Formal Lab Session - 28th July 2009
Introduction
In the past few lab sessions the team has transformed JM109 E. coli cells with five BioBricks from the Spring Distribution and attempted to grow the transformant cells on LB + amp plates. The five BioBricks include BBa_C0056 (cI in the lambda phage), BBa_B1002 (terminator sequence), BBa_C0077 (CinR activator), BBa_C0076 (autoinducer synthetase) and BBa_R0077 (promoter which is CinR and HSL regulated - RBS+).
It was found that when the five sets of transformant E. coli cells were plated out, the cells possessing BBa_C0056, BBa_B1002, BBa_C0076 and BBa_R0077 seemed to produce colonies (although it must be noted that there was a little uncertainty about whether the E. coli transformed with BBa_C0076 did actually produce colonies). Three colonies from each of the four transformant plates were taken and each used to inoculate tubes containing 5ml LB solution (making a total of 12 tubes). These tubes were then placed in the orbital incubator overnight for mini-preps.
Today's plan is to conduct mini-preps on the four sets of E. coli transformant cells
Practical Outline

Dan getting reacquainted with lab techniques
Here's the objectives for today's lab session:
- Carry out mini-preps of the E. coli cells which have taken up the BioBricks BBa_C0056, BBa_B1002, BBa_C0076 and BBa_R0077
- Run these mini-prep samples on the gel to check for DNA
Observations and Hypothesis
When the 12 tubes (each with 5ml LB plus E. coli cells) were taken out of the orbital incubator it was seen that the tubes possessing JM109 cells + BBa_C0076 were clear compared to the other 'cloudy' tubes. This means that the transformant E.coli in the other tubes grew successfully but there was no E. coli growth in the tube where BBa_C0076 transformants should have been.
This means that the so-called colonies found on the LB + amp plate containing the E. coli + BBa_C0076 were not colonies at all! This means that the attempts to transform both BBa_C0077 and BBa_C0076 have failed.
After analysis of the situation, the team studied the plasmid in which the two BioBricks are found. It turns out that both BioBricks are carried by the same plasmid backbone - pSB2K3 - and that this plasmid DOES NOT CONFER AMPICILLIN RESISTANCE! The resistance it bears is kanamycin resistance - this is why the attempted transformations have not grown colonies on the plates. This error has slowed down progress but it has also taught the team some valuable lessons in organisation and lab planning.
Procedure
Mini-Preps for BBa_C0056, BBa_B1002 and BBa_R0077 transformants
The mini-preps carried out on the three sets of transformed JM109 cells was done using Phil's Mini Method protocol. There weren't any particular changes to the protocol but a suggestion was made to use the aspirator/vacuum pump when removing the supernatant in step 2 (where Solution I + RNAse is added) - this method ensures rapid removal of supernatant and also saves on waste flasks.
When using the Speed Vac machine in step 9, the Speed Vac protocol was observed to ensure the machine doesn't get damaged by incorrect handling.
Once the mini-prep procedure was completed the plasmid samples were stored in the -20C freezer for later use.
DNA Gel Electrophoresis
To show that we have DNA in our samples and that the DNA is not anything but our desired BioBricks (within their correct plasmids) we carried out DNA gel electrophoresis on our plasmids. Although we loaded the samples as the electrophoresis protocol suggests there was a change made to the preparation of the DNA samples; each of the nine samples consisted of:
- 10ul of distilled water
- 5ul of plasmid DNA (along with BioBrick)
- 2ul of loading buffer/dye
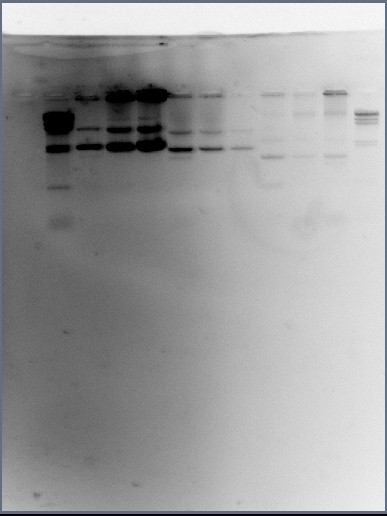
The DNA gel electrophoresis was allowed to commence for 40 minutes under 100volts. Once this time had elapsed a photograph was taken of the gel under UV light using the GelDoc system. This is the photograph:
Not a lot can be determined from this photograph because none of the mini-prep samples have been digested by any restriction enzymes yet. However one thing to note is the consistency in the three sets of wells - all of the BBa_C0056 mini-prep samples (samples 1-3) look similar in layout, the BBa_B1002 mini-prep samples (samples 4-6) also look similar and the BBa_R0077 DNA samples (samples 7-9) look very similar. Although this gel photograph is not enough to make conclusions as to whether the plasmids are actually there, there is enough evidence to suggest that there is no contamination and that DNA is certainly present.
