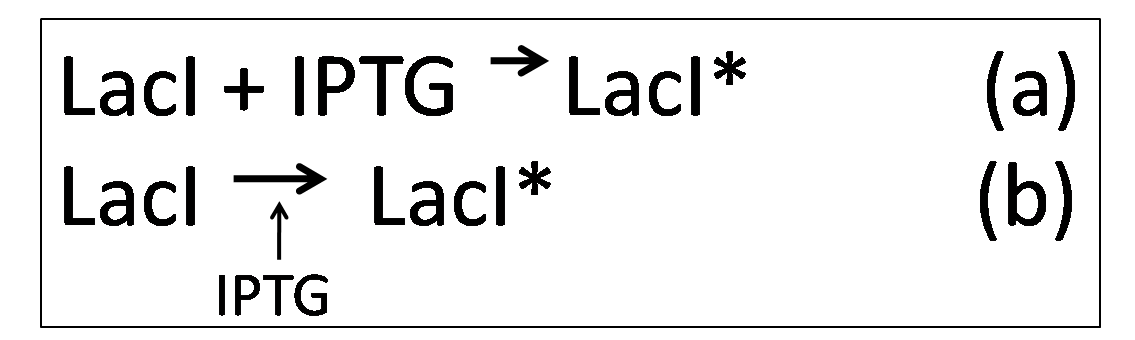Team:Newcastle/SporulationTuning
From 2009.igem.org
(→Introduction) |
(→KinA Expression Model) |
||
| (12 intermediate revisions not shown) | |||
| Line 7: | Line 7: | ||
==Introduction== | ==Introduction== | ||
| - | The | + | The bacterium ''Bacillus subtilis'' used in our project is a gram-positive soil bacterium which, under certain conditions, commits itself to a developmental pathway leading to the production of spores.<sup>[1]</sup> In this part of our project, we aim to control sporulation in our bacterial population, so that we can decide how much of the population becomes spores, and how much continue as vegetative cells. Should the cell sporulate, it becomes a [https://2009.igem.org/Team:Newcastle/Metalintakeefflux ‘metal container’], trapping the sequestered cadmium in its spore. |
| - | After the cell sequesters cadmium into its spore, it should not germinate or the sequestered cadmium will be released back into the environment | + | After the cell sequesters cadmium into its spore, it should not germinate, or the sequestered cadmium will be released back into the environment. Therefore, the [https://2009.igem.org/Team:Newcastle/Chassis chassis] comes into play, where the ''sleB'' and ''cwlJ'' germination-defective mutants are put into use. |
| - | In order to control sporulation, our team | + | In order to control sporulation, our team proposed the idea of inducing the synthesis of ''kinA'', with IPTG as a sporulation initiation signal. |
| - | [https://2009.igem.org/Team:Newcastle/SporulationTuning/Introduction#KinA KinA] is a major kinase which provides phosphate input to | + | [https://2009.igem.org/Team:Newcastle/SporulationTuning/Introduction#KinA ''KinA''] is a major kinase which provides phosphate input to a phosphorelay, which in turn, activates the sporulation pathway upon starvation via the phosphorylated Spo0A transcription factor,<sup>[2]</sup> which governs entry into the sporulation pathways of the bacterium ''Bacillus subtilis''.<sup>[3]</sup> |
:''[[Team:Newcastle/SporulationTuning/Introduction#Spo0A| ... Click to read more ...]] | :''[[Team:Newcastle/SporulationTuning/Introduction#Spo0A| ... Click to read more ...]] | ||
| Line 21: | Line 21: | ||
==Novelty in this sub-project== | ==Novelty in this sub-project== | ||
| - | In this sub-project, instead of allowing the cell to decide whether or not to sporulate, we | + | In this sub-project, instead of allowing the cell to decide whether or not to sporulate, we influence its decision. In order to execute our plan, we used the concentration of ''kinA'', induced by IPTG, to control sporulation. |
| - | + | This sub-project consists of two main models: the Sporulation Tuning and the Sin Operon models. These two models are meant to work together, as mentioned below in the [https://2009.igem.org/Team:Newcastle/SporulationTuning#Modelling modelling section], with the Sporulation Tuning model controlling sporulation, and the Sin Operon model repressing sporulation, creating a more realistic model. | |
==Modelling== | ==Modelling== | ||
| - | The Sin (sporulation inhibition) Operon Model was one of the earlier models built. As its name suggests, it | + | The Sin (sporulation inhibition) Operon Model was one of the earlier models built. As its name suggests, it models the repression of sporulation. The Sin Operon Model was built in CellML. |
