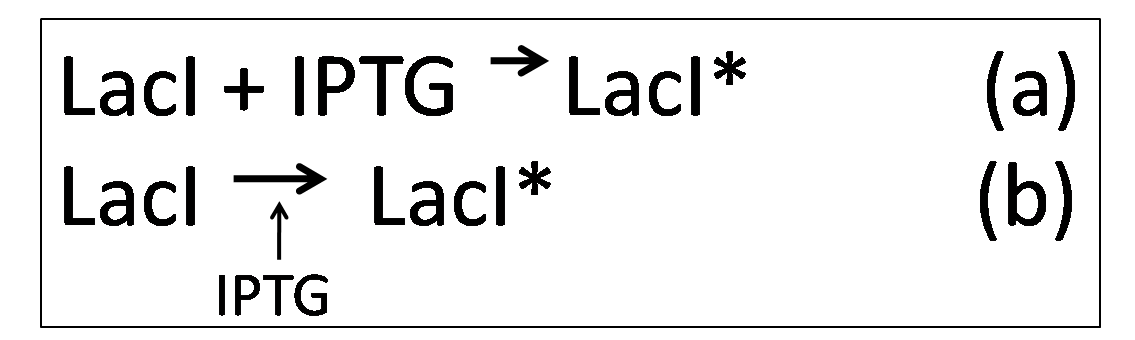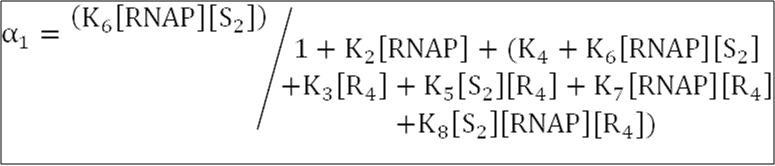Team:Newcastle/SporulationTuning
From 2009.igem.org
Sporulation Tuning
Introduction
In this section of our project, we hope to control sporulation in our bacterial population, such that we can decide how much of the population becomes spores, and how much continue as vegetative cells. Should the cell sporulate, it would become a ‘metal container’, trapping the sequestered cadmium in its spore.
After the cell sequesters cadmium into its spore, it should not germinate or the sequestered cadmium will be released back into the environment as a result. Therefore, the role of chassis comes into play, where the sleB and cwlJ germination-defective mutants are put into use. More information about this other sub-project of ours can be found here.
In order to control sporulation, our team is proposing the idea of inducing the synthesis of KinA, with IPTG as a sporulation initiation signal.
KinA is a major kinase which provides phosphate input to the phosphorelay, which in turn, activates the sporulation pathway upon starvation via the phosphorylated Spo0A transcription factor,[1] which governs entry into the sporulation pathways of the bacterium Bacillus subtilis.[2]
Spo0A
From our research, we found that entry into the sporulation pathway is governed by a member of the response regulator family of transcription facts, known as Spo0A.[2] Spo0A is activated by phosphorylation on an aspartyl residue located in the N-terminal portion of the protein.[2]
Unlike other response regulators, Spo0A is indirectly phosphorylated by a multicomponent phosphorelay involving at least three kinases called KinA, KinB, and KinC, which phosphorylate Spo0F, and the resulting Spo0F~P, in turn, transfers the phosphoryl group to Spo0B. Finally, Spo0B~P transfers the phosphoryl group to, and thereby activates, Spo0A.[2] As such, the phophoryl groups are drained from the relay by the action of dedicated phosphotases that dephosphorylate Spo0F~P and Spo0A~P.[2]
