Team:TUDelft/24 July 2009
From 2009.igem.org
(→24 July 2009) |
|||
| Line 1: | Line 1: | ||
{{Template:TUDelftiGEM2009}} | {{Template:TUDelftiGEM2009}} | ||
| + | |||
| + | {{Template:TUDelftiGEM2009_LabNotebook}} | ||
='''24 July 2009'''= | ='''24 July 2009'''= | ||
| Line 7: | Line 9: | ||
The biobricks received from MIT for Delay [S03335, S03473] which were streaked yesterday have the colonies for future. | The biobricks received from MIT for Delay [S03335, S03473] which were streaked yesterday have the colonies for future. | ||
| + | |||
===Tim Vos=== | ===Tim Vos=== | ||
Glycerol stocks were prepared for the remaining cultures grown in all the culture tubes [R0040, J23008, J23031, B0015, K081013, S03335, S03473, I714031, E0840, J23100]that were incubated yesterday and stored in -80°C freezer. | Glycerol stocks were prepared for the remaining cultures grown in all the culture tubes [R0040, J23008, J23031, B0015, K081013, S03335, S03473, I714031, E0840, J23100]that were incubated yesterday and stored in -80°C freezer. | ||
| Line 15: | Line 18: | ||
===Orr=== | ===Orr=== | ||
Today, I performed the mfold analysis to find out the secondary structures of the key3c and the lock3c. I also started looking at the COSMIC code that is intended to model conjugation. So far I only downloaded the source codes, and still need to find out how to run it. | Today, I performed the mfold analysis to find out the secondary structures of the key3c and the lock3c. I also started looking at the COSMIC code that is intended to model conjugation. So far I only downloaded the source codes, and still need to find out how to run it. | ||
| + | |||
| + | ===Calin=== | ||
| + | Updated Matlab code to fix mRNA / protein confusion. Got growth in all tubes and plates. MIT parts placed in fridge. Tube cultures went to Sriram's miniprep for plasmid extraction. Did some GFP / RFP non-quantitative measurements. These can be done in the gel imaging machine. <br> | ||
| + | [[Image:Gfprfp24072009.png]] | ||
{{Template:TUDelftiGEM2009_end}} | {{Template:TUDelftiGEM2009_end}} | ||
Revision as of 14:41, 25 July 2009
Lab Notebook
|
|
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
24 July 2009
Sriram
The cultures were grown in all the culture tubes [R0040, J23008, J23031, B0015, K081013, S03335, S03473, I714031, E0840, J23100]incubated yesterday except RBS[B0034] and pLacI[R0010], which may be because the colony we picked for those biobricks were not perfect. The plasmid DNA was extracted from all the grown cultures and stored in -20°C freezer.
The biobricks received from MIT for Delay [S03335, S03473] which were streaked yesterday have the colonies for future.
Tim Vos
Glycerol stocks were prepared for the remaining cultures grown in all the culture tubes [R0040, J23008, J23031, B0015, K081013, S03335, S03473, I714031, E0840, J23100]that were incubated yesterday and stored in -80°C freezer.
Daniel
Since the top10 competent cells newly prepared by us using the Openwetware protocol, didn't transform well for Tim Weenink we are in a bit crisis for competent cells. Anyway to make sure whether the cells work or not I did transformation of Biobricks RBS[B0034] and pLacI[R0010] in the newly created top10 competent cells and kept in drawer to grow in the weekend.
Orr
Today, I performed the mfold analysis to find out the secondary structures of the key3c and the lock3c. I also started looking at the COSMIC code that is intended to model conjugation. So far I only downloaded the source codes, and still need to find out how to run it.
Calin
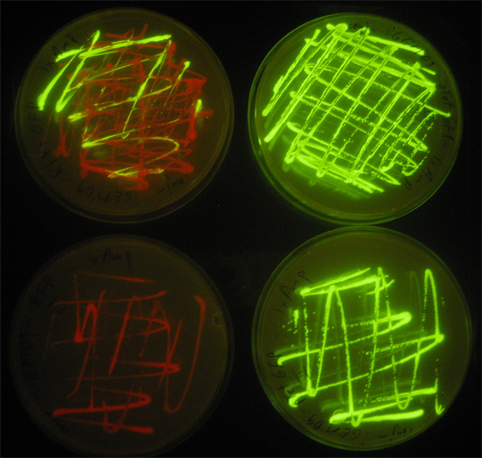
Updated Matlab code to fix mRNA / protein confusion. Got growth in all tubes and plates. MIT parts placed in fridge. Tube cultures went to Sriram's miniprep for plasmid extraction. Did some GFP / RFP non-quantitative measurements. These can be done in the gel imaging machine.
 "
"