The second model built was the simple model of KinA expression. After satisfactory results were obtained, the sporulation phosphorelay was modelled into the KinA Expression Model, and is known as the Sporulation Tuning Model. Both the KinA Expression and Sporulation Tuning models were built using COPASI. | The second model built was the simple model of KinA expression. After satisfactory results were obtained, the sporulation phosphorelay was modelled into the KinA Expression Model, and is known as the Sporulation Tuning Model. Both the KinA Expression and Sporulation Tuning models were built using COPASI. | ||
| - | The Sin Operon and Sporulation Tuning model | + | The Sin Operon and Sporulation Tuning model work hand in hand as components of the [https://2009.igem.org/Team:Newcastle/PopulationDynamics Population Dynamics model]. |
===KinA Expression Model=== | ===KinA Expression Model=== | ||
| - | Under normal conditions, LacI represses | + | Under normal conditions, LacI represses ''kinA''. |
[[Image:TeamNewcastleKinAExpLacIKinA.png|110px|center]] | [[Image:TeamNewcastleKinAExpLacIKinA.png|110px|center]] | ||
| Line 41: | Line 41: | ||
| - | However, in the presence of IPTG, KinA can be expressed, as IPTG binds to | + | However, in the presence of IPTG, KinA can be expressed, as IPTG binds to ''lacI'', deactivating it. Equations (a) and (b) describes how IPTG binds to LacI, forming LacI*, which is the deactivated form of LacI |
[[Image:TeamNewcastleKinAExpLacIIPTG1.png|220px|center]] | [[Image:TeamNewcastleKinAExpLacIIPTG1.png|220px|center]] | ||
| - | :''[[Team:Newcastle/Modelling/KinAExpression#KinA_Expression_Model| Click to view | + | :''[[Team:Newcastle/Modelling/KinAExpression#KinA_Expression_Model| Click to view more of the KinA Expression Model equations]]'' |
<br> | <br> | ||
| Line 61: | Line 61: | ||
===Sporulation Tuning Model=== | ===Sporulation Tuning Model=== | ||
| - | The expression of KinA has been modelled as seen above, therefore we can now proceed further into the Sporulation Tuning Model, which is built from the KinA Expression Model | + | The expression of KinA has been modelled as seen above, therefore we can now proceed further into the Sporulation Tuning Model, which is built from the KinA Expression Model using COPASI. |
| - | To proceed with the modelling of our Sporulation Tuning Model, we have decided that in response to an unidentified | + | To proceed with the modelling of our Sporulation Tuning Model, we have decided that in response to an unidentified stimulus, where KinA autophosphorylates and then donates its phosphate groups to the response regulator Spo0F, the unidentified stimuli will be termed as 'sporulation signal'.<sup>[5]</sup> |
The following equations describe the model: | The following equations describe the model: | ||
| Line 80: | Line 80: | ||
:''[[Team:Newcastle/Modelling/SporulationTuning#Sporulation_Tuning_Model| Click to view more of the Sporulation Tuning Model equations]]'' | :''[[Team:Newcastle/Modelling/SporulationTuning#Sporulation_Tuning_Model| Click to view more of the Sporulation Tuning Model equations]]'' | ||
| + | |||
| + | <br> | ||
====Results==== | ====Results==== | ||
| Line 87: | Line 89: | ||