Conclusion
Carry out restriction enzyme digests of the 9 mini-prepped DNA to ensure that the correct DNA has been transformed - the enzymes to be used are EcoRI and PstI.
| July
|
| M | T | W | T | F | S | S
|
|
|
| [http://2009.igem.org/wiki/index.php?title=Team:Newcastle/Labwork/1_July_2009&action=edit 1]
| [http://2009.igem.org/wiki/index.php?title=Team:Newcastle/Labwork/2_July_2009&action=edit 2]
| [http://2009.igem.org/wiki/index.php?title=Team:Newcastle/Labwork/3_July_2009&action=edit 3]
| [http://2009.igem.org/wiki/index.php?title=Team:Newcastle/Labwork/4_July_2009&action=edit 4]
| [http://2009.igem.org/wiki/index.php?title=Team:Newcastle/Labwork/5_July_2009&action=edit 5]
|
| [http://2009.igem.org/wiki/index.php?title=Team:Newcastle/Labwork/6_July_2009&action=edit 6]
| [http://2009.igem.org/wiki/index.php?title=Team:Newcastle/Labwork/7_July_2009&action=edit 7]
| [http://2009.igem.org/wiki/index.php?title=Team:Newcastle/Labwork/8_July_2009&action=edit 8]
| [http://2009.igem.org/wiki/index.php?title=Team:Newcastle/Labwork/9_July_2009&action=edit 9]
| [http://2009.igem.org/wiki/index.php?title=Team:Newcastle/Labwork/10_July_2009&action=edit 10]
| [http://2009.igem.org/wiki/index.php?title=Team:Newcastle/Labwork/11_July_2009&action=edit 11]
| [http://2009.igem.org/wiki/index.php?title=Team:Newcastle/Labwork/12_July_2009&action=edit 12]
|
| [http://2009.igem.org/wiki/index.php?title=Team:Newcastle/Labwork/13_July_2009&action=edit 13]
| [http://2009.igem.org/wiki/index.php?title=Team:Newcastle/Labwork/14_July_2009&action=edit 14]
| [http://2009.igem.org/wiki/index.php?title=Team:Newcastle/Labwork/15_July_2009&action=edit 15]
| [http://2009.igem.org/wiki/index.php?title=Team:Newcastle/Labwork/16_July_2009&action=edit 16]
| [http://2009.igem.org/wiki/index.php?title=Team:Newcastle/Labwork/17_July_2009&action=edit 17]
| [http://2009.igem.org/wiki/index.php?title=Team:Newcastle/Labwork/18_July_2009&action=edit 18]
| [http://2009.igem.org/wiki/index.php?title=Team:Newcastle/Labwork/19_July_2009&action=edit 19]
|
| [http://2009.igem.org/wiki/index.php?title=Team:Newcastle/Labwork/20_July_2009&action=edit 20]
| [http://2009.igem.org/wiki/index.php?title=Team:Newcastle/Labwork/21_July_2009&action=edit 21]
| [http://2009.igem.org/wiki/index.php?title=Team:Newcastle/Labwork/22_July_2009&action=edit 22]
| [http://2009.igem.org/Team:Newcastle/Labwork/23_July_2009 23]
| [http://2009.igem.org/Team:Newcastle/Labwork/24_July_2009 24]
| [http://2009.igem.org/wiki/index.php?title=Team:Newcastle/Labwork/25_July_2009&action=edit 25]
| [http://2009.igem.org/wiki/index.php?title=Team:Newcastle/Labwork/26_July_2009&action=edit 26]
|
| [http://2009.igem.org/Team:Newcastle/Labwork/27_July_2009 27]
| [http://2009.igem.org/Team:Newcastle/Labwork/28_July_2009 28]
| [http://2009.igem.org/Team:Newcastle/Labwork/29_July_2009 29]
| [http://2009.igem.org/Team:Newcastle/Labwork/30_July_2009 30]
| [http://2009.igem.org/Team:Newcastle/Labwork/31_July_2009 31]
|
|
| August
|
| M | T | W | T | F | S | S
|
|
|
|
|
|
| [http://2009.igem.org/wiki/index.php?title=Team:Newcastle/Labwork/1_August_2009&action=edit 1]
| [http://2009.igem.org/wiki/index.php?title=Team:Newcastle/Labwork/2_August_2009&action=edit 2]
|
| [http://2009.igem.org/wiki/index.php?title=Team:Newcastle/Labwork/3_August_2009&action=edit 3]
| [http://2009.igem.org/Team:Newcastle/Labwork/4_August_2009 4]
| [http://2009.igem.org/Team:Newcastle/Labwork/5_August_2009 5]
| [http://2009.igem.org/wiki/index.php?title=Team:Newcastle/Labwork/6_August_2009&action=edit 6]
