Team:Newcastle/Promoter Library
From 2009.igem.org
Promoter Library
Introduction
In order to effectively model genetic circuits, databases of parts with known characteristics are essential. Without such databases, computationally designed circuits may not be feasible to implement in vivo. Unfortunately, parameters such as promoter strength, RBS affinity, transcription, translation and decay rates are hard to find in the literature, and those which do exist are rarely measured under standardised conditions.
Several efforts to produce parts libraries are currently underway.As part of our contribution to synthetic biology, we decided to create a promoter library for B. subtilis.
Novelty in this sub-project
The novelty in this sub-project lies in the generation of a large number of B. subtilis promoters, based upon carefully considered variations to an existing promoter region. We chose the promoter region of the transcription factor SigmaA, since it is the most widely-used promoter in B. subtilis. Over 300 genes are controlled by SigA. Their promoter regions have highly conserved -10 and -35 regions, surrounded by variable sequences.
Design
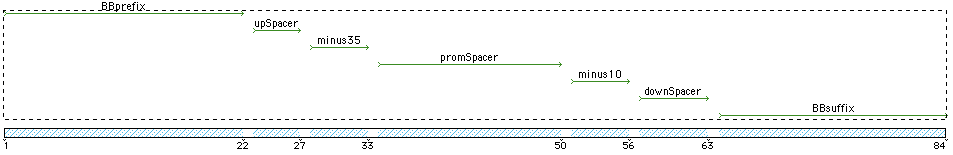
Template used to create the promoter sequences:
Prefix...UpSpacer...-35...PromSpacer...-10...DownSpacer..Suffix
- BBPrefix - StandardBioBrickPrefix EcoR1 and Xba1 - gaattcgcggccgcttctagag
- UpSpacer - attta - consensus sigA 5 bases 5'
- -35 - TTGACA
- PromSpacer - length 17 - consensus ttttatttaaattatga
- -10 - TATAAT
- DownSpacer (from ackA)- ggaaaag
- BBSuffix - Standard BioBrick suffix Spe1 and Pst1 - tactagtagcggccgctgcag
Variances
We designed three variances to create a library of promoters with different strengths.
Variance 1
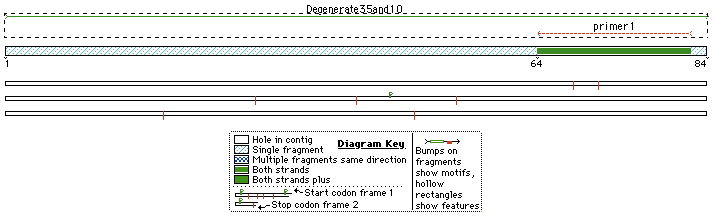
In this variance we degenerate -35 and -10 sequences.
BBPrefix...UpSpacer...N-35... PromSpacer...N-10...DownSpacer..BBSuffix
Variance 2
In the second variance we use N's for the promoter sequence between -35 and -10 regions
BBPrefix...UpSpacer...-35... 18N’s...-10...DownSpacer..BBSuffix
Variance 3
In the third variance we change the last two bases of minus 35 region and the first two bases of -10 region.
BBPrefix...UpSpacerNN...-35...NNPromSpacerNN...-10...NNDownSpacer..BBSuffix
Lab Work Strategies
- Synthesis other strand using polymerase and a primer
- Cut promoter fragments using EcoR1 and Pst1
- Cut pSB1AT3 vector using EcoR1 and Pst1
- Ligate the insert and the plasmid backbone, and transform E. coli DH5alpha
- Subclone into Bacillus integration vector with gfp and insert at amyE
News
Events
- 20 – 21 June 2009 - Europe workshop (London)
- 23 – 24 June 2009 - UK iGEM meetup (Edinburgh)
- 23 October Practice Presentation (Newcastle)
- 23 October T-shirts are ready
- 27 October Practice Presentation (Sunderland)
- 27 October Poster is ready
- 30 October – 2 November 2009 - Jamboree (Boston)
Social Net
- Newcastle iGEM Twitter
- [http://www.facebook.com/home.php#/group.php?gid=131709337641 Newcastle on Facebook]
- [http://www.youtube.com/user/newcastle2009igem Newcastle Youtube Channel]
 "
"