:''[[Team:Newcastle/Modelling/SporulationTuning#Results| Click to view the Sporulation Tuning results]]'' | :''[[Team:Newcastle/Modelling/SporulationTuning#Results| Click to view the Sporulation Tuning results]]'' | ||
| + | |||
| + | <br> | ||
===Sin (sporulation inhibition) Operon Model=== | ===Sin (sporulation inhibition) Operon Model=== | ||
| - | In order to create a more realistic model of our sporulation system | + | In order to create a more realistic model of our sporulation system the team has decided to include the Sin (sporulation inhibition) Operon Model, which the team designed in CellML. |
The sin operon controls the production and activity of the repressor SinR, which in its active tetrameric form, inhibits sporulation by repressing stage II and spo0A promoters. On the other hand, the accumulation of Spo0A~P induces the expression of SinI, which binds to and inactivates SinR. | The sin operon controls the production and activity of the repressor SinR, which in its active tetrameric form, inhibits sporulation by repressing stage II and spo0A promoters. On the other hand, the accumulation of Spo0A~P induces the expression of SinI, which binds to and inactivates SinR. | ||
| Line 98: | Line 102: | ||
| - | + | :''[[Team:Newcastle/Modelling/SinOperon| Click to view more of the Sin Operon Model equations]]'' | |
| - | :'' | + | |
| + | <br> | ||
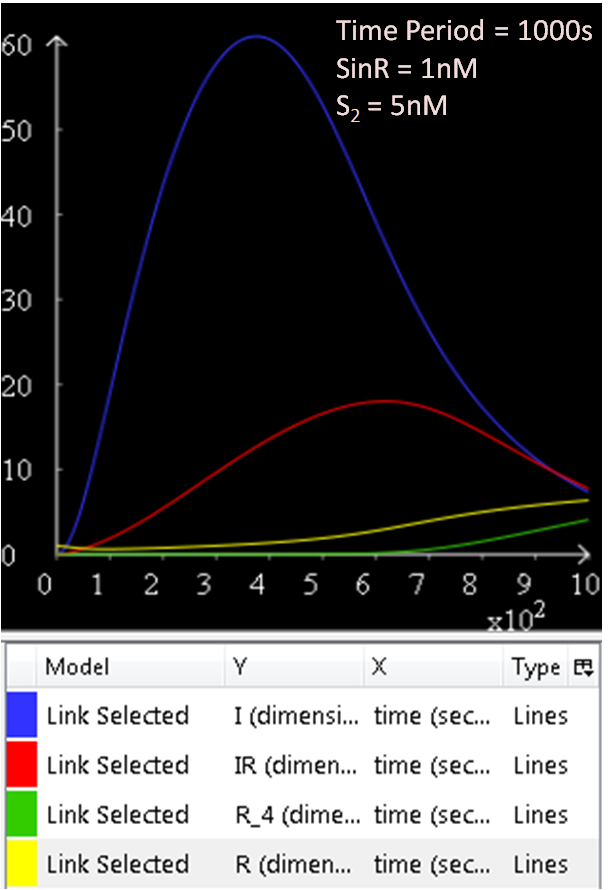
====Results==== | ====Results==== | ||
| Line 106: | Line 111: | ||
| - | + | :''[[Team:Newcastle/Modelling/SinOperon#Results| Click to view the Sin Operon Model results]]'' | |
| - | :'' | + | |
| + | <br> | ||
==BioBrick constructs== | ==BioBrick constructs== | ||
| - | The BioBrick we have designed is to contain an IPTG inducable kinA gene, using pSpac, allowing us to test the theory about KinA in the lab. | + | The BioBrick we have designed is to contain an IPTG inducable ''kinA'' gene, using pSpac, allowing us to test the theory about KinA in the lab. |
'''BBa_K174010''' | '''BBa_K174010''' | ||
| Line 137: | Line 143: | ||
==Lab Work Strategies== | ==Lab Work Strategies== | ||
| - | The lab work executed | + | The lab work executed was be to induce sporulation in the presence of IPTG, and this involved microscopy and testing cultures for sporulation (e.g. by their heat resistance). |
[[Image:Newcastle KinA-sporulation1.JPG|center|400px]] | [[Image:Newcastle KinA-sporulation1.JPG|center|400px]] | ||
| Line 163: | Line 169: | ||
[9] Voigt, C. A., Wolf, D. M., Arkin, A. P. (2004) The ''Bacillus subtilis sin'' Operon: An Evolvable Network Motif. Genetics 169: 1187-1202 | [9] Voigt, C. A., Wolf, D. M., Arkin, A. P. (2004) The ''Bacillus subtilis sin'' Operon: An Evolvable Network Motif. Genetics 169: 1187-1202 | ||
| + | |||
| + | <br> | ||
Latest revision as of 02:11, 22 October 2009
Sporulation Tuning
Introduction
The bacterium Bacillus subtilis used in our project is a gram-positive soil bacterium which, under certain conditions, commits itself to a developmental pathway leading to the production of spores.[1] In this part of our project, we aim to control sporulation in our bacterial population, so that we can decide how much of the population becomes spores, and how much continue as vegetative cells. Should the cell sporulate, it becomes a ‘metal container’, trapping the sequestered cadmium in its spore.
