Team:Newcastle/Labwork/28 August 2009
From 2009.igem.org
Babyneurone (Talk | contribs) (→PCR) |
Babyneurone (Talk | contribs) (→PCR) |
||
| Line 33: | Line 33: | ||
| 50 | | 50 | ||
|- | |- | ||
| - | + | |1 | |
| - | | | + | |27 |
| - | | | + | |15 |
| - | | | + | |pGFPconf2 [2.5] |
| + | |pGFPconf3 [2.5] | ||
| + | |168 DNA [2] | ||
| + | |Moltaq [1] | ||
| + | | 50 | ||
| + | |- | ||
| + | |1 | ||
| + | |27 | ||
| + | |15 | ||
| + | |pGFPconf2 [2.5] | ||
| + | |pGFPconf3 [2.5] | ||
| + | |168 DNA [2] | ||
| + | |Moltaq [1] | ||
| + | | 50 | ||
| + | |- | ||
| + | |1 | ||
| + | |27 | ||
| + | |15 | ||
| + | |pGFPconf2 [2.5] | ||
| + | |pGFPconf3 [2.5] | ||
| + | |168 DNA [2] | ||
| + | |Moltaq [1] | ||
| + | | 50 | ||
| + | |- | ||
| + | |1 | ||
| + | |27 | ||
| + | |15 | ||
| + | |pGFPconf2 [2.5] | ||
| + | |pGFPconf3 [2.5] | ||
| + | |168 DNA [2] | ||
| + | |Moltaq [1] | ||
| + | | 50 | ||
| + | |- | ||
| + | |1 | ||
| + | |27 | ||
| + | |15 | ||
| + | |pGFPconf2 [2.5] | ||
| + | |pGFPconf3 [2.5] | ||
| + | |168 DNA [2] | ||
| + | |Moltaq [1] | ||
| + | | 50 | ||
==<u>Stochastic Switch team</u>== | ==<u>Stochastic Switch team</u>== | ||
Revision as of 13:47, 8 September 2009
Contents |
Lab Work - 28/08/09
Metal Sensing Team
Summary
Today's lab experiment will be focussing on a second attempt to PCR the genome of Bacillus subtilis 168 cells which have taken up and inserted the pGFP-rrnB plasmid. In a couple of the previous lab sessions, the Metal Sensing team focussed on growing up some B. subtilis transformant cells (i.e. those which had taken up the plasmid pGFP-rrnB) and carrying out a PCR reaction on them using four synthesized primers to make sure that the genome carries out the insert.
PCR
Exp.No.2 We used the same protocol as used on the 21/08/2009 to carry out our PCR. The only difference is that this time we have only 6 samples, and the annealing temperature will be changed to 50 degree ( instead of the previous 56 degree). Also, we got some B.subtilis genomic DNA from Prof.Colin Hardwood's lab; last time the PCR reaction was carried out on crude cell extracts.
Sample 1 : B.subtilis 168 genomic DNA + pGFPconf2 + pGFPconf3 = control Sample 2 : B.subtilis transformed with pGFP-rrnB + pGFPconf2 + pGFPconf1 Sample 3 : B.subtilis transformed with pGFP-rrnB + pGFPconf4 + pGFPconf3 Sample 4 : B.subtilis transformed with pGFP-rrnB + pGFPconf1 + pGFPconf4 Sample 5 : B.subtilis transformed with pGFP-rrnB + pGFPconf2 + pGFPconf3 Sample 6 : PCR water + pGFPconf2 + pGFPconf3 ( no template DNA ) = control
See Exp.No.2 PCR experiment Plan
| No. | H2O | PCR Mix | Primer Forward | Primer Reverse | Template | Pol.Used | Total Vol. |
|---|---|---|---|---|---|---|---|
| 1 | 27 | 15 | pGFPconf2 [2.5] | pGFPconf3 [2.5] | 168 DNA [2] | Moltaq [1] | 50 |
| 1 | 27 | 15 | pGFPconf2 [2.5] | pGFPconf3 [2.5] | 168 DNA [2] | Moltaq [1] | 50 |
| 1 | 27 | 15 | pGFPconf2 [2.5] | pGFPconf3 [2.5] | 168 DNA [2] | Moltaq [1] | 50 |
| 1 | 27 | 15 | pGFPconf2 [2.5] | pGFPconf3 [2.5] | 168 DNA [2] | Moltaq [1] | 50 |
| 1 | 27 | 15 | pGFPconf2 [2.5] | pGFPconf3 [2.5] | 168 DNA [2] | Moltaq [1] | 50 |
| 1 | 27 | 15 | pGFPconf2 [2.5] | pGFPconf3 [2.5] | 168 DNA [2] | Moltaq [1] | 50
Stochastic Switch teamSummaryToday the pSB1AT3 DNA fragment cut from the gel yesterday was purified using the gel fragment cleaning kit, and as well as this a PCR was carried out with both the ara primers on Bacillus genomic DNA, and the sspb primers on E.coli genomic DNA prepared yesterday. We expected to see a ~170bp band for the ara regulatory sequence, and a ~570bp band for the sspb coding sequence. We also continued to extract the plasmid backbone, pSB1AT3, cut with EcoRI and SpeI sites from the gel. PCRPCR was carried out with both the ara primers on Bacillus genomic DNA, and the sspb primers on E.coli genomic DNA prepared yesterday. We expected to see a ~170bp band for the ara regulatory sequence, and a ~570bp band for the sspb coding sequence. The Metal team were also doing PCR work today so we made our solutions as a whole to save time and resources. Overall there were 10 reactions. We did our two PCRs as well as negative controls that contained everything other than the genomic DNA.
