Modeling > Parameters
Parameters
Constructing ODEs is only the first step of simulating our design. The parameters, actually, play a significant role in the modeling process. Here are two sets of parameters(T3 RNA polymerase and P2) we used.
We have done an systematic literature review to select parameters for our model. However, not every parameter can be found from existing data, which means we have to guess part of the parameters from trial and error. To check whether we have guessed correctly, we do the sensitivity test. The sensitivity test works like this: We first give both salicylate(food) and arabinose(bell) to make bistable turn to CI state(have memory). After a period of time, we give the E. coli arabinose stimulus and GFP output will raise after a short while. We select the highest concentration point of GFP in the second procedure. The sensitivity of a parameter is calculated by using the following equations. The closer the sensitivity is to zero, the more reasonable the parameter is.
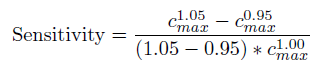
Assumptions
Our model consists of 53 parameters, which makes the modeling process very difficult. To reduce the amount of work without losing quality, we proposed the following assumptions.
The average transcription speed in E.coli is 70nt/s. Assuming all the transcription in our circuit works in such speed, we can calculate the maximum transcription rate for each transcription equation by using this formula:
Maximum Transcription Rate = Transcription Speed(nt/min)/Gene Length(bp)=4200/Gene Length (nM/min)
Ref: http://gchelpdesk.ualberta.ca/CCDB/cgi-bin/STAT_NEW.cgi, NTU-Singapore iGEM2008 Team
The average translation speed in E.coli is 40Aa/s. Also assuming all the translation in our circuit works in the same speed, we can calculate the theoretical transcription rate. However, in wetlab, we can use different rbs to regulate the translation process, thus, the translation rate can be written as:
Translation Rate = RBS * Translation Speed(Aa/min)/Protein Length(Aa) = 2400RBS/Protein Length (min^-1)
This transformation does not change the degree of freedom of our system. However, this does limit the range of parameters since the strength of RBS can not be too extreme.
Ref: http://gchelpdesk.ualberta.ca/CCDB/cgi-bin/STAT_NEW.cgi, NTU-Singapore iGEM2008 Team
We have assume the period of cell division is 30 mins, which means the "degradation rate" in our model is actually the sum of degradation rate of the substance(1/half life) and cell division rate(1/30 mins).
From Ref1(Belasco 1993) and Ref2(Genome Biology 2006, 7:R99), we have decided that all the mRNA in our system have a half life of 4.4 mins.
Modeling - T3 RNA polymerase
We have construct two models, the difference of which is in the AND Gate 2 module. In this section, we'll demonstrate the parameters of our first model, in which T3 RNA polymerase mRNA with amber mutation and Aa-tRNA cooperate to produce T3 RNA polymerase protein.
