Formal Lab Session - 18th September 2009
Overview
Stochastic Switch Team
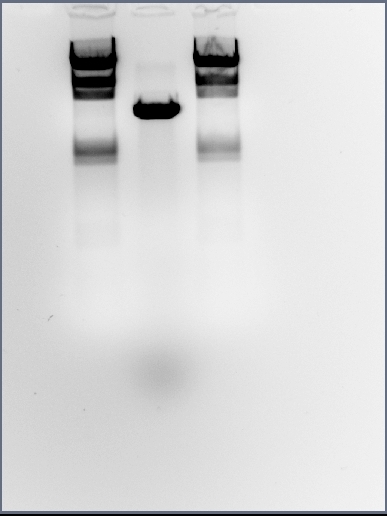
Today we did midipreps of the ara transformation and digested 10ul to run on a gel (EcoRI + PstI) Two bands could be seen one for the backbone and one correctly sied for the insert (~200bp) so this midiprep can be taken forward in order to do the ara-sspb double clone.
Today we also made up some fresh DNA ladder as we have had problems with degradation.
- 40ul of 500ug/ml promega lambda HindIII digest (20ug in total)
- 280ul H2O
- 80ul loading dye
=400ul which was aliquoted into 4 tubes; 3 put in the freezer 1 in the fridge.
Metal Sensor Team
Introduction
Practical Outline
This is the initial list of tasks which need to be carried out by the end of the day:
- Start the Bacillus subtilis transformation process ("Day of transformation" steps of protocol) in the morning.
- Design primers for checking B. subtilis cotC integration
- Plate out some TPA2 cells for production of competent DH5-alpha E. coli cells
- Run digests from yesterday (midi-prep digests) on gel
- If successful transform B. subtilis with midi-prep DNA
- If unsuccessful prepare another ligation attempt
- Carry out a second attempt at digesting the 5 midi-prep samples from the 11/09/09 Lab Session, i.e. cotC-GFP-smtA, kinA, pGFP-rrnB, pMUTIN4 and pSB1AT3
Procedure
Starting Bacillus subtilis transformations
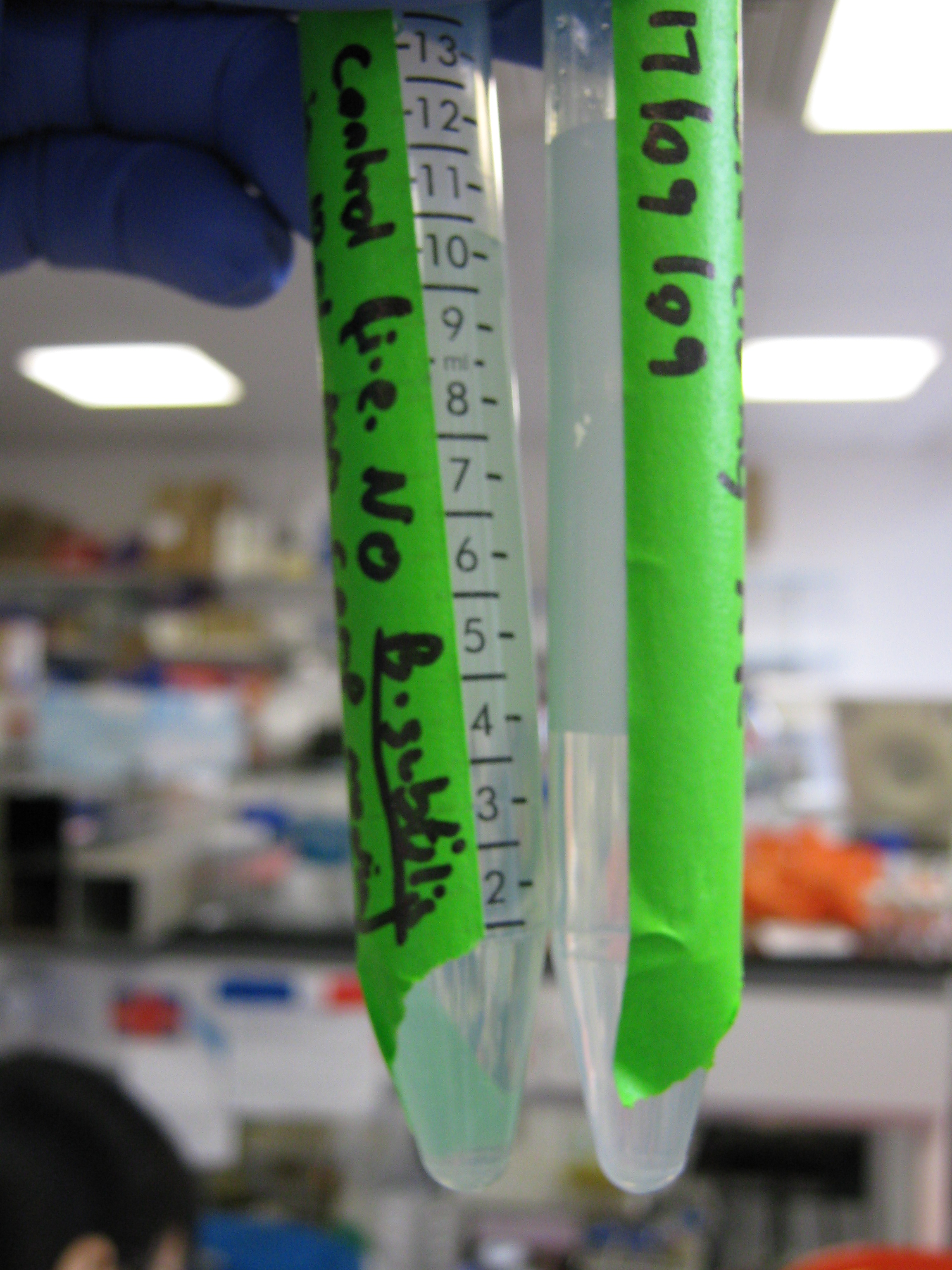
The tube on the right, which contains
B. subtilis cells (grown overnight) is slightly cloudier than the control
The team started the first stage of the Bacillus subtilis transformation process by carrying out the beginning steps of the "Day of transformation" section shown in the protocol. If the midi-prep test digest analysis shows that the ligations and the transformation of cotC-GFP-smtA in the pMUTIN4 plasmid have worked then some of the remaining midi-prep DNA can be used to transform B. subtilis when the time is right.
Designing primers for cotC BioBrick integration
