Team:Cambridge/Project/Amplification/Characterisation
From 2009.igem.org
(→Introduction) |
(→Introduction) |
||
| Line 13: | Line 13: | ||
== Introduction == | == Introduction == | ||
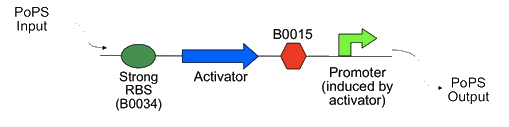
| - | The Cambridge 2007 iGEM team build 15 "amplifiers," constructs with RFP and GFP reporters that amplified the PoPS output of the promoter pBad (<partinfo>BBa_I0500</partinfo>, as described below: | + | The Cambridge 2007 iGEM team build 15 "amplifiers," constructs with RFP and GFP reporters that amplified the PoPS output of the promoter pBad/AraC(<partinfo>BBa_I0500</partinfo>, as described below: |
[[Image:amplifier07.jpg]] | [[Image:amplifier07.jpg]] | ||
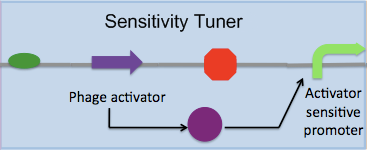
| - | We re-designed these constructs as follows... | + | We re-designed these constructs to be PoPS converters as follows... |
[[Image:thresholddevice3.jpg]] | [[Image:thresholddevice3.jpg]] | ||
| Line 51: | Line 51: | ||
|} | |} | ||
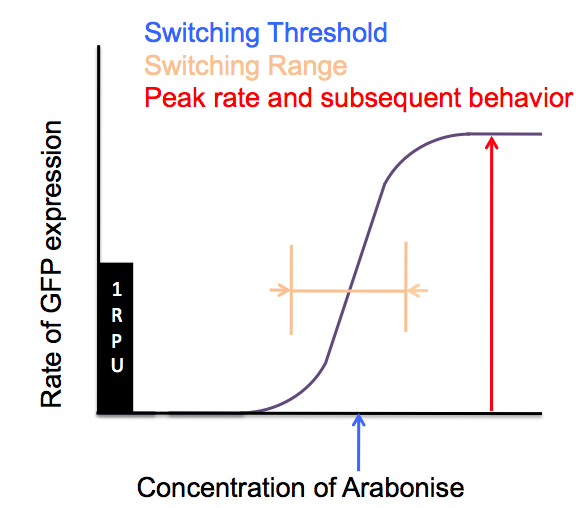
| - | In order to characterize these phage activator/promoter constructs, we used the corresponding Cambridge 2007 amplifier as an illustration of how our Sensitivity Tuners alter the behaviour of pBad. | + | In order to characterize these phage activator/promoter constructs, we used the corresponding Cambridge 2007 amplifier as an illustration of how our Sensitivity Tuners alter the behaviour of pBad/AraC. These parts are very useful for characterisation as they contain fluorescent reporters; the parts we designed, which lack an input promoter and fluorescent reporters, are more useful parts for other iGEM teams to incorporate into their own projects. For characterisation, we moved the Cambridge 2007 amplifiers onto a low copy plasmid in order to make meaningful comparisons with <partinfo>BBa_J69591</partinfo>, the standard promoter. We looked at four major characteristics relating input (arabinose) to output (GFP) and how they are modified compared to pBad/AraC on its own. |
| - | + | ||
| - | + | ||
| - | + | ||
[[Image:characterization1.jpg]] | [[Image:characterization1.jpg]] | ||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
== Recreating Previous Work == | == Recreating Previous Work == | ||
Revision as of 20:42, 21 October 2009
Categories :
Project :
-
Overview
Sensitivity Tuner
--- Characterisation
--- Modelling
Colour Generators
--- Carotenoids (Orange/Red)
--- Melanin (Brown)
--- Violacein (Purple/Green)
The Future
Safety
Notebook :
Team Logistics :
The Sensitivity Tuner
Introduction
The Cambridge 2007 iGEM team build 15 "amplifiers," constructs with RFP and GFP reporters that amplified the PoPS output of the promoter pBad/AraC(, as described below:
We re-designed these constructs to be PoPS converters as follows...
