Team:Newcastle/Labwork/14 September 2009
From 2009.igem.org
Lab Work - 14/09/09
Metal Sensing team
Introduction
In the last lab session the team carried out a successful PCR reaction on the cotC-GFP-smtA BioBrick and cleaned up the products. The team also midi-prepped cotC-GFP-smtA, kinA, pGFP-rrnB, pMUTIN4 and pSB1AT3 which will prove useful for other team members too. In addition, transformant E. coli cells containing the kinA BioBrick and the cotC-GFP-smtA BioBrick were grown up in 5ml LB cultures and then frozen down.
However there were also things that we didn't accomplish. We had previously carried out a PCR reaction involving the two elements of our cadmium-sensitive logic AND gate - arsR and cadA. But when the products were cleaned a large amount of ethanol remained in the sample and so when placed into agarose gel the sample dispersed. We then proceded to carry out a second attempt at the same PCR reaction.
In today's practical we hope to ligate pMUTIN4 and the cotC-GFP-smtA BioBrick together - firstly cutting both fragments with BamHI and HindIII and then ligating the DNA together. We also hope to run the arsR and cadA promoter PCR products on agarose gel (in DNA gel electrophoresis) and if successful, clean up the sample. Then run all of the sample through agarose gel and excise the band produced, also extracting the gel from it afterwards. If all these events happen, we will also cut both the arsR and cadA fragments with BamHI and NheI and attempt overnight ligation.
Practical Outline
Today's lab session will continue straight on from the last lab session and fulfil the following objectives:
- Run products of PCR reaction involving the arsR BioBrick (BBa_J33206 in pSB1A2 missing promoter) and cadA promoter region
- If successful clean up product, run on agarose gel through electrophoresis, excise band and carry out gel extraction.
- Cut cleaned and excised fragment with BamHI and NheI
- Carry out ligation (possibly an overnight ligation)
- Cut pMUTIN4 and cotC (i.e. PCR product 3) with BamHI and HindIII
- Ligate the fragments (possibly overnight)
Procedure
1) Cadmium-sensing logic AND gate
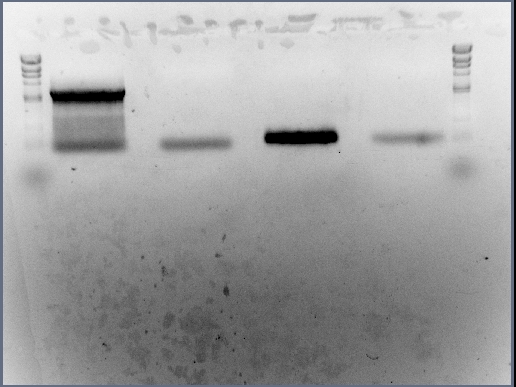
The first step to take was to run the PCR products of reactions involving the arsR BioBrick (BBa_J33206 without it's original promoter) and the cadA promoter region through agarose gel in DNA gel electrophoresis. The initial views of the team was to carry out a test on the samples on some gel before excising the rest of the DNA when in gel. However we decided to place all of the DNA into gel and excise the band immediately, without conducting tests.
All of the DNA from the PCR reactions involving the cadA promoter region and the arsR BioBrick (BBa_J33206 without promoter) was inserted into agarose gel (in two separate wells of course) and DNA gel electrophoresis allowed to commence for 20 minutes under 80 volts.
We also ran ara and pGFP-rrnB DNA on the gel (this is for the Stochastic Switch team).
The bands were then excised and cleaned using Sigma-Aldrich's GenElute Gel Extraction kit (along with the protocol). There was no changes to the protocol but a few important points to note:
- The 'Spin Method' was the preferred route in the protocol to follow.
- The mass of the gel bands were as follows:
- arsR band = 0.285 grams
- cadA promoter region band = 0.354 grams
- ara band = 0.212 grams
- pGFP-rrnB band = 0.233 grams
- arsR band = 860ul of solution to be added
- cadA promoter region band = 1060ul of solution to be added
- ara band = 640ul of solution to be added
- pGFP-rrnB band = 700ul of solution to be added
- arsR band = 285ul of isopropanol to be added
- cadA promoter region band = 354ul of isopropanol to be added
- ara band = 212ul of isopropanol to be added
- pGFP-rrnB band = 233ul of isopropanol to be added
Once the arsR (the BBa_J33206 BioBrick without promoter region) and cadA promoter DNA samples were cleaned, they were both cut using BamHI and NheI in the following way:
| arsR BioBrick | cadA promoter | ||
|---|---|---|---|
| DNA (ul) | 10 | 10 | |
| Restriction Buffer (ul) | 2 | 2 | |
| BamHI Restriction Enzyme(ul) | 1 | 1 | |
| NheI Restriction Enzyme(ul) | 1 | 1 | |
| Distilled water(ul) | 6 | 6 |
The two fragments then underwent a FastLigation reaction. This ligation only took 10 minutes. This is what we inserted into the ligation reaction:
- Linearised vector DNA = BBa_J33206 (arsR) without promoter - 1ul
- Insert DNA = cadA promoter region - 7ul
- 5x Rapid Ligation Buffer = 4ul
- Water = 6ul
- T4 DNA ligase = 1ul
Note: cadA promoter region DNA concentration was 17ng/ul; arsR BioBrick DNA concentration was 40ng/ul
2) Ligating the cotC BioBrick with pMUTIN4
Firstly the cotC BioBrick and the pMUTIN4 DNA was cut with BamHI and HindIII:
| cotC BioBrick (reaction 1) | cotC BioBrick (reaction2) | pMUTIN4 plasmid | ||
|---|---|---|---|---|
| DNA (ul) | 30 | 10 | 20 | |
| Restriction Buffer (ul) | 5 | 5 | 5 | |
| BamHI Restriction Enzyme(ul) | 1 | 1 | 1 | |
| HindIII Restriction Enzyme(ul) | 1 | 1 | 1 | |
| Distilled water(ul) | 13 | 33 | 23 |
These digests were left to commence for 1 hour. Once this time had elapsed the two fragments of DNA were ligated together in the following way:
- 1ul of pMUTIN4
- 6ul of cotC
- 4ul of buffer solution
- 1ul DNA ligase
- 7ul sterile distilled water
3) Transforming E. coli cells with two ligated products
The two ligated products - the arsR-cadA promoter AND gate and the cotC-GFP-smtA BioBrick in pMUTIN4 - were then transformed into DH5-alpha E. coli cells using Phil's transformation by heat shock protocol.
News
Events
- 20 – 21 June 2009 - Europe workshop (London)
- 23 – 24 June 2009 - UK iGEM meetup (Edinburgh)
- 23 October Practice Presentation (Newcastle)
- 23 October T-shirts are ready
- 27 October Practice Presentation (Sunderland)
- 27 October Poster is ready
- 30 October – 2 November 2009 - Jamboree (Boston)
Social Net
 "
"