It is key that if the team manage to transform Bacillus subtilis with the cotC-GFP-smtA BioBrick in pMUTIN4 plasmid, they can determine that the BioBrick has successfully integrated into the B. subtilis genome. We have designed the BioBrick so that it integrates within the cotC gene of the B. subtilis genome so any primers designed should flank this region. Unfortunately, due to time restrictions, we didn't manage to carry this step out; it will be carried out another day.
Plating out TPA2 cells
It has been noted that the stock of competent DH5-alpha E. coli cells stored in the -80C freezer has been greatly reduced. To generate some more competent E. coli cells, some TPA2 cells (DH5-alpha E. coli) from the stocks freezer were taken and spread on an LB + agar plate. This is to fulfil the first step of the "Pre-culture" stage in the "Preparing Competent Cells" protocol. The plate will then be grown overnight in the 37C incubator.
Analysing midi-prep digests
The midi-prep digest, which we are analysing today, was carried out in yesterday's lab session. The team hopes that the midi-prep DNA is that of cotC-GFP-smtA ligated into pMUTIN4 plasmid and only an analytical digest will determine this.
Just as a recap: we had attempted to ligate the cotC BioBrick into pMUTIN4 and had attempted to transform DH5-alpha E. coli with this product. Several colonies were found on the LB + chloramphenicol plate the trasnformants had been plated onto which suggested both the ligations and the transformations had worked well. Mini-preps were then carried out for 12 colonies in the 16/09/09 Lab Session and digested. When ran on gel, the 2 bands created did not match the expected sizes but there was consistency for most of the samples. This led us to think that it was the restriction enzymes to blame and ruled out contamination.
Based on this decision, the Metal Sensing team conducted a midi-prep on colony 8 LB culture (seen as colony 8 had the clearest bands) and had digested the DNA (see the 17/09/09 Lab Session). The team then attempted to run 10ul of the digest on agrose gel through DNA gel electrophoresis but the DNA ladder was faulty and the bands seemed faint. Today's midi-prep analysis is a second attempt of this procedure.