Added:
These can be found in the 'stock solutions' box in the -20 freezer.
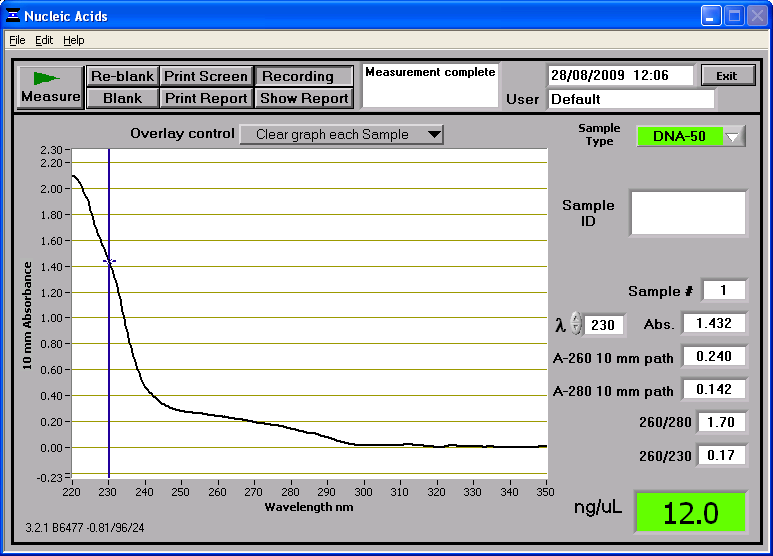
The final solutions can be put into the PCR machine and the appropriate program set. The PCR products can then be run on a gel which we did in the afternoon. DNA ExtractionWe took the eppendorf tube containing 184mg of gel from the fridge and solved the gel in 552ul(184*3) of gel solubilization solution. We left the tube containing the mix in 55C hot water bath for 10 minutes and vortexed every three minutes. We used two collection tubes and added 184 ul of isopropanol, 700ul of wash solution and 50ul of elute for each tube at the relevant steps. The first collection tube had 700ul of the solved gel whereas the second one had 450ul. 110ul of Elute was placed in an eppendorf tube and preheated in 65C prior to use. After adding the elute we left the tubes on the bench for a minute to incubate before the centrifuge. We got 100ul of DNA from the first tube and 80ul of from the second tube. We then combined the tubes, measured the DNA concentration and placed the tube to -20C fridge where midipreps are stored. DNA ConcentrationConcentration of the DNA was measured as 12ng/ul using the spectrophotometer Promoter Library Sub-ProjectIntroductionSo far, the work involved in this sub-project has seen pGFP-rrnB plasmid DNA digested with restriction enzymes EcoRI and NheI. The resulting 8kb DNA fragment was then extracted by firstly running the digested DNA through agarose gel in electrophoresis and then by excising the heavies band formed. The fragment DNA was then extracted from this removed piece of agarose by gel extraction kit and the 50ul of resulting DNA was stored in the -20C freezer. When running the digested pGFP-rrnB plasmid DNA though the gel to extract the 8Kb fragment, we did not see the bands expected to be seen. Therefore another restriction enzyme digestion of the pGFP-rrnB DNA with EcoRI and NheI was attempted (with different concentrations of the enzymes and DNA). In addition more pGFP-rrnB DNA was digested but with EcoRI and NheI separately. All of the resulting digests were stored in the -20C freezer. In today's experiments we hope to run 10ul of pGFP-rrnB treated with EcoRI and 10ul of pGFP-rrnB + NheI on some agarose gel for analysis. Additionally, 5ul of the 8Kb pGFP-rrnB digest fragment should be run on the gel. DNA gel Electrophoresis of samples0.8% agarose gel was prepared and poured in the manner exhibited in the DNA gel electrophoresis protocol. Once this step had been carried out, the DNA samples (once dye had been added) were loaded in the wells as follows:
Resulting gel photograph
News
Events
Social Net
|
 "
"