...and generated our own set of Sensitivity Tuners:
| P2 ogr activator | PSP3 pag activator | phiR73 delta activator | |
|---|---|---|---|
| PF promoter | |||
| PO promoter | |||
| PP promoter | |||
| Psid promoter | |||
| PLL promoter |
In order to characterize these phage activator/promoter constructs, we used the corresponding Cambridge 2007 amplifier as an illustration of how our Sensitivity Tuners alter the behaviour of pBad/AraC. These parts are very useful for characterisation as they contain fluorescent reporters; the parts we designed, which lack an input promoter and fluorescent reporters, are more useful parts for other iGEM teams to incorporate into their own projects. For characterisation, we moved the Cambridge 2007 amplifiers onto a low copy plasmid in order to make meaningful comparisons with , the standard promoter. We looked at four major characteristics relating input (arabinose) to output (GFP) and how they are modified compared to pBad/AraC on its own.
Recreating Previous Work
We began by recreating the 2007 team's data with some select amplifier constructs. We have the advantage over the 2007 team in that we have a better plate reader that is able to take OD600 absorbance readings at the same time as taking RFP and GFP output readings. For our transformations, we used the E. coli host strain BW27783. This host strain constitutively expresses arabinose transporters and is unable to metabolise arabinose, making it an ideal host for arabinose titration experiments.
