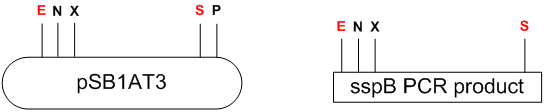Team:Newcastle/ Stochastic Switch cloning strategy
From 2009.igem.org
| Line 27: | Line 27: | ||
[[Image:Sspb into pSB1AT3.png|center|550px]] | [[Image:Sspb into pSB1AT3.png|center|550px]] | ||
| - | [[Image:TeamNewcastleSSpbpSB1AT3.png|center| | + | |
| + | [[Image:TeamNewcastleSSpbpSB1AT3.png|center|250px]] | ||
==Primers== | ==Primers== | ||
| Line 38: | Line 39: | ||
TGTGACACTAGTATTACTTCACAACGCGTAATGC | TGTGACACTAGTATTACTTCACAACGCGTAATGC | ||
| - | + | ||
Revision as of 16:52, 21 October 2009
Contents |
Stochastic Switch Cloning Strategy
We have the following cloning strategies for testing our construct.
Cloning strategies: Stochastic switch
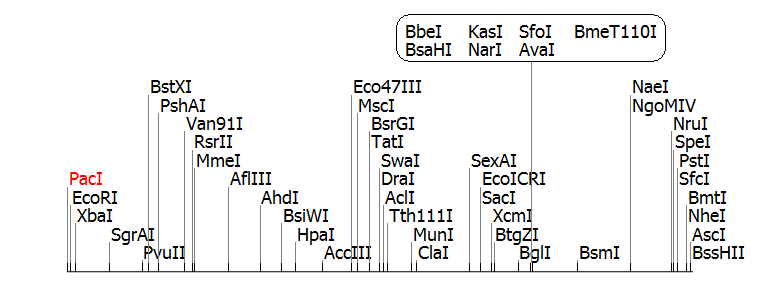
The following is the clone manager file for our synthesised brick; using clone manager it is easy to simulate digests and ligation reactions, in order to make sure the enzymes you use in the will give the result you expect.
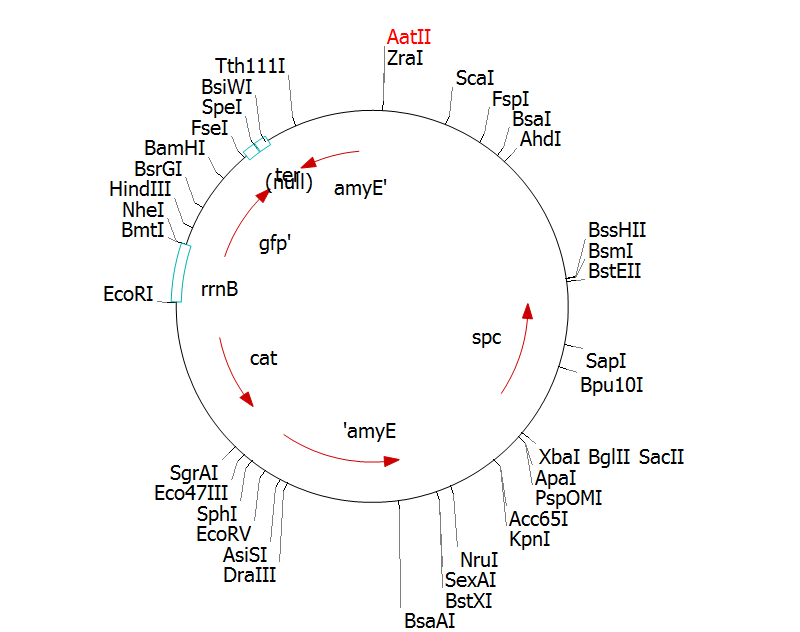
The stochastic brick fragment is to be cloned into the integration vector pGFP-rrnB. This is because plasmid DNA is very unstable in bacillus. From pGFP-rrnb it will integrate into the 168 chromosome at amyE. We can clone the fragment by cutting pGFP-rrnB with EcoRI and NheI, cutting the fragment with EcoRI and NheI, and ligating.
We also need to clone the brick into a BioBrick compatible vector for submission to MIT. Digesting the fragment with EcoR1 and Pst1 and ligation into a EcoR1 and Pst1 cut biobrick vector will do this.
Cloning strategies: Degradation controller
SspB
SspB proteintargets proteins tagged with modified degradation tag, ssrA, and deliver them to ClpXP for proteolysis. Modified ssrA tags are fused onto the 3’ end of a gene. In our project Hin protein is tagged with the modified ssrA. Hence we will be able to control the degradation of Hin proteins by controlling the expression of SspB.The sequence is taken from E. coli strain MG1655. Genbank accession number is NC_000913.2.
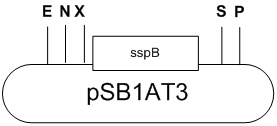
The 498 bp long sspB CDS is amplified by PCR with dangling end primers with EcoRI-NotI-XbaI restriction sites at the 5’ and SpeI at the 3’. The resulting PCR product is cut with EcoRI and SpeI and inserted into a Biobrick compatible vector cut with the same enzymes.
Primers
We used the following primers to amplify SspB:
Forward primer: GATCTGGAATTCGCGGCCGCTTCTAGATGGATTTGTCACAGCTAAC
Reverse Primer: TGTGACACTAGTATTACTTCACAACGCGTAATGC
News
Events
- 20 – 21 June 2009 - Europe workshop (London)
- 23 – 24 June 2009 - UK iGEM meetup (Edinburgh)
- 23 October Practice Presentation (Newcastle)
- 23 October T-shirts are ready
- 27 October Practice Presentation (Sunderland)
- 27 October Poster is ready
- 30 October – 2 November 2009 - Jamboree (Boston)
Social Net
 "
"