After the cell sequesters cadmium into its spore, it should not germinate, or the sequestered cadmium will be released back into the environment. Therefore, the chassis comes into play, where the sleB and cwlJ germination-defective mutants are put into use.
In order to control sporulation, our team proposed the idea of inducing the synthesis of kinA, with IPTG as a sporulation initiation signal.
KinA is a major kinase which provides phosphate input to a phosphorelay, which in turn, activates the sporulation pathway upon starvation via the phosphorylated Spo0A transcription factor,[2] which governs entry into the sporulation pathways of the bacterium Bacillus subtilis.[3]
Novelty in this sub-project
In this sub-project, instead of allowing the cell to decide whether or not to sporulate, we influence its decision. In order to execute our plan, we used the concentration of kinA, induced by IPTG, to control sporulation.
This sub-project consists of two main models: the Sporulation Tuning and the Sin Operon models. These two models are meant to work together, as mentioned below in the modelling section, with the Sporulation Tuning model controlling sporulation, and the Sin Operon model repressing sporulation, creating a more realistic model.
Modelling
The Sin (sporulation inhibition) Operon Model was one of the earlier models built. As its name suggests, it models the repression of sporulation. The Sin Operon Model was built in CellML.
The second model built was the simple model of KinA expression. After satisfactory results were obtained, the sporulation phosphorelay was modelled into the KinA Expression Model, and is known as the Sporulation Tuning Model. Both the KinA Expression and Sporulation Tuning models were built using COPASI.
The Sin Operon and Sporulation Tuning model work hand in hand as components of the Population Dynamics model.
KinA Expression Model
Under normal conditions, LacI represses kinA.
However, in the presence of IPTG, KinA can be expressed, as IPTG binds to lacI, deactivating it. Equations (a) and (b) describes how IPTG binds to LacI, forming LacI*, which is the deactivated form of LacI
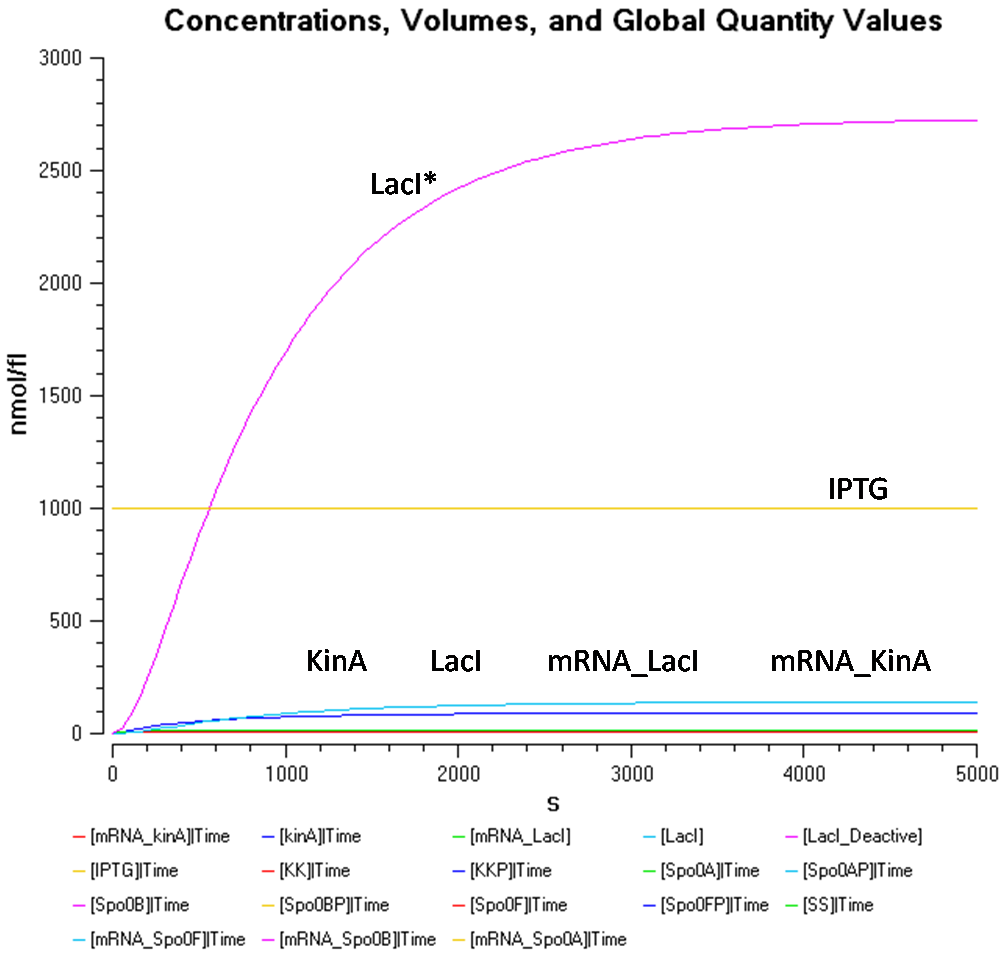
Results
Sporulation Tuning Model
The expression of KinA has been modelled as seen above, therefore we can now proceed further into the Sporulation Tuning Model, which is built from the KinA Expression Model using COPASI.