It can be seen that the gel photograph shows some erroneous results; there is only one band in sight when there should be two (pMUTIN4 and cotC-GFP-smtA bands) and the band formed lies just underneath the 4,361 band from the HindIII DNA ladder. This size neither matches pMUTIN4 or cotC-GFP-smtA indicating a huge error.
Conclusion
We will have to abandon the potential Bacillus subtilis transformations and consider carrying out a second attempt at ligating together pMUTIN4 and cotC-GFP-smtA.
Analysing cotC, kinA, pGFP-rrnB, pMUTIN4 and pSB1AT3 midi-preps
Today, we only got as far as digesting the five midi-prep samples with restriction enzymes (this should linearise the plasmids) - tomorrow the team will run products on gel. The restriction digests were allowed to run for 1 hour at 37C (in waterbath). The contents of the digest reactions were as follows:
Contents of restriction enzyme digests
| | DNA (ul) | EcoRI (ul) | PstI (ul) | BamHI (ul) | Buffer (ul) | Water (ul)
|
| cotC-GFP-smtA
| 10 | 1 | 1 | 0 | 2 | 6
|
| kinA
| 10 | 1 | 1 | 0 | 2 | 6
|
| pGFP-rrnB
| 10 | 1 | 0 | 0 | 2 | 7
|
| pMUTIN4
| 10 | 0 | 0 | 1 | 2 | 7
|
| pSB1AT3
| 10 | 1 | 1 | 0 | 2 | 6
|
Once the digests were completed, the samples were placed in the -20C freezer for storage.
New Tasks
In light of the ligation/transformation not working, the team needs to start again with the cotC-GFP-smtA work. This involves carrying out a PCR amplification of the cotC BioBrick, digesting both cotC and pMUTIN4, ligating the two fragments together and then using this DNA to transform E. coli cells.