| Parameters | Brief Introduction | Value | Unit | Reference/Sensitivity
|
| k_1 | Max Transcription rate of tRNA | 46.67 | nM/min | Assumption, 0.19
|
| k_2 | Synthesis rate of Aa-tRNA | 0.08 | min^-1 | 0.09
|
| k_3 | Max Transcription rate of T7RNAP | 1.5625 | nM/min | Assumption, 0.00
|
| k_4 | Max Translation rate of T7RNAP | 2.68*0.05 | min^-1 | Assumption, 0.00
|
| k_5 | Max Transcription rate of trigger CI | 5.6 | nM/min | Assumption, 0.00
|
| k_6 | Transcription rate of bistable CI | 5.6 | nM/min | Assumption, 0.00
|
| k_6' | Transcription rate of bistable CI | 1 | nM/min | 0.00
|
| k_7 | Transcription rate of T3RNAP | 1.75 | nM/min | Assumption, 1.34
|
| k_7' | Transcription rate of T3RNAP | 1 | nM/min | 0.00
|
| k_8 | Translation rate of trigger CI | 9.6*0.045 | min^-1 | Assumption, 0.00
|
| k_8' | Translation rate of bistable CI | 9.6*0.3 | min^-1 | Assumption, 0.00
|
| k_9 | Max Transcription rate of CI434 | 5.92 | nM/min | Assumption, 0.00
|
| k_10 | Transcription rate of CI434 | 10.14*0.5 | min^-1 | Assumption, 0.00
|
| k_11 | Max Translation rate of T3RNAP | 3*0.15 | min^-1 | Assumption, 1.34
|
| k_12 | Max Transcription rate of GFP from Sal | 5.25 | nM/min | Assumption, 0.00
|
| k_12' | Max Trasncription rate of GFP from T3RNAP | 5.25 | nM/min | Assumption, 1.00
|
| k_13 | Translation rate of GFP | 9*0.6 | min^-1 | Assumption, 1.00
|
| k_s | rate of AND Gate 1 | 0.3 | nM^-1 | 0.00
|
| k_s' | rate of AND Gate 2 | 0.3 | nM^-1 | 0.18
|
| K_1 | dissociation constant of AraC,tRNA | 14 | nM | 0.03
|
| K_3 | dissociation constant of Sal,T7RNAP | 0.5 | nM | 0.00
|
| K_5 | dissociation constant of T7RNAP,trigger CI | 3 | nM | [http://bionumbers.hms.harvard.edu/bionumber.aspx?s=y&id=103592&ver=1 Ref]
|
| K_6 | dissociation constant of CI,bistable CI | 40 | nM | Ref: iGEM2007 PKU Team
|
| K_6' | dissociation constant of CI434,bistable CI | 50 | nM | Ref: iGEM2007 PKU Team
|
| K_7 | dissociation constant of CI,T3RNAP | 40 | nM | Ref: iGEM2007 PKU Team
|
| K_7' | dissociation constant of CI434,T3RNAP | 50 | nM | Ref: iGEM2007 PKU Team
|
| K_9 | dissociation constant of CI,CI434 | 40 | nM | Ref: iGEM2007 PKU Team
|
| K_12 | dissociation constant of Sal,GFP | 0.5 | nM | 0.00
|
| K_12' | dissociation constant of T3RNAP,GFP | 55 | nM | Ref: The FEBS journal 2006,273:17
|
| n_1 | Hill co-effiency of AraC,tRNA | 2 | |
|
| n_3 | Hill co-effiency of Sal,T7RNAP | 2 | |
|
| n_5 | Hill co-effiency of T7RNAP,CI | 2 | |
|
| n_6 | Hill co-effiency of CI,bistable CI | 4 | | Ref: iGEM2007 PKU Team
|
| n_6' | Hill co-effiency of CI434,bistable CI | 2 | | Ref: iGEM2007 PKU Team
|
| n_7 | Hill co-effiency of CI,T3RNAP | 4 | | Ref: iGEM2007 PKU Team
|
| n_7' | Hill co-effiency of CI434,T3RNAP | 2 | | Ref: iGEM2007 PKU Team
|
| n_9 | Hill co-effiency of CI,CI434 | 4 | | Ref: iGEM2007 PKU Team
|
| n_12 | Hill co-effiency of Sal,GFP | 2 | |
|
| n_12' | Hill co-effiency of T3RNAP,GFP | 2 | |
|