To proceed with the modelling of our Sporulation Tuning Model, we have decided that in response to an unidentified stimulus, where KinA autophosphorylates and then donates its phosphate groups to the response regulator Spo0F, the unidentified stimuli will be termed as 'sporulation signal'.[5]
The following equations describe the model:
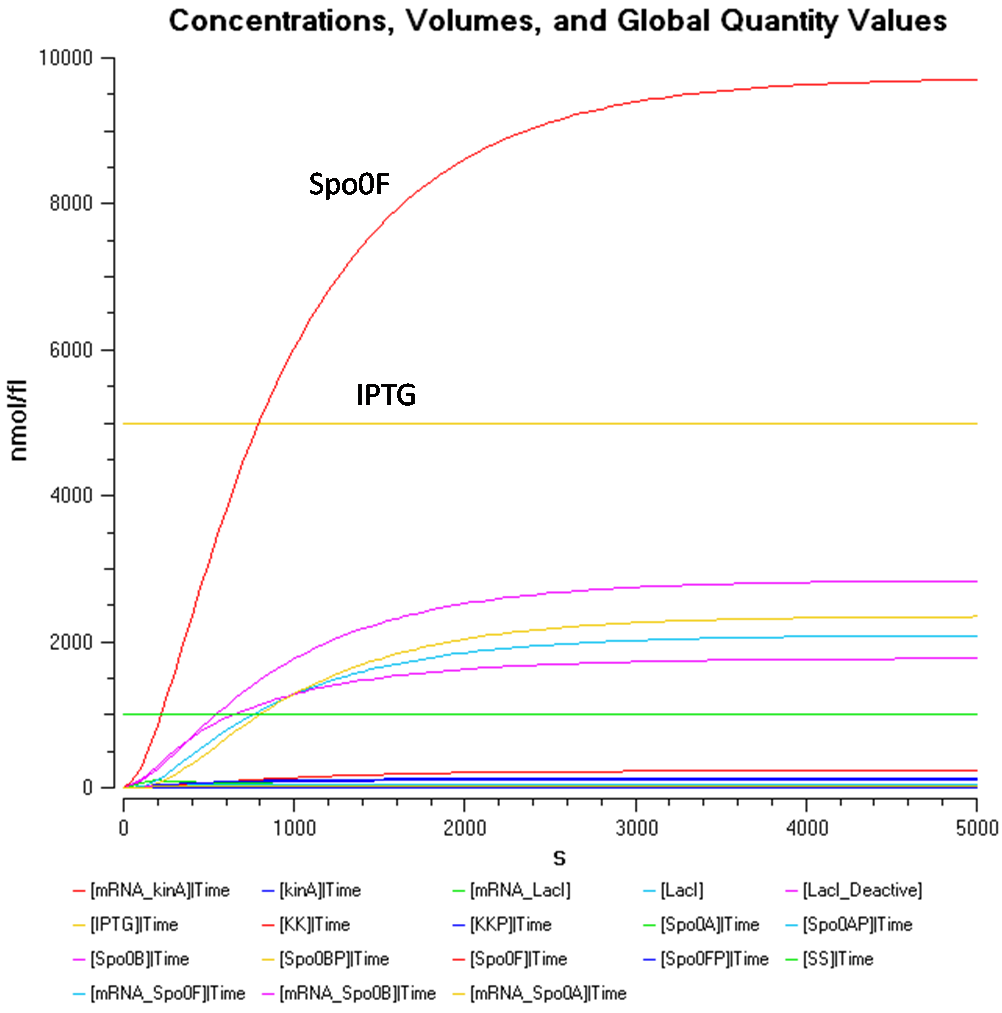
Results
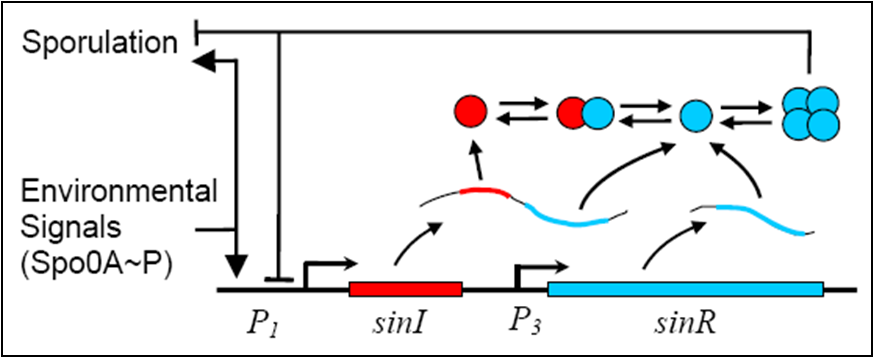
Sin (sporulation inhibition) Operon Model
In order to create a more realistic model of our sporulation system the team has decided to include the Sin (sporulation inhibition) Operon Model, which the team designed in CellML.