Today, we will do the following:
- Carry out a PCR amplification of the cotC-GFP-smtA BioBrick (which is found within the pMK-RQ plasmid) using primers cotC_forward and cotC_reverse
PCR amplification of cotC-GFP-smtA
Sporulation Tuning/Chassis Team
Introduction
- We cultured two sets of transformation cells yesterday, two Mini prep need to be processed today.
- After Mini Prep, we digested those plasmid to find the strain which contains right clone.
- Continued with yesterday's PCR result, a PCR clean-up process was carried out today.
Experiment procedure
Mini Prep for L1 and L2
- Followed the Phil's Mini Prep protocal.
Digest the Mini Prep result
- Prepare the digest reaction.
dd H2O 7ul
10X fast digest buffer 2ul
Fast enzyme 1 0.5ul
Fast enzyme 2 0.5ul
Mini Prep Plasmid 10ul
------------------------------
20ul
- Incubate 1 hour at 37 degree.
- For digestion of L1 Mini Prep result:
enzyme 1 EcoRI
enzyme 2 SpeI
- For digestion of L2 Mini Prep result:
enzyme 1 XbaI
enzyme 2 SpeI
Control plasmid : pSB1AT3:sleB
Run the digested result on 0.8% agarose gel
- 2ul stop buffer added before loading sample
Conclusion
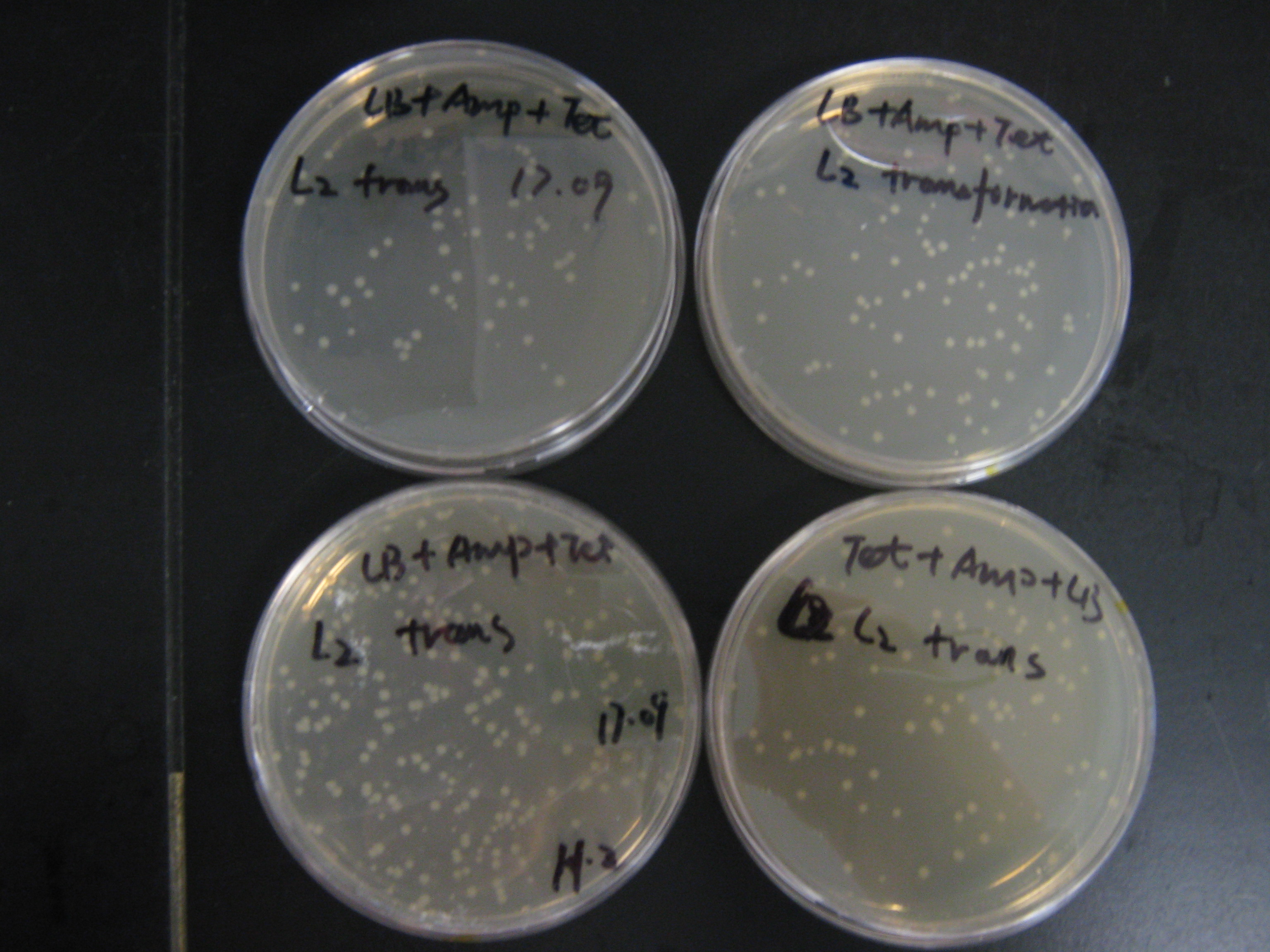
L1: all stains worked well except No.10.
L2: Strain No.4,5,6,11 were not the right clone.
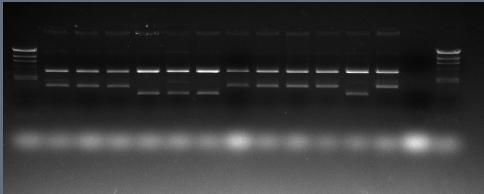
L2: cwlJ and sleB double clone
Further plan
- Culture the right strain for Midi prep of plasmid.
- PCR sleB+cwlJ part and ligate to pMutin4 plasmid.
- Midi Prep L1 and L2 strain.