| γ_1 | Degradation rate of tRNA | 1/30+1/60 | min^-1 | Since half life of tRNA is very long,
we decided to use 60 mins instead
|
| γ_2 | Degradation rate of Aa-tRNA | 1/30+1/40 | min^-1 |
|
| γ_2' | Real Degradation rate of Aa-tRNA | 1/40 | min^-1 |
|
| γ_3 | Degradation rate of T7RNAP mRNA | 1/30+1/4.4 | min^-1 | Assumption
|
| γ_4 | Degradation rate of T7RNAP | 1/30+1/40 | min^-1 |
|
| γ_5 | Degradation rate of trigger CI mRNA | 1/30+1/4.4 | min^-1 | Assumption
|
| γ_6 | Degradation rate of bistable CI mRNA | 1/30+1/4.4 | min^-1 | Assumption
|
| γ_7 | Degradation rate of T3RNAP mRNA | 1/30+1/4.4 | min^-1 | Assumption
|
| γ_8 | Degradation rate of CI | 1/30+1/44 | min^-1 | Ref: iGEM2007 PKU Team
|
| γ_9 | Degradation rate of CI434 mRNA | 1/30+1/4.4 | min^-1 | Assumption
|
| γ_10 | Degradation rate of CI434 | 1/30+1/11 | min^-1 | Ref: iGEM 2007 PKU Team
|
| γ_11 | Degradation rate of T3RNAP | 1/30+1/30 | min^-1 |
|
| γ_12 | Degradation rate of GFP mRNA | 1/30+1/4.4 | min^-1 | Assumption
|
| γ_13 | Degradation rate of GFP | 1/30+1/60 | min^-1 | Since half life of GFP is very long,
we use 60 mins instead
|
Overall, the sensitivity of parameters from trial and error is generally low. The bi-stable-related parameters' sensitivity indicates that the bi-stable model, which is the core in the circuit, is very stable. With all of these facts, we have concluded that this model is reasonable.
Modeling - P2
Here's our second model, the one with P2 instead of T3 RNA polymerase
| Parameters | Brief Introduction | Value | Unit | Reference/Sensitivity
|
| k_1 | Max Transcription rate of tRNA | 46.67 | nM/min | Assumption, 1.41
|
| k_2 | Synthesis rate of Aa-tRNA | 0.08 | min^-1 | 0.81
|
| k_3 | Max Transcription rate of T7RNAP | 1.5625 | nM/min | Assumption, 0.00
|
| k_4 | Max Translation rate of T7RNAP | 2.68*0.05 | min^-1 | Assumption, 0.00
|
| k_5 | Max Transcription rate of trigger CI | 5.6 | nM/min | Assumption, 0.00
|
| k_6 | Transcription rate of bistable CI | 5.6 | nM/min | Assumption, 0.00
|
| k_6' | Transcription rate of bistable CI | 1 | nM/min | 0.00
|
| k_7 | Transcription rate of P2 | 16.8 | nM/min | Assumption, 1.23
|
| k_7' | Transcription rate of P2 | 1 | nM/min | 0.00
|
| k_8 | Translation rate of trigger CI | 9.6*0.05 | min^-1 | Assumption, 0.00
|
| k_8' | Translation rate of bistable CI | 9.6*0.5 | min^-1 | Assumption, 0.00
|
| k_9 | Max Transcription rate of CI434 | 5.92 | nM/min | Assumption, 0.00
|
| k_10 | Transcription rate of CI434 | 10.14*1 | min^-1 | Assumption, 0.00
|
| k_11 | Max Translation rate of P2 | 28.8*0.0045 | min^-1 | Assumption, 1.23
|
| k_12 | Max Transcription rate of GFP from Sal | 5.25 | nM/min | Assumption, 0.00
|
| k_12' | Max Trasncription rate of GFP from P2 | 5.25 | nM/min | Assumption, 1.00
|
| k_13 | Translation rate of GFP | 9*0.6 | min^-1 | Assumption, 1.00
|
| k_s | rate of AND Gate 1 | 0.3 | nM^-1 | 0.00
|
| k_s' | rate of AND Gate 2 | 0.01 | nM^-1 | 1.29
|
