Team:Newcastle/Labwork/11 August 2009
From 2009.igem.org
Contents |
Lab Session 11/08/09
Sporulation Tuning/ Chassis team
Previously, we failed to pour the LB + Em + Kan plates successfully due to the smaller amount of volume. Today, from our lessons learnt, we attempted again, and it was a success.
This time however, we set up the water bath to a temperature of 55oC and placed the autoclaved LB + agar solution in it. The water bath ensured that the LB + agar solution would cool to only a minimum of 55oC and the same mistake of the mixture solidifying upon addition of the cold water would not happen.
Stochastic Switch Team
Today we run the gel for our gfp-rrnb integration vector. We had restriction digest using EcoRI and HindIII. This gave us a 500bp of DNA on the gel.
Preparation of the gel
- 3.2 gr of agarise were solved in 400ml of 1xTAE buffer to make 0.8% agarose gel
- The solution was microwaved for 5 minutes and left to cool down for an hour.
Restriction digest
- We prepared a final solution of 10ul for the restriction digest
- We used
- 4.75ul H2O
- 1ul 10xBuffer(BufferE)
- 0.25ul BSA
- 0.5ul HindIII
- 0.5ul EcoRI
- 3ul DNA (gfp-rrnb)
- The solutions were incubate for an hour at 37C.
Running the gel
- we cleaned the comb and the tray
- We used 100ml of the agarose gel
- We poured the solution into a flask tube and added 0.8ul of ethium bromide.
- Poured the final solution into the tray. It was left for 15 minutes
- Loaded the wells with DNA ladder, uncut DNa, cut DNA and the ladder again. Because of the yellow band on the tray, it was very difficult to load the samples./ We intended to leave some gap between uncut and cut DNAs but we loaded them next to each other. The wells were also closed a bit. This situation may have been because the gel was not left long enought to set.
- Run the gel for 0.5 hour. But for a better resolution we decided to run it for another 0.5 hour.
Results
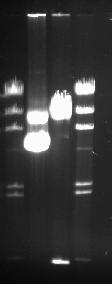
We saw the small 500bp DNA in the cut DNA's lane. Our DNA was highly concentrated and as shown below, the images are highly saturated.
The order of the lanes from left to right: DNA ladder, Uncut DNA, Cut DNA, DNA ladder
Metal Sensor Team
Introduction and summary
The Metal Sensor team have been given the task of test-transforming Bacillus subtilis 168 with vector plasmid GFP-rrnb. This involves the Bacillus transformation protocol. So far we have made up all of the solutions involved in both the MM competence medium and starvation medium and have grown overnight cultures of Bacillus subtilis in MM competence medium. We have also incubated a falcon tube containing MM competence media without B. subtilis to act as a contamination control. Today, the rest of the Bacillus transformation protocol will be completed. In addition to this, LB plates containing chloramphenicol will need to be produced.
News
Events
- 20 – 21 June 2009 - Europe workshop (London)
- 23 – 24 June 2009 - UK iGEM meetup (Edinburgh)
- 23 October Practice Presentation (Newcastle)
- 23 October T-shirts are ready
- 27 October Practice Presentation (Sunderland)
- 27 October Poster is ready
- 30 October – 2 November 2009 - Jamboree (Boston)
Social Net
 "
"