| [http://2009.igem.org/Team:Newcastle/Labwork/7_August_2009 7]
| [http://2009.igem.org/wiki/index.php?title=Team:Newcastle/Labwork/8_August_2009&action=edit 8]
| [http://2009.igem.org/wiki/index.php?title=Team:Newcastle/Labwork/9_August_2009&action=edit 9]
|
| [http://2009.igem.org/Team:Newcastle/Labwork/10_August_2009 10]
| [http://2009.igem.org/Team:Newcastle/Labwork/11_August_2009 11]
| [http://2009.igem.org/Team:Newcastle/Labwork/12_August_2009 12]
| [http://2009.igem.org/Team:Newcastle/Labwork/13_August_2009 13]
| [http://2009.igem.org/Team:Newcastle/Labwork/14_August_2009 14]
| [http://2009.igem.org/wiki/index.php?title=Team:Newcastle/Labwork/15_August_2009&action=edit 15]
| [http://2009.igem.org/wiki/index.php?title=Team:Newcastle/Labwork/16_August_2009&action=edit 16]
|
| [http://2009.igem.org/Team:Newcastle/Labwork/17_August_2009 17]
| [http://2009.igem.org/Team:Newcastle/Labwork/18_August_2009 18]
| [http://2009.igem.org/Team:Newcastle/Labwork/19_August_2009 19]
| [http://2009.igem.org/Team:Newcastle/Labwork/20_August_2009 20]
| [http://2009.igem.org/Team:Newcastle/Labwork/21_August_2009 21]
| [http://2009.igem.org/wiki/index.php?title=Team:Newcastle/Labwork/22_August_2009&action=edit 22]
| [http://2009.igem.org/wiki/index.php?title=Team:Newcastle/Labwork/23_August_2009&action=edit 23]
|
| [http://2009.igem.org/Team:Newcastle/Labwork/24_August_2009 24]
| [http://2009.igem.org/Team:Newcastle/Labwork/25_August_2009 25]
| [http://2009.igem.org/Team:Newcastle/Labwork/26_August_2009 26]
| [http://2009.igem.org/Team:Newcastle/Labwork/27_August_2009 27]
| [http://2009.igem.org/Team:Newcastle/Labwork/28_August_2009 28]
| [http://2009.igem.org/wiki/index.php?title=Team:Newcastle/Labwork/29_August_2009&action=edit 29]
| [http://2009.igem.org/wiki/index.php?title=Team:Newcastle/Labwork/30_August_2009&action=edit 30]
|
| [http://2009.igem.org/wiki/index.php?title=Team:Newcastle/Labwork/31_August_2009&action=edit 31]
|
|
| September
|
| M | T | W | T | F | S | S
|
|
| [http://2009.igem.org/Team:Newcastle/Labwork/1_September_2009 1]
| [http://2009.igem.org/Team:Newcastle/Labwork/2_September_2009 2]
| [http://2009.igem.org/Team:Newcastle/Labwork/3_September_2009 3]
| [http://2009.igem.org/Team:Newcastle/Labwork/4_September_2009 4]
| [http://2009.igem.org/wiki/index.php?title=Team:Newcastle/Labwork/5_September_2009&action=edit 5]
| [http://2009.igem.org/wiki/index.php?title=Team:Newcastle/Labwork/6_September_2009&action=edit 6]
|
| [http://2009.igem.org/Team:Newcastle/Labwork/7_September_2009 7]
| [http://2009.igem.org/Team:Newcastle/Labwork/8_September_2009 8]
| [http://2009.igem.org/Team:Newcastle/Labwork/9_September_2009 9]
| [http://2009.igem.org/Team:Newcastle/Labwork/10_September_2009 10]
| [http://2009.igem.org/Team:Newcastle/Labwork/11_September_2009 11]
| [http://2009.igem.org/wiki/index.php?title=Team:Newcastle/Labwork/12_September_2009&action=edit 12]
| [http://2009.igem.org/wiki/index.php?title=Team:Newcastle/Labwork/13_September_2009&action=edit 13]
|
| [http://2009.igem.org/Team:Newcastle/Labwork/14_September_2009 14]
| [http://2009.igem.org/Team:Newcastle/Labwork/15_September_2009 15]
| [http://2009.igem.org/Team:Newcastle/Labwork/16_September_2009 16]
| [http://2009.igem.org/Team:Newcastle/Labwork/17_September_2009 17]
| [http://2009.igem.org/Team:Newcastle/Labwork/18_September_2009 18]
| [http://2009.igem.org/wiki/index.php?title=Team:Newcastle/Labwork/19_September_2009&action=edit 19]
| [http://2009.igem.org/wiki/index.php?title=Team:Newcastle/Labwork/20_September_2009&action=edit 20]
