ||
| + | </html> | ||
| + | <br> | ||
| + | ---- | ||
| + | <br> | ||
<html><center> | <html><center> | ||
| - | <font size=" | + | <font size="12" color="#007CBC">Strategy</font> |
</center></html> | </center></html> | ||
<br> | <br> | ||
---- | ---- | ||
| - | |||
<br> | <br> | ||
==Cloning strategy== | ==Cloning strategy== | ||
| + | |||
| + | <font size="3">'''LovTap BioBrick'''</font> | ||
| + | |||
Our aim was to create a biobrick containing LovTAP under the influence of an inducible promoter. This part needed to contain (in order) the inducible promoter (LacI), RBS (ribosome binding site), the LovTAP gene (that was generously sent by Pr. Sosnick form the University of Chicago) and finally a terminator (Term). This entire part was of course to be flanked by the standard prefix (E,X) and suffix (S,P). In order to synthesize this part, we proceeded in three steps : | Our aim was to create a biobrick containing LovTAP under the influence of an inducible promoter. This part needed to contain (in order) the inducible promoter (LacI), RBS (ribosome binding site), the LovTAP gene (that was generously sent by Pr. Sosnick form the University of Chicago) and finally a terminator (Term). This entire part was of course to be flanked by the standard prefix (E,X) and suffix (S,P). In order to synthesize this part, we proceeded in three steps : | ||
| Line 30: | Line 65: | ||
With this biobrick, we have an inducible system : when we add IPTG, the promoter activates the expression of the LovTAP protein. | With this biobrick, we have an inducible system : when we add IPTG, the promoter activates the expression of the LovTAP protein. | ||
| - | ==Read | + | |
| + | |||
| + | <font size="3">'''Read-Out n°1 BioBrick'''</font> | ||
| + | |||
| + | This BioBrick was synthesized using the protocol we developed with the Klenow fragment. To do this, two long primers which together contained the sequence for the Trp promoter were ordered; these primers could self-anneal in the reaction mix, and the second strands that were missing at the extremities of the primers were fully synthesized using the Klenow fragment, instead of a classic extension using the TAQ DNA Polymerase. | ||
| + | |||
| + | For more information, here is the [https://2009.igem.org/wiki/index.php?title=Team:EPF-Lausanne/Protocols/Klenow complete protocol]. | ||
| + | |||
| + | |||
| + | |||
| + | <font size="3">'''Read-Out n°2 BioBrick'''</font> | ||
| + | |||
| + | This BioBrick was created using our new method, which we named "1.5-step PCR". This method is similar to a 2-step PCR, however the difference is that in the 1.5-step, the first cycle of amplification is omitted, so that in the initial step we directly add the longer primers to amplify the DNA from the plasmid. The last step is the same as for the 2-step PCR, where you add the iGEM (short) primers to amplify the entire sequence, from the extremities of the long primers which had been added previously. By suppressing one step of the "classic" 2-step PCR protocol, it allows us to save a considerable amount of time. | ||
| + | |||
| + | For more details on this method, look at the [https://2009.igem.org/wiki/index.php?title=Team:EPF-Lausanne/Protocols/1.5_steps_PCR protocol]. | ||
| + | |||
| + | ==Read-Out systems== | ||
<div style="text-align:justify;"> | <div style="text-align:justify;"> | ||
Once we have our protein LovTAP produced, we need a read out system to assess whether the protein was functional or not. | Once we have our protein LovTAP produced, we need a read out system to assess whether the protein was functional or not. | ||
| Line 46: | Line 97: | ||
[[Image: systems.jpg|800 px|center|Readouts]] | [[Image: systems.jpg|800 px|center|Readouts]] | ||
| + | |||
| + | ==Modeling== | ||
| + | |||
| + | In order to anticipate the output of the Read Out 2 system, we made a mathematical model of this circuit and solved this first order ordinary differential equation system. | ||
| + | |||
| + | The system was modeled as [https://2009.igem.org/wiki/index.php?title=Team:EPF-Lausanne/ODEmodeling explained here] | ||
| + | |||
| + | |||
| + | [[Image:FluoRed.png|frame|center|Simulation of the Read Out 2 system and experimental data taken from [https://2009.igem.org/Team:EPF-Lausanne/LOVTAP_Results#Characterization_of_the_entire_system Experiment N°2]]] | ||
| + | |||
Latest revision as of 16:33, 21 October 2009




Cloning strategy
LovTap BioBrick
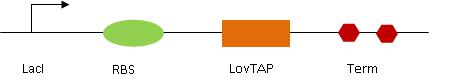
Our aim was to create a biobrick containing LovTAP under the influence of an inducible promoter. This part needed to contain (in order) the inducible promoter (LacI), RBS (ribosome binding site), the LovTAP gene (that was generously sent by Pr. Sosnick form the University of Chicago) and finally a terminator (Term). This entire part was of course to be flanked by the standard prefix (E,X) and suffix (S,P). In order to synthesize this part, we proceeded in three steps :
1. We ligated the LovTAP gene into the terminator (we used a double terminator).
2. We also ligated LacI into RBS.
3. Finally we ligated these two parts together to obtain our biobrick (LacI-RBS in the LovTAP-term part).
So finally we got :
With this biobrick, we have an inducible system : when we add IPTG, the promoter activates the expression of the LovTAP protein.
Read-Out n°1 BioBrick
This BioBrick was synthesized using the protocol we developed with the Klenow fragment. To do this, two long primers which together contained the sequence for the Trp promoter were ordered; these primers could self-anneal in the reaction mix, and the second strands that were missing at the extremities of the primers were fully synthesized using the Klenow fragment, instead of a classic extension using the TAQ DNA Polymerase.
