Team:Newcastle/Chassis
From 2009.igem.org
(→Introduction) |
|||
| Line 122: | Line 122: | ||
==Lab Work Strategies== | ==Lab Work Strategies== | ||
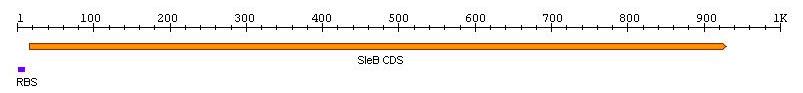
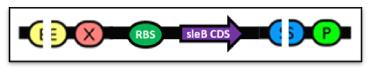
| - | + | [[Image:TeamNewcastleChassisFigure1.jpg|center|450px|thumb|<center>Figure 1: RBS + ''sleB''</center>]] | |
| - | + | ||
| - | [[Image:TeamNewcastleChassisFigure1.jpg|center| | + | |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| + | To find out more about our lab strategies, click [https://2009.igem.org/Team:Newcastle/Chassis/LabStrategies ''here''] | ||
==<b>References:</b>== | ==<b>References:</b>== | ||
Revision as of 20:47, 21 October 2009
Chassis
Introduction
The main aim of our project is to sequester cadmium in the environment into the spores of our engineered B. subtilis, but what happens after the cadmium has been sequestered?
Do we attempt to retrieve the sequestered cadmium? Or, do we simply leave the sequestered cadmium in the spores of our engineered B. subtilis?
For our project, we have chosen the latter. We will not be attempting to retrieve the sequestered cadmium. However, then comes the question of, would there not be chances of the cadmium entering the environment again?
Our solution to this question would be to disable germination of the spores, thus retrieval of the sequestered cadmium becomes unnecessary, as the spores can persist intact for thousands of years.
In order to disable germination of the spores, we would require non-germinating spores, and we were fortunate enough that Prof. Anne Moir from Sheffield University kindly sent us two non-germination spores, with inactivated genes, namely cwlD, and sleB and cwlJ.
While we would like to disable germination for the spores that contain sequestered cadmium, not all the cells would have sequestered cadmium, and it is also essential that we still have some cells germinating, so that our population of bacteria can continue to live and grow, reaching a balance, and not simply deplete totally.
Therefore, a mechanism is needed to allow us to choose to turn on germination, when the cell is not a "metal container".
Using the treatment protocol for the non-germination spores from Prof. Anne Moir, we performed lab experiments for the two non-germination spores, and concluded that the double-knockout mutant, sleB and cwlJ would be more ideal for our project as it had more colonies growing after treatment, and less colonies growing without treatment, as compared to the single knock-out mutant, cwlD.
We propose that we could use IPTG as a switch for germination.
Novelty in this sub-project
In this sub-project, we are disabling germination, using non-germinating spores with the inactivated genes, sleB and cwlJ. In order to control germination, we intend to use IPTG as a switch, via the promoter, pSpac
Wet Lab
Click on the dates to go the the particular lab session.
| Summary of Lab Sessions for Chassis | |
|---|---|
| | |
| 04/08/09 | Arrival of the non-germination spores. Preparation of the buffer solution required for the treatment of the spores |
| 07/08/09 | Preparation of the lysozyme stock solution required for treatment of the spores |
| 10/08/09 | Re-preparation of the buffer solution required for the treatment of the spores. Pouring of agar plates with the appropriate antibiotics. |
| 11/08/09 | Re-pouring the agar plates with the appropriate antibiotics |
| 12/08/09 | Treatment of the non-germinating cwlD spores using Method A |
| 13/08/09 | Results for the treatment of the cwlD spores using Method A |
| 17/08/09 | Repeat experiment for the treatment of the cwlD spores using Method A |
| 18/08/09 | Successful results for the treatment of the cwlD spores using Method A. Performed treatment for the double-knockout mutants sleB and cwlJ spores using Method A |
| 19/08/09 | Successful results for the treatment of the double-knockout mutants sleB and cwlJ spores using Method A |
| 25/08/09 | Freezing down of the treated non-germinating spores, cwlD, and sleB and cwlJ. |
| 02/09/09 | PCR-ing of gene sleB and cwlJ using primers previously designed and ordered. |
| 03/09/09 | Attempt to PCR-ing of gene sleB and cwlJ using primers previously designed and ordered again. |
| 04/09/09 | Redesign PCR primers |
| 08/09/09 | Cloning of sleB |
BioBrick constructs
BBa_K174012
sleB, Bacillus subtilis germination gene with RBS
Length: 932bp
Click [http://partsregistry.org/wiki/index.php?title=Part:BBa_K174012 here] for more information on this part.
BBa_K174013
cwlJ, Bacillus subtilis germination gene with RBS
Length: 441bp
Click [http://partsregistry.org/wiki/index.php?title=Part:BBa_K174013 here] for more information on this part.
BBa_K184014
cwlJ and sleB, Bacillus subtilis germination genes
Click [http://partsregistry.org/wiki/index.php?title=Part:BBa_K174014 here] for more information on this part.
Lab Work Strategies
To find out more about our lab strategies, click here
References:
[1] Popham, D., Helin, J., Costello, C & Setlow, P. (1996). Muramic lactam in peptidoglycan of Bacillus subtilis spores is required for spore outgrowth but not for spore rehydration or heat resistance. Proc. Natl. Acad. Sci. 93; 15403-15410
[2] Chirakkal, H., O'Rourke, M., Atrih, A., Foster, S. J., Moir, A. (2002.) Analysis of spore cortex lytic enzymes and related proteins in Bacillus subtilis endospore germination. Microbiology 148; 2383-2392
News
Events
- 20 – 21 June 2009 - Europe workshop (London)
- 23 – 24 June 2009 - UK iGEM meetup (Edinburgh)
- 23 October Practice Presentation (Newcastle)
- 23 October T-shirts are ready
- 27 October Practice Presentation (Sunderland)
- 27 October Poster is ready
- 30 October – 2 November 2009 - Jamboree (Boston)
Social Net
- Newcastle iGEM Twitter
- [http://www.facebook.com/home.php#/group.php?gid=131709337641 Newcastle on Facebook]
- [http://www.youtube.com/user/newcastle2009igem Newcastle Youtube Channel]
 "
"