| July
|
| M | T | W | T | F | S | S
|
|
|
| [http://2009.igem.org/wiki/index.php?title=Team:Newcastle/Labwork/1_July_2009&action=edit 1]
| [http://2009.igem.org/wiki/index.php?title=Team:Newcastle/Labwork/2_July_2009&action=edit 2]
| [http://2009.igem.org/wiki/index.php?title=Team:Newcastle/Labwork/3_July_2009&action=edit 3]
| [http://2009.igem.org/wiki/index.php?title=Team:Newcastle/Labwork/4_July_2009&action=edit 4]
| [http://2009.igem.org/wiki/index.php?title=Team:Newcastle/Labwork/5_July_2009&action=edit 5]
|
| [http://2009.igem.org/wiki/index.php?title=Team:Newcastle/Labwork/6_July_2009&action=edit 6]
| [http://2009.igem.org/wiki/index.php?title=Team:Newcastle/Labwork/7_July_2009&action=edit 7]
| [http://2009.igem.org/wiki/index.php?title=Team:Newcastle/Labwork/8_July_2009&action=edit 8]
| [http://2009.igem.org/wiki/index.php?title=Team:Newcastle/Labwork/9_July_2009&action=edit 9]
| [http://2009.igem.org/wiki/index.php?title=Team:Newcastle/Labwork/10_July_2009&action=edit 10]
| [http://2009.igem.org/wiki/index.php?title=Team:Newcastle/Labwork/11_July_2009&action=edit 11]
| [http://2009.igem.org/wiki/index.php?title=Team:Newcastle/Labwork/12_July_2009&action=edit 12]
|
| [http://2009.igem.org/wiki/index.php?title=Team:Newcastle/Labwork/13_July_2009&action=edit 13]
| [http://2009.igem.org/wiki/index.php?title=Team:Newcastle/Labwork/14_July_2009&action=edit 14]
| [http://2009.igem.org/wiki/index.php?title=Team:Newcastle/Labwork/15_July_2009&action=edit 15]
| [http://2009.igem.org/wiki/index.php?title=Team:Newcastle/Labwork/16_July_2009&action=edit 16]
| [http://2009.igem.org/wiki/index.php?title=Team:Newcastle/Labwork/17_July_2009&action=edit 17]
| [http://2009.igem.org/wiki/index.php?title=Team:Newcastle/Labwork/18_July_2009&action=edit 18]
| [http://2009.igem.org/wiki/index.php?title=Team:Newcastle/Labwork/19_July_2009&action=edit 19]
|
| [http://2009.igem.org/wiki/index.php?title=Team:Newcastle/Labwork/20_July_2009&action=edit 20]
| [http://2009.igem.org/wiki/index.php?title=Team:Newcastle/Labwork/21_July_2009&action=edit 21]
| [http://2009.igem.org/wiki/index.php?title=Team:Newcastle/Labwork/22_July_2009&action=edit 22]
| [http://2009.igem.org/Team:Newcastle/Labwork/23_July_2009 23]
| [http://2009.igem.org/Team:Newcastle/Labwork/24_July_2009 24]
| [http://2009.igem.org/wiki/index.php?title=Team:Newcastle/Labwork/25_July_2009&action=edit 25]
| [http://2009.igem.org/wiki/index.php?title=Team:Newcastle/Labwork/26_July_2009&action=edit 26]
|
| [http://2009.igem.org/Team:Newcastle/Labwork/27_July_2009 27]
| [http://2009.igem.org/Team:Newcastle/Labwork/28_July_2009 28]
| [http://2009.igem.org/Team:Newcastle/Labwork/29_July_2009 29]
| [http://2009.igem.org/Team:Newcastle/Labwork/30_July_2009 30]
| [http://2009.igem.org/Team:Newcastle/Labwork/31_July_2009 31]
|
|
| August
|
| M | T | W | T | F | S | S
|
|
|
|
|
|
| [http://2009.igem.org/wiki/index.php?title=Team:Newcastle/Labwork/1_August_2009&action=edit 1]
| [http://2009.igem.org/wiki/index.php?title=Team:Newcastle/Labwork/2_August_2009&action=edit 2]
|
| [http://2009.igem.org/wiki/index.php?title=Team:Newcastle/Labwork/3_August_2009&action=edit 3]
| [http://2009.igem.org/Team:Newcastle/Labwork/4_August_2009 4]
| [http://2009.igem.org/Team:Newcastle/Labwork/5_August_2009 5]
| [http://2009.igem.org/wiki/index.php?title=Team:Newcastle/Labwork/6_August_2009&action=edit 6]
| [http://2009.igem.org/Team:Newcastle/Labwork/7_August_2009 7]