For more information, here is the complete protocol.
Read-Out n°2 BioBrick
This BioBrick was created using our new method, which we named "1.5-step PCR". This method is similar to a 2-step PCR, however the difference is that in the 1.5-step, the first cycle of amplification is omitted, so that in the initial step we directly add the longer primers to amplify the DNA from the plasmid. The last step is the same as for the 2-step PCR, where you add the iGEM (short) primers to amplify the entire sequence, from the extremities of the long primers which had been added previously. By suppressing one step of the "classic" 2-step PCR protocol, it allows us to save a considerable amount of time.
For more details on this method, look at the protocol.
Read-Out systems
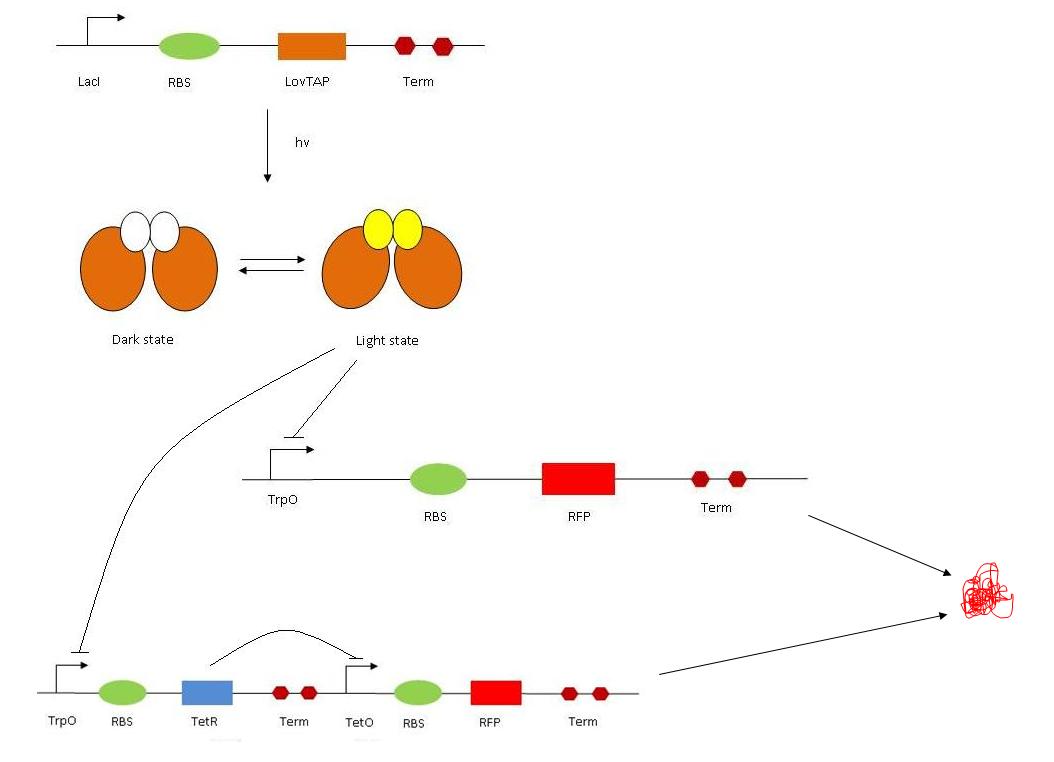
Once we have our protein LovTAP produced, we need a read out system to assess whether the protein was functional or not. Therefore, we designed two different read out systems :
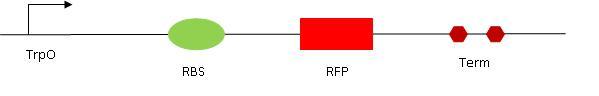
1. The read out 1 (RO1) contains the tryptophan operon followed by RBS, RFP and Term. In normal conditions, RFP should be expressed (so we should see some red fluorescence). This is because the Trp promoter is constitutively ON. The Trp repressor (TrpR) only binds to DNA if there is tryptophane available in the medium. The activated LovTAP protein (when illuminated at 470 nm) binds to the Trp operon and repress the RFP gene, exactly as the TrpR in presence of Trp would do. We should therefore observe a decrease in red fluorescence. In order to characterize the read out, we tested it in different conditions : with or without TRP, with different amount of TRP. With addition of Trp, we would expect the fluorescence to decrease.
2. The read out 2 (RO2) is composed of the Trp op, RBS, TetR (gene), Term and then TetR op, RBS, RFP and Term. It is a double repressor system : TrpR/LovTAP bind to the Trp op, therefore repressing the expression of the tetracyline repressor (TetR). On the other hand, TetR (if expressed) inhibits the production of RFP by acting on the TetR op. The final result is that, if LovTAP is active, red fluorescence should increase as RFP is expressed. Trp has the same effect as LovTAP. For the characterization of the system, we also used ATC (anhydrotetracyclin). ATC inhibits the TetR protein, making it unable to bind to its operator sequence. Therefore, if present, it has the same effect has inhibiting the TetR protein synthesis. The overall effect is then the same as Trp(LovTAP), when it present, the fluoresence increases. Eventually, we have three ways of modulating the fluorescence : adding Trp, shining light (if LovTAP is present) and adding ATC would all make the red fluorescence to increase.
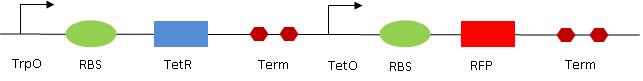
So everything together gives the following system :
Modeling
In order to anticipate the output of the Read Out 2 system, we made a mathematical model of this circuit and solved this first order ordinary differential equation system.
The system was modeled as explained here

 "
"