Spo0A is also subjected to control at the levels of its synthesis and activity by a positive feedback loop in which the response regulator stimulates the synthesis of the RNA polymerase σ factor σH, which, in turn stimulates transcription of the gene for Spo0A, as well as the genes for the phosphorelay components KinA and Spo0F.[1]
It is important to note that activating Spo0A via the phosphorelay is essential as it is responsible for allowing Spo0A to accumulate in a gradual manner, and this slow accumulation plays a critical role in the ability of the regulatory protein to trigger sporulation.[2] Experiments have also been carried out and results have shown that the activated form of Spo0A failed to trigger sporulation during growth as it had bypassed phosphorylation, and only a small proportion of cells progressed to the early stage of sporulation.[2]
Sporulation
From our research, we found that phosphatases may be viewed as negative regulators that provide access for negative signals to influence the cell’s decision whether to sporulate or to continue vegetative growth.[3] The phosphotases that dephosphorylates Spo0A, preventing its activation are Spo0E, YisI, and YnzD.[4] Therefore, in order to ensure that sporulation occurs under the appropriate conditions, the phosphorelay must integrate the competition between signal input provided by the kinases and signal cancellation carried out by the phosphatises, which determines the decision to sporulate or not,[3][4] by governing flux through the relay and hence the level of Spo0A~P, which must reach a threshold concentration to trigger sporulation.[2]
Other than the external and internal signals that are integrated into the phosphorelay, it is important to note that sporulation is also initiated by nutrient starvation, cell density, and cell cycle progression.[4]