| [http://2009.igem.org/wiki/index.php?title=Team:Newcastle/Labwork/8_August_2009&action=edit 8]
| [http://2009.igem.org/wiki/index.php?title=Team:Newcastle/Labwork/9_August_2009&action=edit 9]
|
| [http://2009.igem.org/Team:Newcastle/Labwork/10_August_2009 10]
| [http://2009.igem.org/Team:Newcastle/Labwork/11_August_2009 11]
| [http://2009.igem.org/Team:Newcastle/Labwork/12_August_2009 12]
| [http://2009.igem.org/Team:Newcastle/Labwork/13_August_2009 13]
| [http://2009.igem.org/Team:Newcastle/Labwork/14_August_2009 14]
| [http://2009.igem.org/wiki/index.php?title=Team:Newcastle/Labwork/15_August_2009&action=edit 15]
| [http://2009.igem.org/wiki/index.php?title=Team:Newcastle/Labwork/16_August_2009&action=edit 16]
|
| [http://2009.igem.org/Team:Newcastle/Labwork/17_August_2009 17]
| [http://2009.igem.org/Team:Newcastle/Labwork/18_August_2009 18]
| [http://2009.igem.org/Team:Newcastle/Labwork/19_August_2009 19]
| [http://2009.igem.org/Team:Newcastle/Labwork/20_August_2009 20]
| [http://2009.igem.org/Team:Newcastle/Labwork/21_August_2009 21]
| [http://2009.igem.org/wiki/index.php?title=Team:Newcastle/Labwork/22_August_2009&action=edit 22]
| [http://2009.igem.org/wiki/index.php?title=Team:Newcastle/Labwork/23_August_2009&action=edit 23]
|
| [http://2009.igem.org/Team:Newcastle/Labwork/24_August_2009 24]
| [http://2009.igem.org/Team:Newcastle/Labwork/25_August_2009 25]
| [http://2009.igem.org/Team:Newcastle/Labwork/26_August_2009 26]
| [http://2009.igem.org/Team:Newcastle/Labwork/27_August_2009 27]
| [http://2009.igem.org/Team:Newcastle/Labwork/28_August_2009 28]
| [http://2009.igem.org/wiki/index.php?title=Team:Newcastle/Labwork/29_August_2009&action=edit 29]
| [http://2009.igem.org/wiki/index.php?title=Team:Newcastle/Labwork/30_August_2009&action=edit 30]
|
| [http://2009.igem.org/wiki/index.php?title=Team:Newcastle/Labwork/31_August_2009&action=edit 31]
|
|
| September
|
| M | T | W | T | F | S | S
|
|
| [http://2009.igem.org/Team:Newcastle/Labwork/1_September_2009 1]
| [http://2009.igem.org/Team:Newcastle/Labwork/2_September_2009 2]
| [http://2009.igem.org/Team:Newcastle/Labwork/3_September_2009 3]
| [http://2009.igem.org/Team:Newcastle/Labwork/4_September_2009 4]
| [http://2009.igem.org/wiki/index.php?title=Team:Newcastle/Labwork/5_September_2009&action=edit 5]
| [http://2009.igem.org/wiki/index.php?title=Team:Newcastle/Labwork/6_September_2009&action=edit 6]
|
| [http://2009.igem.org/Team:Newcastle/Labwork/7_September_2009 7]
| [http://2009.igem.org/Team:Newcastle/Labwork/8_September_2009 8]
| [http://2009.igem.org/Team:Newcastle/Labwork/9_September_2009 9]
| [http://2009.igem.org/Team:Newcastle/Labwork/10_September_2009 10]
| [http://2009.igem.org/Team:Newcastle/Labwork/11_September_2009 11]
| [http://2009.igem.org/wiki/index.php?title=Team:Newcastle/Labwork/12_September_2009&action=edit 12]
| [http://2009.igem.org/wiki/index.php?title=Team:Newcastle/Labwork/13_September_2009&action=edit 13]
|
| [http://2009.igem.org/Team:Newcastle/Labwork/14_September_2009 14]
| [http://2009.igem.org/Team:Newcastle/Labwork/15_September_2009 15]
| [http://2009.igem.org/Team:Newcastle/Labwork/16_September_2009 16]
| [http://2009.igem.org/Team:Newcastle/Labwork/17_September_2009 17]
| [http://2009.igem.org/Team:Newcastle/Labwork/18_September_2009 18]
| [http://2009.igem.org/wiki/index.php?title=Team:Newcastle/Labwork/19_September_2009&action=edit 19]
| [http://2009.igem.org/wiki/index.php?title=Team:Newcastle/Labwork/20_September_2009&action=edit 20]
|