|
| [http://2009.igem.org/Team:Newcastle/Labwork/21_September_2009 21]
| [http://2009.igem.org/Team:Newcastle/Labwork/22_September_2009 22]
| [http://2009.igem.org/Team:Newcastle/Labwork/23_September_2009 23]
| [http://2009.igem.org/Team:Newcastle/Labwork/24_September_2009 24]
| [http://2009.igem.org/Team:Newcastle/Labwork/25_September_2009 25]
| [http://2009.igem.org/wiki/index.php?title=Team:Newcastle/Labwork/26_September_2009&action=edit 26]
| [http://2009.igem.org/wiki/index.php?title=Team:Newcastle/Labwork/27_September_2009&action=edit 27]
|
| [http://2009.igem.org/Team:Newcastle/Labwork/28_September_2009 28]
| [http://2009.igem.org/Team:Newcastle/Labwork/29_September_2009 29]
| [http://2009.igem.org/Team:Newcastle/Labwork/30_September_2009 30]
|
|
| October
|
| M | T | W | T | F | S | S
|
|
|
|
| [http://2009.igem.org/Team:Newcastle/Labwork/1_October_2009 1]
| [http://2009.igem.org/Team:Newcastle/Labwork/2_October_2009 2]
| [http://2009.igem.org/wiki/index.php?title=Team:Newcastle/Labwork/3_October_2009&action=edit 3]
| [http://2009.igem.org/wiki/index.php?title=Team:Newcastle/Labwork/4_October_2009&action=edit 4]
|
| [http://2009.igem.org/Team:Newcastle/Labwork/5_October_2009 5]
| [http://2009.igem.org/Team:Newcastle/Labwork/6_October_2009 6]
| [http://2009.igem.org/Team:Newcastle/Labwork/7_October_2009 7]
| [http://2009.igem.org/Team:Newcastle/Labwork/8_October_2009 8]
| [http://2009.igem.org/Team:Newcastle/Labwork/9_October_2009 9]
| [http://2009.igem.org/wiki/index.php?title=Team:Newcastle/Labwork/10_October_2009&action=edit 10]
| [http://2009.igem.org/wiki/index.php?title=Team:Newcastle/Labwork/11_October_2009&action=edit 11]
|
| [http://2009.igem.org/Team:Newcastle/Labwork/12_October_2009 12]
| [http://2009.igem.org/Team:Newcastle/Labwork/13_October_2009 13]
| [http://2009.igem.org/Team:Newcastle/Labwork/14_October_2009 14]
| [http://2009.igem.org/Team:Newcastle/Labwork/15_October_2009 15]
| [http://2009.igem.org/Team:Newcastle/Labwork/16_October_2009 16]
| [http://2009.igem.org/wiki/index.php?title=Team:Newcastle/Labwork/17_October_2009&action=edit 17]
| [http://2009.igem.org/wiki/index.php?title=Team:Newcastle/Labwork/18_October_2009&action=edit 18]
|
| [http://2009.igem.org/wiki/index.php?title=Team:Newcastle/Labwork/19_October_2009&action=edit 19]
| [http://2009.igem.org/wiki/index.php?title=Team:Newcastle/Labwork/20_October_2009&action=edit 20]
| [http://2009.igem.org/wiki/index.php?title=Team:Newcastle/Labwork/21_October_2009&action=edit 21]
| [http://2009.igem.org/wiki/index.php?title=Team:Newcastle/Labwork/22_October_2009&action=edit 22]
| [http://2009.igem.org/wiki/index.php?title=Team:Newcastle/Labwork/23_October_2009&action=edit 23]
| [http://2009.igem.org/wiki/index.php?title=Team:Newcastle/Labwork/24_October_2009&action=edit 24]
| [http://2009.igem.org/wiki/index.php?title=Team:Newcastle/Labwork/25_October_2009&action=edit 25]
|
| [http://2009.igem.org/wiki/index.php?title=Team:Newcastle/Labwork/26_October_2009&action=edit 26]
| [http://2009.igem.org/wiki/index.php?title=Team:Newcastle/Labwork/27_October_2009&action=edit 27]
| [http://2009.igem.org/wiki/index.php?title=Team:Newcastle/Labwork/28_October_2009&action=edit 28]
| [http://2009.igem.org/wiki/index.php?title=Team:Newcastle/Labwork/29_October_2009&action=edit 29]
| [http://2009.igem.org/wiki/index.php?title=Team:Newcastle/Labwork/30_October_2009&action=edit 30]
| [http://2009.igem.org/wiki/index.php?title=Team:Newcastle/Labwork/31_October_2009&action=edit 31]
|
|
 "
"