| K_1 | dissociation constant of AraC,tRNA | 14 | nM | 0.20
|
| K_3 | dissociation constant of Sal,T7RNAP | 0.5 | nM | 0.00
|
| K_5 | dissociation constant of T7RNAP,trigger CI | 3 | nM | [http://bionumbers.hms.harvard.edu/bionumber.aspx?s=y&id=103592&ver=1 Ref]
|
| K_6 | dissociation constant of CI,bistable CI | 40 | nM | Ref: iGEM2007 PKU Team
|
| K_6' | dissociation constant of CI434,bistable CI | 50 | nM | Ref: iGEM2007 PKU Team
|
| K_7 | dissociation constant of CI,P2 | 40 | nM | Ref: iGEM2007 PKU Team
|
| K_7' | dissociation constant of CI434,P2 | 50 | nM | Ref: iGEM2007 PKU Team
|
| K_9 | dissociation constant of CI,CI434 | 40 | nM | Ref: iGEM2007 PKU Team
|
| K_12 | dissociation constant of Sal,GFP | 0.5 | nM | 0.00
|
| K_12' | dissociation constant of P2,GFP | 35 | nM | 1.29
|
| n_1 | Hill co-effiency of AraC,tRNA | 2 | |
|
| n_3 | Hill co-effiency of Sal,T7RNAP | 2 | |
|
| n_5 | Hill co-effiency of T7RNAP,CI | 2 | |
|
| n_6 | Hill co-effiency of CI,bistable CI | 4 | | Ref: iGEM2007 PKU Team
|
| n_6' | Hill co-effiency of CI434,bistable CI | 2 | | Ref: iGEM2007 PKU Team
|
| n_7 | Hill co-effiency of CI,P2 | 4 | | Ref: iGEM2007 PKU Team
|
| n_7' | Hill co-effiency of CI434,P2 | 2 | | Ref: iGEM2007 PKU Team
|
| n_9 | Hill co-effiency of CI,CI434 | 4 | | Ref: iGEM2007 PKU Team
|
| n_12 | Hill co-effiency of Sal,GFP | 2 | |
|
| n_12' | Hill co-effiency of P2,GFP | 2 | |
|
| γ_1 | Degradation rate of tRNA | 1/30+1/60 | min^-1 | Since half life of tRNA is very long,
we decided to use 60 mins instead
|
| γ_2 | Degradation rate of Aa-tRNA | 1/30+1/40 | min^-1 |
|
| γ_2' | Real Degradation rate of Aa-tRNA | 1/40 | min^-1 |
|
| γ_3 | Degradation rate of T7RNAP mRNA | 1/30+1/4.4 | min^-1 | Assumption
|
| γ_4 | Degradation rate of T7RNAP | 1/30+1/40 | min^-1 |
|
| γ_5 | Degradation rate of trigger CI mRNA | 1/30+1/4.4 | min^-1 | Assumption
|
| γ_6 | Degradation rate of bistable CI mRNA | 1/30+1/4.4 | min^-1 | Assumption
|
| γ_7 | Degradation rate of P2 mRNA | 1/30+1/4.4 | min^-1 | Assumption
|
| γ_8 | Degradation rate of CI | 1/30+1/44 | min^-1 | Ref: iGEM2007 PKU Team
|
| γ_9 | Degradation rate of CI434 mRNA | 1/30+1/4.4 | min^-1 | Assumption
|
| γ_10 | Degradation rate of CI434 | 1/30+1/11 | min^-1 | Ref: iGEM 2007 PKU Team
|
| γ_11 | Degradation rate of P2 | 1/30+1/30 | min^-1 |
|
| γ_12 | Degradation rate of GFP mRNA | 1/30+1/4.4 | min^-1 | Assumption
|
| γ_13 | Degradation rate of GFP | 1/30+1/60 | min^-1 | Since half life of GFP is very long,
we use 60 mins instead
|
Overall, the sensitivity of parameters from trial and error is also low. The bi-stable is also shows great stability. However, we consider the value in AND Gate 2 is too extreme. Although this fact doesn't indicate that the circuit design is wrong or we can't finish the weblab project theoretically, we decided that it's very necessary to construct the circuit with T3 RNA polymerase.
With all necessities prepared, it's time to see the result!
^Top
|