| [http://2009.igem.org/Team:Newcastle/Labwork/21_September_2009 21]
| [http://2009.igem.org/Team:Newcastle/Labwork/22_September_2009 22]
| [http://2009.igem.org/Team:Newcastle/Labwork/23_September_2009 23]
| [http://2009.igem.org/Team:Newcastle/Labwork/24_September_2009 24]
| [http://2009.igem.org/Team:Newcastle/Labwork/25_September_2009 25]
| [http://2009.igem.org/wiki/index.php?title=Team:Newcastle/Labwork/26_September_2009&action=edit 26]
| [http://2009.igem.org/wiki/index.php?title=Team:Newcastle/Labwork/27_September_2009&action=edit 27]
|
| [http://2009.igem.org/Team:Newcastle/Labwork/28_September_2009 28]
| [http://2009.igem.org/Team:Newcastle/Labwork/29_September_2009 29]
| [http://2009.igem.org/Team:Newcastle/Labwork/30_September_2009 30]
|
|
| October
|
| M | T | W | T | F | S | S
|
|
|
|
| [http://2009.igem.org/Team:Newcastle/Labwork/1_October_2009 1]
| [http://2009.igem.org/Team:Newcastle/Labwork/2_October_2009 2]
| [http://2009.igem.org/wiki/index.php?title=Team:Newcastle/Labwork/3_October_2009&action=edit 3]
| [http://2009.igem.org/wiki/index.php?title=Team:Newcastle/Labwork/4_October_2009&action=edit 4]
|
| [http://2009.igem.org/Team:Newcastle/Labwork/5_October_2009 5]
| [http://2009.igem.org/Team:Newcastle/Labwork/6_October_2009 6]
| [http://2009.igem.org/Team:Newcastle/Labwork/7_October_2009 7]
| [http://2009.igem.org/Team:Newcastle/Labwork/8_October_2009 8]
| [http://2009.igem.org/Team:Newcastle/Labwork/9_October_2009 9]
| [http://2009.igem.org/wiki/index.php?title=Team:Newcastle/Labwork/10_October_2009&action=edit 10]
| [http://2009.igem.org/wiki/index.php?title=Team:Newcastle/Labwork/11_October_2009&action=edit 11]
|
| [http://2009.igem.org/Team:Newcastle/Labwork/12_October_2009 12]
| [http://2009.igem.org/Team:Newcastle/Labwork/13_October_2009 13]
| [http://2009.igem.org/Team:Newcastle/Labwork/14_October_2009 14]
| [http://2009.igem.org/Team:Newcastle/Labwork/15_October_2009 15]
| [http://2009.igem.org/Team:Newcastle/Labwork/16_October_2009 16]
| [http://2009.igem.org/wiki/index.php?title=Team:Newcastle/Labwork/17_October_2009&action=edit 17]
| [http://2009.igem.org/wiki/index.php?title=Team:Newcastle/Labwork/18_October_2009&action=edit 18]
|
| [http://2009.igem.org/wiki/index.php?title=Team:Newcastle/Labwork/19_October_2009&action=edit 19]
| [http://2009.igem.org/wiki/index.php?title=Team:Newcastle/Labwork/20_October_2009&action=edit 20]
| [http://2009.igem.org/wiki/index.php?title=Team:Newcastle/Labwork/21_October_2009&action=edit 21]
| [http://2009.igem.org/wiki/index.php?title=Team:Newcastle/Labwork/22_October_2009&action=edit 22]
| [http://2009.igem.org/wiki/index.php?title=Team:Newcastle/Labwork/23_October_2009&action=edit 23]
| [http://2009.igem.org/wiki/index.php?title=Team:Newcastle/Labwork/24_October_2009&action=edit 24]
| [http://2009.igem.org/wiki/index.php?title=Team:Newcastle/Labwork/25_October_2009&action=edit 25]
|
| [http://2009.igem.org/wiki/index.php?title=Team:Newcastle/Labwork/26_October_2009&action=edit 26]
| [http://2009.igem.org/wiki/index.php?title=Team:Newcastle/Labwork/27_October_2009&action=edit 27]
| [http://2009.igem.org/wiki/index.php?title=Team:Newcastle/Labwork/28_October_2009&action=edit 28]
| [http://2009.igem.org/wiki/index.php?title=Team:Newcastle/Labwork/29_October_2009&action=edit 29]
| [http://2009.igem.org/wiki/index.php?title=Team:Newcastle/Labwork/30_October_2009&action=edit 30]
| [http://2009.igem.org/wiki/index.php?title=Team:Newcastle/Labwork/31_October_2009&action=edit 31]
|
|
 "
"