Team:Newcastle/Labwork/26 August 2009
From 2009.igem.org
Contents |
Lab Work - 26/08/09
Stochastic Switch team
Summary
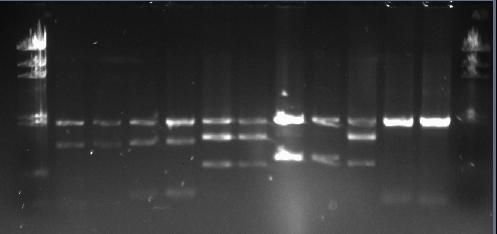
Today Lab started at 2pm, so we had the afternoon to do restriction digests for the remaining biobricks(other than C0178). We used the fast digest system, which takes 5 minutes in a 37 degree waterbath rather than 60 minutes. Below are the volumes required for a double digest using EcoRI and PstI:
- 6ul water
- 2ul 10X Fast digest buffer
- 1ul of fast digest EcoRI enzyme
- 1ul of fast digest PstI enzyme
- 10ul DNA (miniprepped)
Total volume: 20ul
These were left for 5-10 minutes at 37 degrees, then 9ul of the total volume was added to 1ul of loading buffer, and the samples were run on the gel we prepared yesterday.
Today we also prepared some overnight cultures of DH5alpha cells (2X 3ml plain LB) for the Genomic DNA prep which will be carried out tomorrow.
Metal Sensing Team team
Introduction
Practical Outline
The following list consists of the tasks the Metal Sensing team hope to achieve by the end of the day:
- Carry out the restiction enzyme digests of the midi-prep plasmid DNA (pSB1A2 containing BioBrick BBa_J33206) and run them on the gel.
- Carry out the genomic DNA preparations.
Procedure
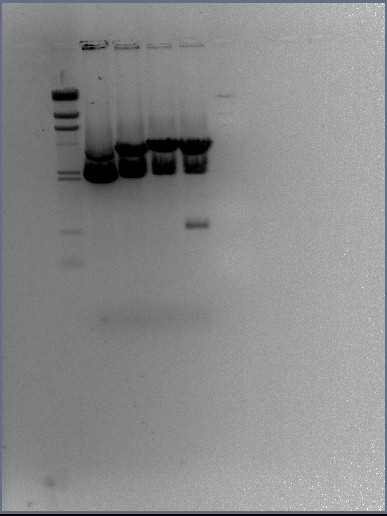
Restriction enzyme digests of the midi-prep DNA
DNA gel electrophoresis of digests
Results
Promoter Library Sub-Project
The aim of this sub-project is to create a library of sigA promoters which differ in strength and to characterise them.
The first step in this process is to digest the pGFP-rrnb plasmid with restriction enzymes EcoRI and NheI. The enzyme digests were prepared in 1 Eppendorf tube in the following way:
- 15ul of pGFP-rrnb plasmid DNA (from a midiprep)
- 3ul NheI (FastDigest)
- 3ul EcoRI (FastDigest)
- 4ul FastDigest buffer
- 15ul distilled water
This makes up a total volume of 40ul The mixture was made in such a way that the DNA, the distilled water and the FastDigest buffer was added first before the enzymes were added. The tube was given a spin pulse and placed in a water bath at 37C for 1 hour.
After this the mixture was placed in the -20C freezer for use tomorrow.
In the meantime, 0.8% agarose gel was prepared in the usual way and poured into the gel tray. The only difference between this gel and others previously poured is that some of the comb was taped so that the wells formed were larger. Once the gel had set, 1xTAE buffer was sprinkled onto the surface and the gel wrapped up in cling film. The gel was placed in the fridge also for use tomorrow.
News
Events
- 20 – 21 June 2009 - Europe workshop (London)
- 23 – 24 June 2009 - UK iGEM meetup (Edinburgh)
- 23 October Practice Presentation (Newcastle)
- 23 October T-shirts are ready
- 27 October Practice Presentation (Sunderland)
- 27 October Poster is ready
- 30 October – 2 November 2009 - Jamboree (Boston)
Social Net
- Newcastle iGEM Twitter
- [http://www.facebook.com/home.php#/group.php?gid=131709337641 Newcastle on Facebook]
- [http://www.youtube.com/user/newcastle2009igem Newcastle Youtube Channel]
 "
"