KinA
Sporulation can be triggered with high efficiency in cells in the exponential phase of growth in rich medium by artificial induction of the synthesis of any one of three histidine kinases tha feed phosphoryl groups into the relay.[2] For our project, we will be using Kin A, a major histidine kinase responsible for activating the sporulation pathway in Bacillus subtilis.[5]
KinA is a soluble cytoplasmic protein that appears to be active as a dimer and is composed of an amino-terminal sensor domain and a carboxy-terminal autokinase domain.[5]
Novelty in this sub-project
Instead of allowing the cell to decide whether or not to sporulate, we hope to influence it's decision. We plan to use kinA as a part of this system.
Modelling
KinA Expression Model
Under normal conditions, LacI represses KinA.
However, in the presence of IPTG, KinA can be expressed, as IPTG binds to LacI, deactivating it. Equations (a) and (b) describes how IPTG binds to LacI, forming LacI*, which is the deactivated form of LacI
Over time, the deactivated form of LacI, LacI* may degrade. This is also taken into consideration while building the model.
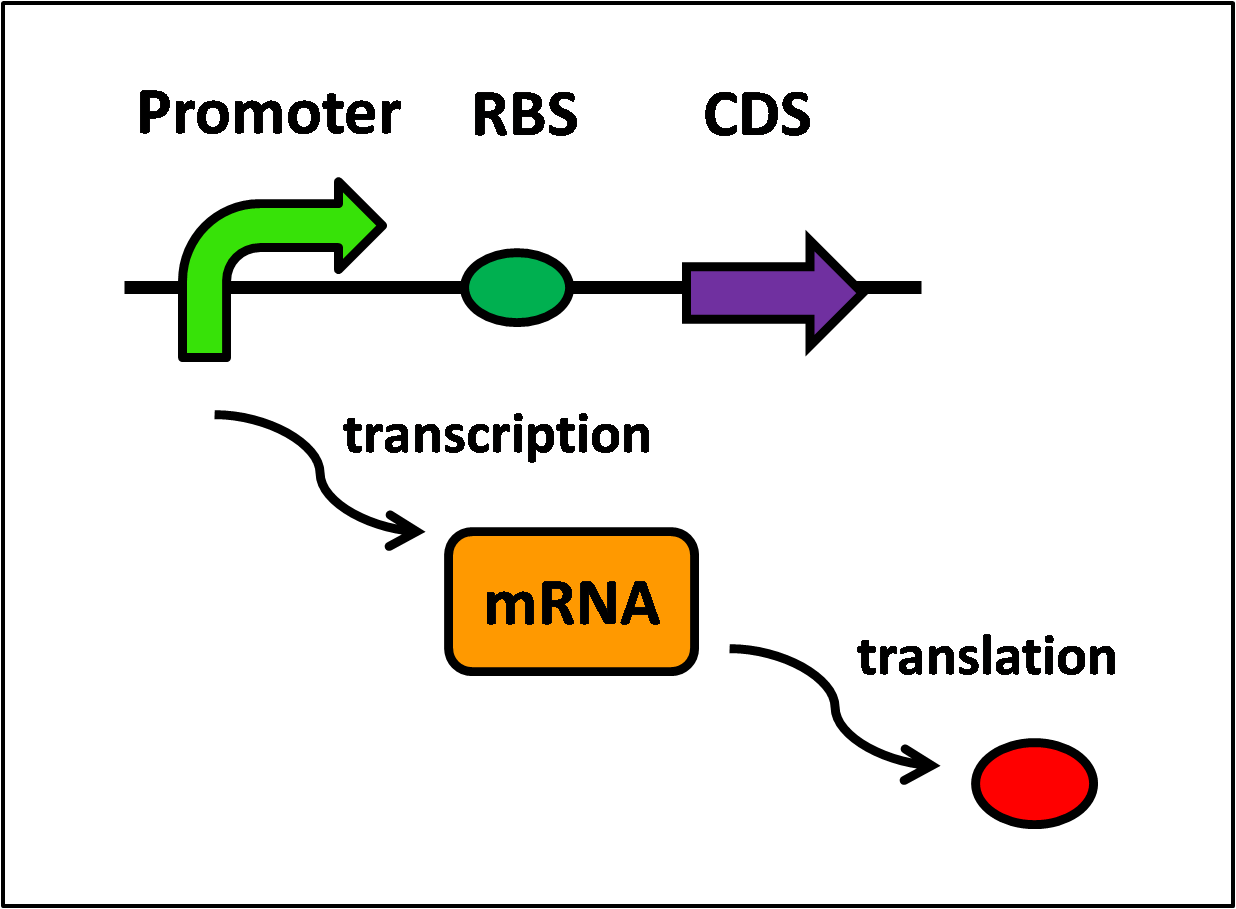
Protein synthesis requires two steps, transcription and translation, and is further illustrated in Figure 1, below.
Referring to Figure 1, the following equations can be written:
- Transcription of LacI,
- Translation of LacI,
where mRNA_LacI is inducing the formation of LacI.
- Transcription of KinA,
where LacI is repressing mRNA_KinA, therefore a lower concentration of LacI would result in a higher concentration of mRNA_KinA.
- Translation of KinA,
where mRNA_KinA is inducing the formation of KinA.
Results
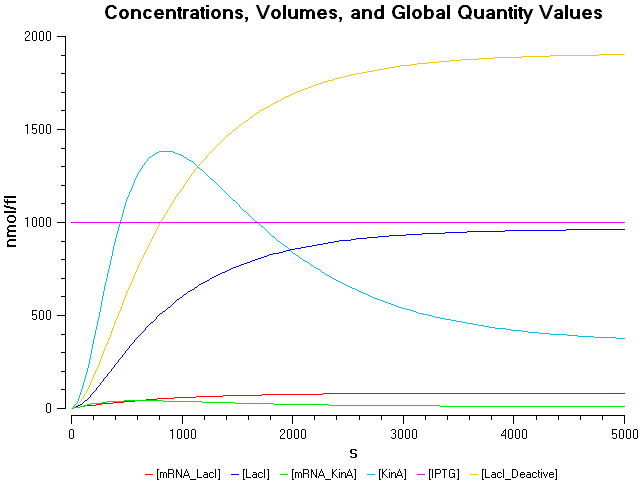
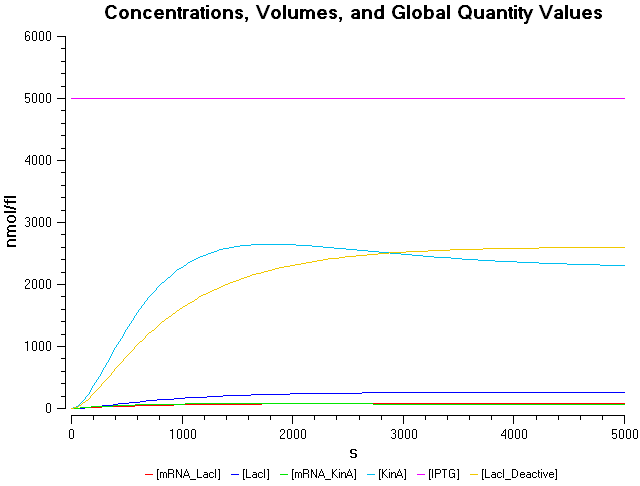
The above equations were modelled in COPASI, and the following graphs show the behaviour of the system over time at different IPTG concentrations.