The sin operon controls the production and activity of the repressor SinR, which in its active tetrameric form, inhibits sporulation by repressing stage II and spo0A promoters. On the other hand, the accumulation of Spo0A~P induces the expression of SinI, which binds to and inactivates SinR.
Results
BioBrick constructs
The BioBrick we have designed is to contain an IPTG inducable kinA gene, using pSpac, allowing us to test the theory about KinA in the lab.
BBa_K174010
KinA
Length: 1818bp
Click [http://partsregistry.org/wiki/index.php?title=Part:BBa_K174010 here] for more information on this part.
BBa_K174011
IPTG inducable KinA sporulation trigger
Length: 1953bp
Click [http://partsregistry.org/wiki/index.php?title=Part:BBa_K174011 here] for more information on this part.
Lab Work Strategies
The lab work executed was be to induce sporulation in the presence of IPTG, and this involved microscopy and testing cultures for sporulation (e.g. by their heat resistance).
To find out more about our lab work strategies,
References
[1] Predich, M., Nair, G., Smith, I. (1992) Bacillus subtilis Early Sporulation Genes kinA, spo0F, and spo0A Are Transcribed by the RNA Polymerase Containing σH. Journal of Bacteriology. Pp 2771-2778
[2] Veening, J-W., Smits, W. K., Kuipers, O. P. (2008) Bistability, Epigenetics, and Bet-Hedging in Bacteria. Annu. Rev. Microbiol. 62: 193-210
[3] Fujita, M., Losick, R. (2005) Evidence that Entry into Sporulation in Bacillus subtilis is Governed by a Gradual Increase in the Level and Activity of the Master Regulator Spo0A. 19: 2236–2244
[4] Sonenshein, A.L., Hoch, J.A., Losick, R., (2002) Bacillus subtilis and Its Closest Relatives From Genes to Cells. ASM Press, United States of America. Pp 476–477
[5] Hilbert, D.W., Piggot, P.J., (June 2004) Compartmentalization of Gene Expression during Bacillus subtilis Spore Formation. Microbiology and Molecular Biology Reviews. Vol. 68, No. 2. Pp 234-262
[6] Eswaramoorthy, P., Guo, T., Fujita, M. (2009) In Vivo Domain-Based Functional Analysis of the Major Sporulation Sensor Kinase, KinA, in Bacillus subtilis. Journal of Bacteriology. Pp 5358-5368
[7] Stephenson, K., Hoch, J. A. (2001) PAS-A domain of phosphorelay sensor kinase A: A catalytic ATP-binding domain involved in the initiation of development in Bacillus subtilis. PNAS. Vol. 98, no. 26: 15251-15256
[8] Veening, J-W., Hamoen, L. W., Kuipers, O. P. (2005) Phosphatases modulate the bistable sporulation gene expression pattern in Bacillus subtilis. Molecular Microbiology 56(6), 1481-1494
[9] Voigt, C. A., Wolf, D. M., Arkin, A. P. (2004) The Bacillus subtilis sin Operon: An Evolvable Network Motif. Genetics 169: 1187-1202
 "
"