- IPTG concentration = 1000 nmol/fl
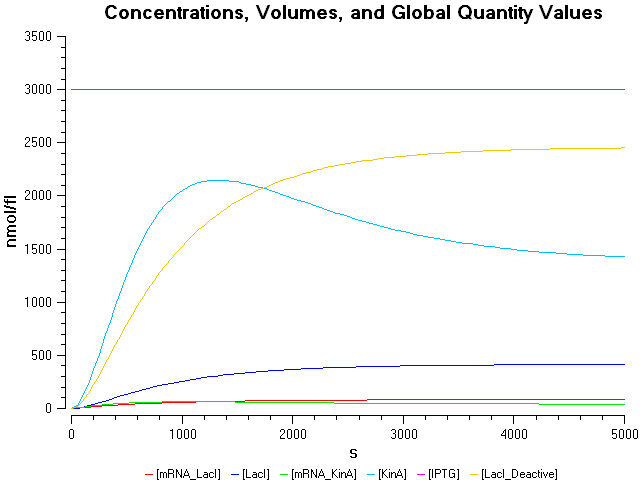
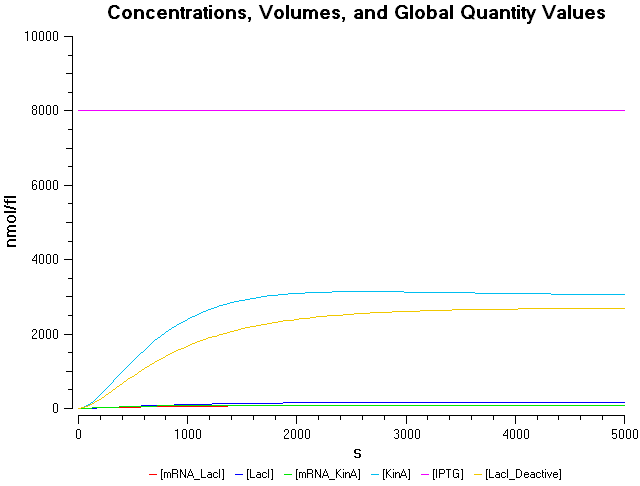
- IPTG concentration = 3000 nmol/fl
- IPTG concentration = 5000 nmol/fl
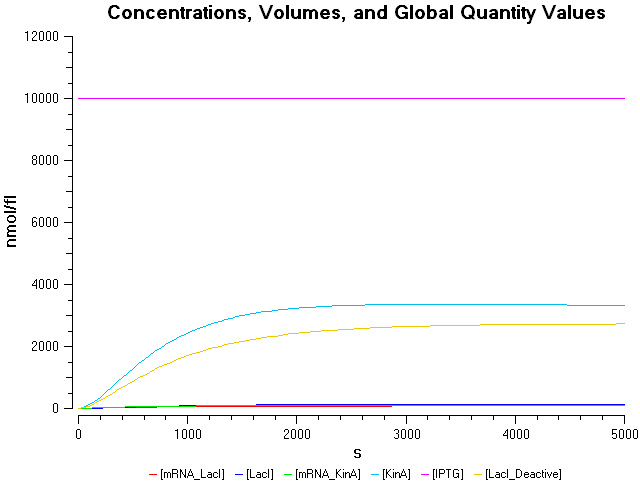
- IPTG concentration = 8000 nmol/fl
- IPTG concentration = 10000 nmol/fl
Sporulation Tuning Model
The expression of KinA has been modelled as seen above, therefore we can now proceed further into the Sporulation Tuning Model, which is built from the KinA Expression Model with COPASI.
To proceed with the modelling of our Sporulation Tuning Model, we have decided that in response to an unidentified stimuli, where KinA autophosphorylates and then donates its phosphate groups to the response regulator Spo0F, the unidentified stimuli will be termed as 'sporulation signal'.[4]
The following equations describe the model:
As Spo0F lacks an output domain and is incapable of activating transcription; it serves only as an intermediary in the phosphorelay. The phosphotransferase Spo0B transfers the phosphate from Spo0F~P to Spo0A.[4]
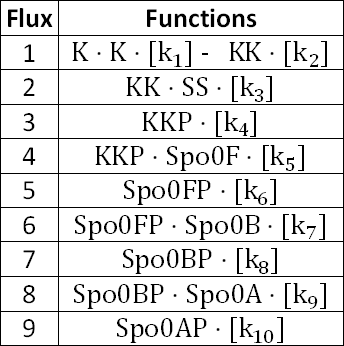
With respect to Equations 1 to 9, as seen above, the corresponding fluxes are as follows:
As mentioned, the phosphotases Spo0E, YisI, and YznD dephosphorylates Spo0A, thus prevents its activation.[4]
Sin (sporulation inhibition) Operon Model
where:
AI, the expression rate of SinI = 0.8 sec-1
γI, the degradation rate of SinI = 0.02 sec-1,
kIon, the rate of the formation of dimers = 0.083 nM-1 sec-1,
kIoff, the off rate for SinI:SinR = 0.5 sec-1,
where:
AR, the expression rate of SinR = 0.014 sec-1
γR, the degradation rate of SinR = 0.002 sec-1
kRoff, the off rate for SinR tetramers = 0.5 sec-1
kRon, the rate for tetramer formation = 0.00125 nM-3 sec-1
where:
k1, the transcription rate from P1 = 0.15 sec-1
γ1, the degradation rate of m1 = 0.005 sec-1
where:
γ3, the degradation rate of m1 = 0.005 sec-1
where:
[S2] is the concentration of Spo0A~P dimers,
[R4] is the concentration of SinR tetramers,
[RNAP] = 30 nM is the concentration of free RNA polymerase available for transcription.
The equilibrium association constants Ki, where RT = 1.62, are defined as:
K2 = exp(-∆G2/RT) x 1e-9, ∆G2 = -10.5
K3 = exp(-∆G3/RT) x 1e-9, ∆G2 = -12.5
K4 = exp(-∆G4/RT) x 1e-9, ∆G2 = -9.0
K5 = exp(-∆G5/RT) x 1e-9, ∆G2 = -21.5
K6 = exp(-∆G6/RT) x 1e-9, ∆G2 = -21.5
K7 = exp(-∆G7/RT) x 1e-9, ∆G2 = -22.5
K8 = exp(-∆G8/RT) x 1e-9, ∆G2 = -33.5
BioBrick constructs
The BioBrick we have designed is to contain an IPTG inducable kinA gene, using pSpac, allowing us to test the theory about KinA in the lab.
BBa_K174010
KinA
Length: 1818bp
Click [http://partsregistry.org/wiki/index.php?title=Part:BBa_K174010 here] for more information on this part.
BBa_K174011
IPTG inducable KinA sporulation trigger
Length: 1953bp
Click [http://partsregistry.org/wiki/index.php?title=Part:BBa_K174011 here] for more information on this part.
Lab Work Strategies
The lab work will mainly be to test our BioBrick using IPTG.
Other Presentations and Diagrams
References
[1] Veening, J-W., Smits, W. K., Kuipers, O. P. (2008) Bistability, Epigenetics, and Bet-Hedging in Bacteria. Annu. Rev. Microbiol. 62: 193-210
[2] Fujita, M., Losick, R. (2005) Evidence that Entry into Sporulation in Bacillus subtilis is Governed by a Gradual Increase in the Level and Activity of the Master Regulator Spo0A. 19: 2236–2244
[3] Sonenshein, A.L., Hoch, J.A., Losick, R., (2002) Bacillus subtilis and Its Closest Relatives From Genes to Cells. ASM Press, United States of America. Pp 476–477
[4] Hilbert, D.W., Piggot, P.J., (June 2004) Compartmentalization of Gene Expression during Bacillus subtilis Spore Formation. Microbiology and Molecular Biology Reviews. Vol. 68, No. 2. Pp 234-262
[5] Eswaramoorthy, P., Guo, T., Fujita, M. (2009) In Vivo Domain-Based Functional Analysis of the Major Sporulation Sensor Kinase, KinA, in Bacillus subtilis. Journal of Bacteriology. Pp 5358-5368
News
Events
- 20 – 21 June 2009 - Europe workshop (London)
- 23 – 24 June 2009 - UK iGEM meetup (Edinburgh)
- 23 October Practice Presentation (Newcastle)
- 23 October T-shirts are ready
- 27 October Practice Presentation (Sunderland)
- 27 October Poster is ready
- 30 October – 2 November 2009 - Jamboree (Boston)
Social Net
- Newcastle iGEM Twitter
- [http://www.facebook.com/home.php#/group.php?gid=131709337641 Newcastle on Facebook]
- [http://www.youtube.com/user/newcastle2009igem Newcastle Youtube Channel]
 "
"