Results: The 2007 team's hopes for future work included investigating a problem they attributed to the toxicity of high levels of activator in the cell. Overnight OD600 readings of cells transformed with their amplifier constructs indicated cell death. However, these OD readings were conducted separately from their RFP and GFP output measurements. The 2009 team gathered data on the plate reader capable of taking OD600 absorbance readings as well as RFP and GFP output readings; no OD600 readings suggested cell death due to toxicity.
- graphs*
Characterisation
We moved all 15 activator constructs onto pSB3K3, a low copy plasmid. The standard promoter for 1 RPU, J69591, is also on pSB3K3 and has a GFP reporter, so we can make meaningful comparisons on the plate reader.
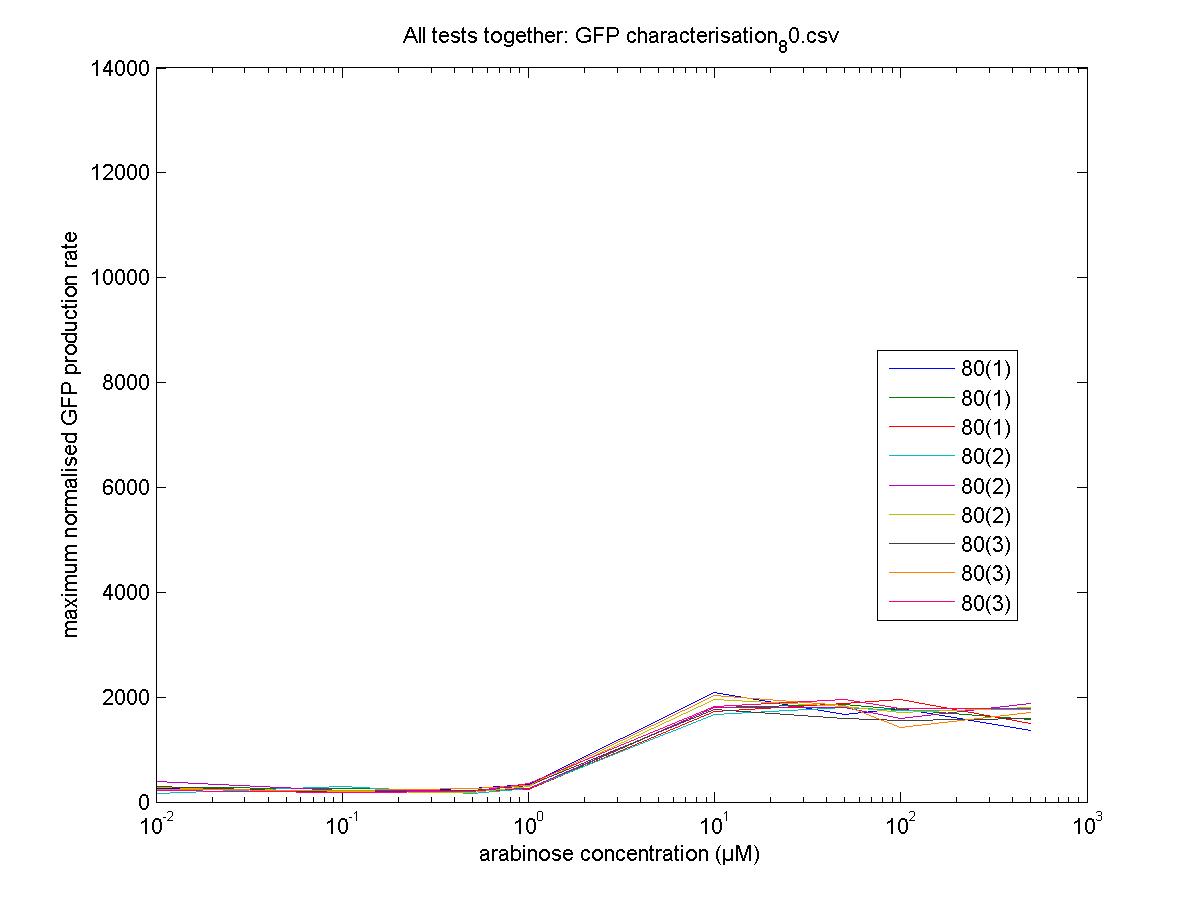
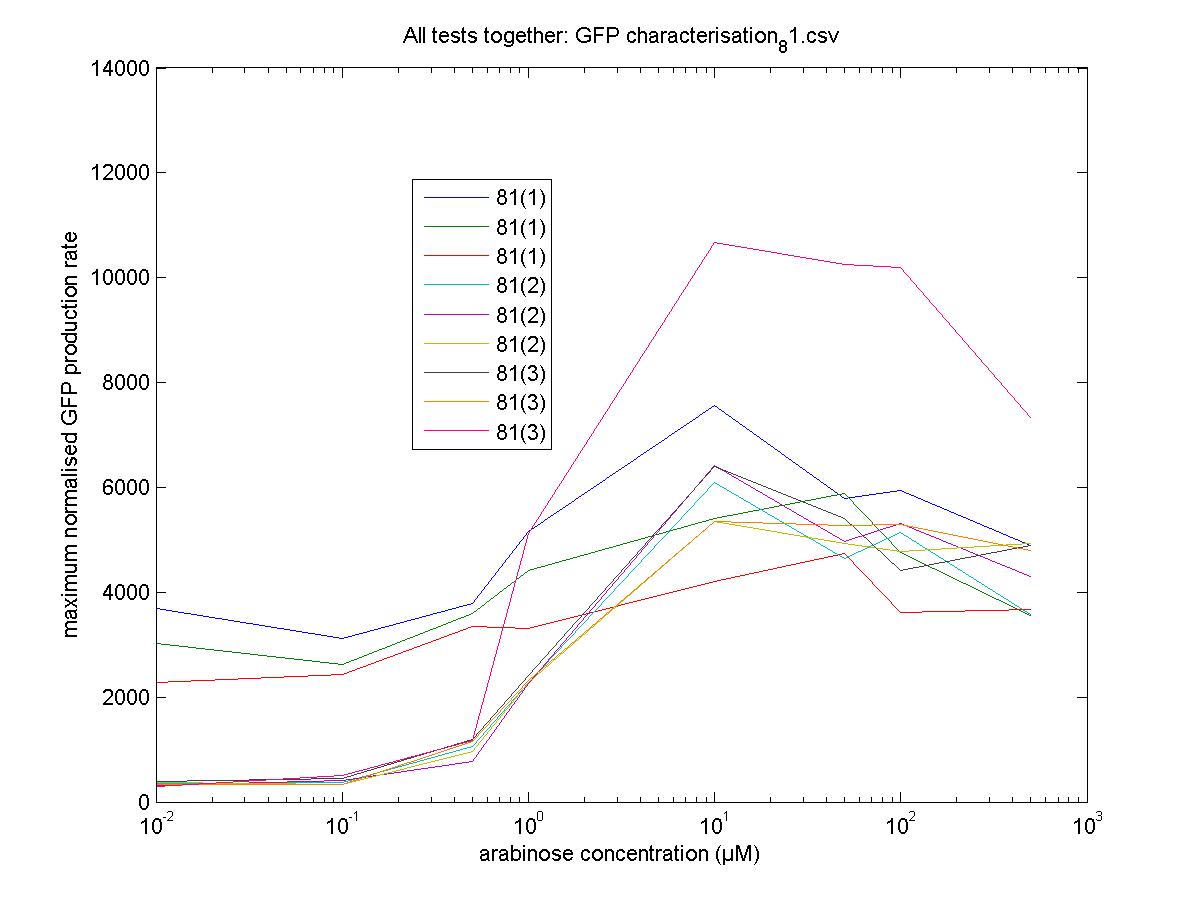
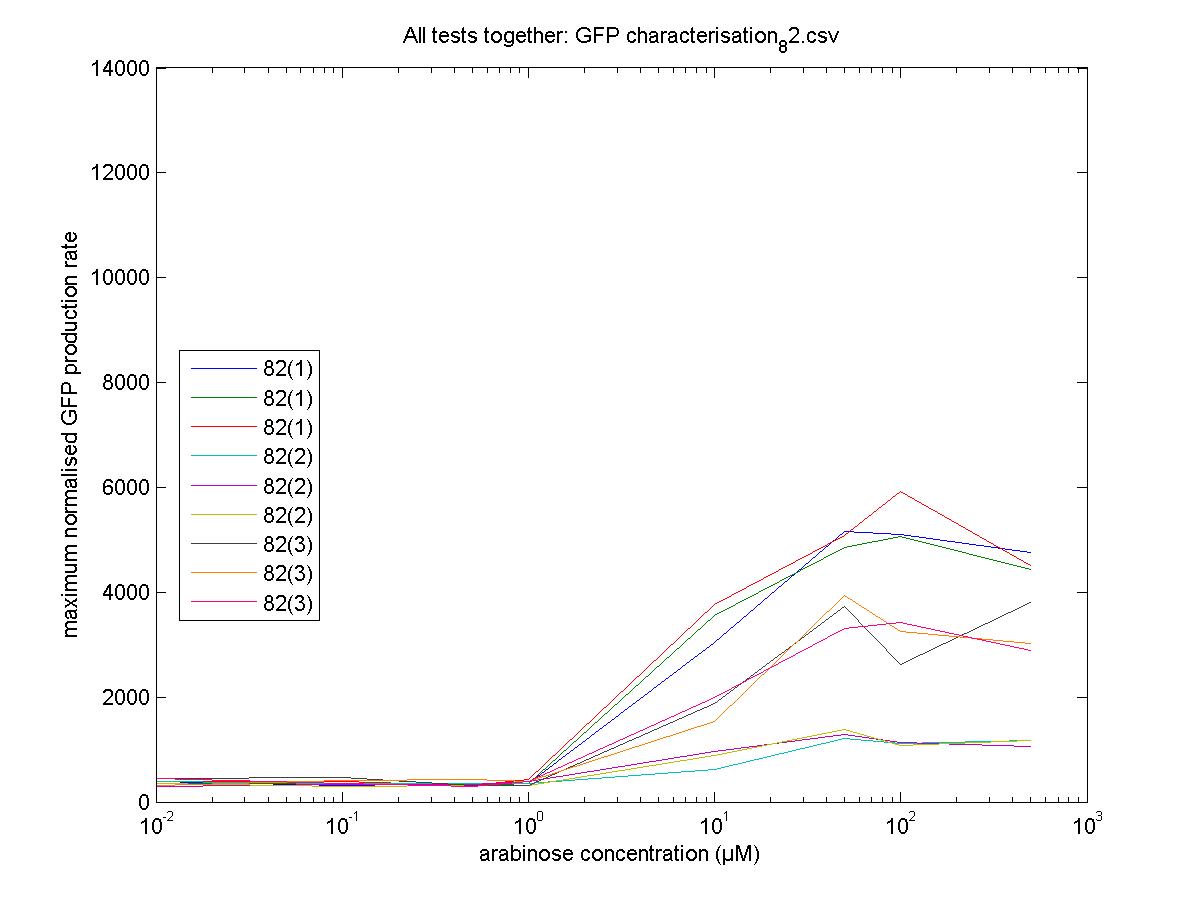
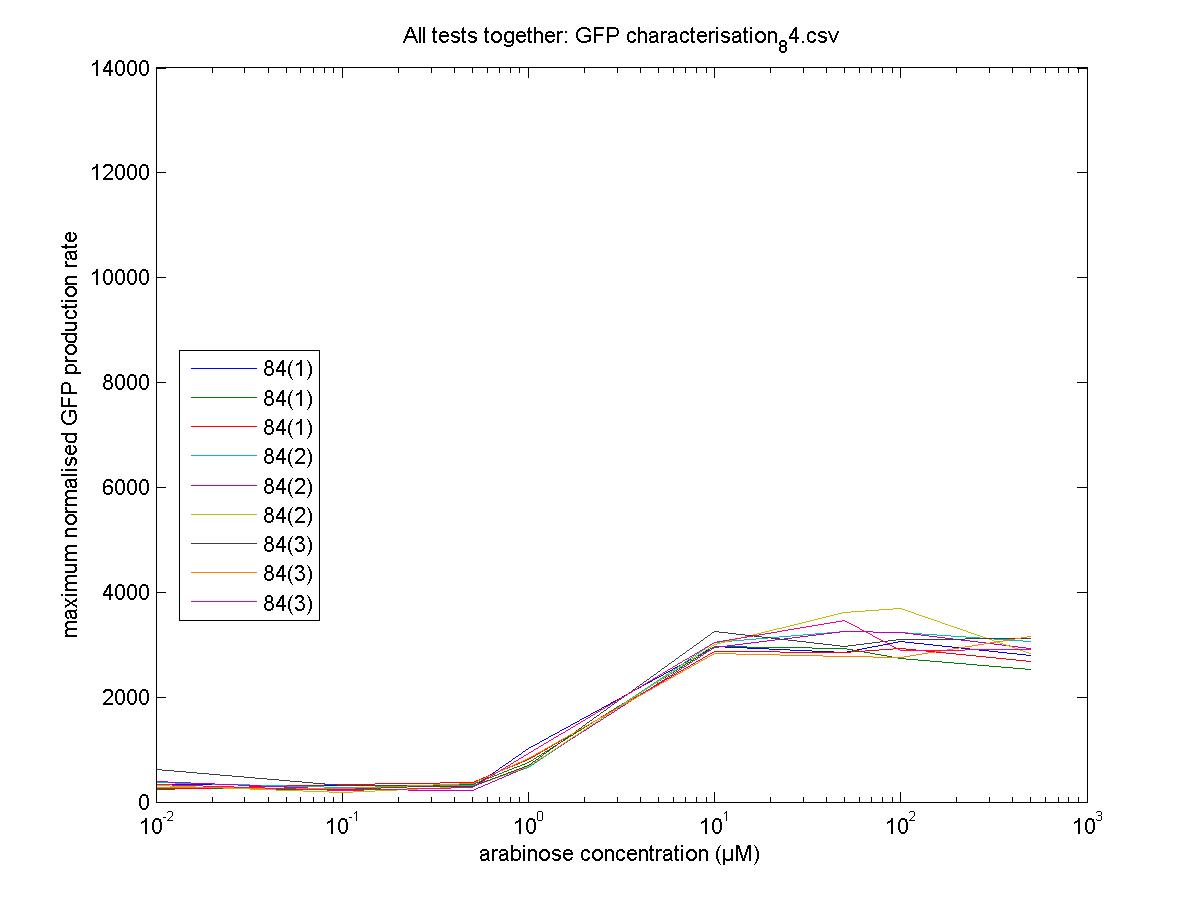
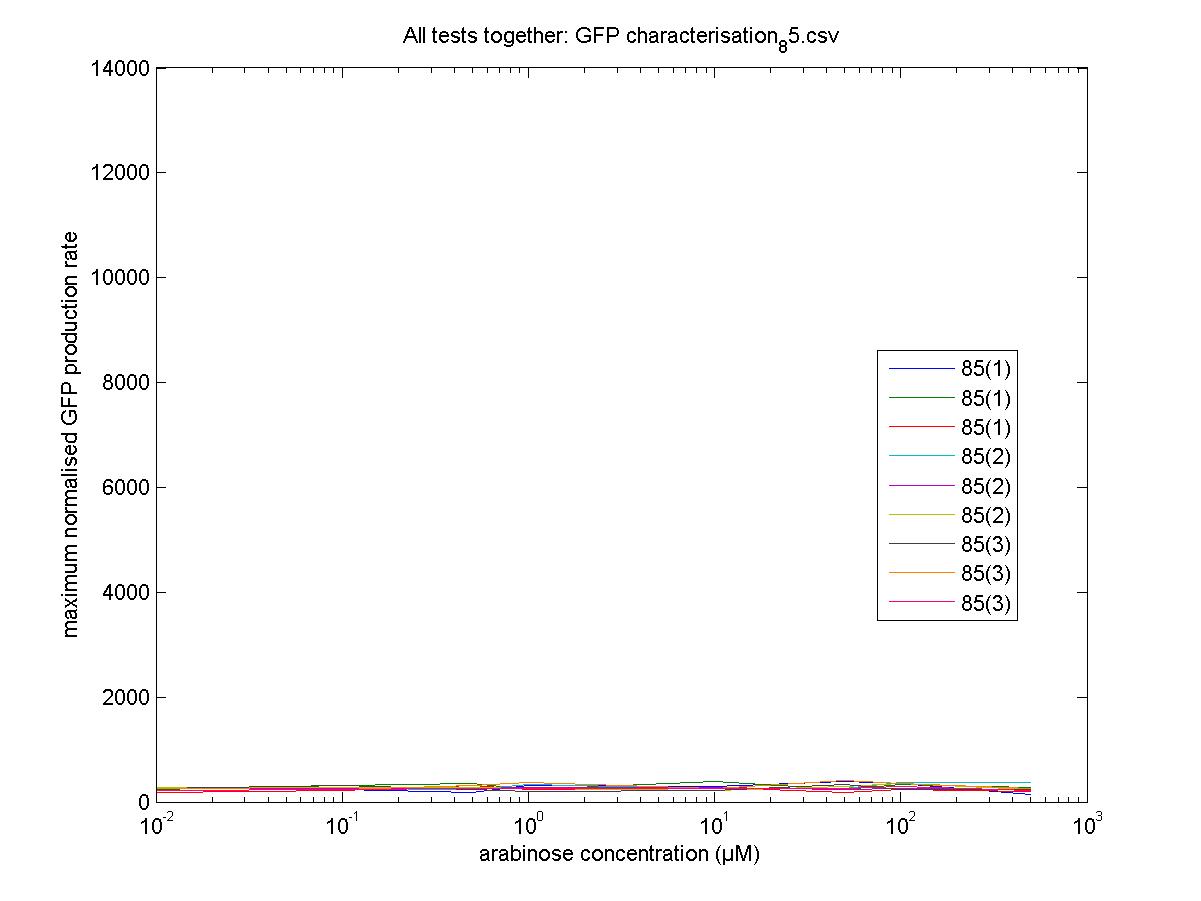
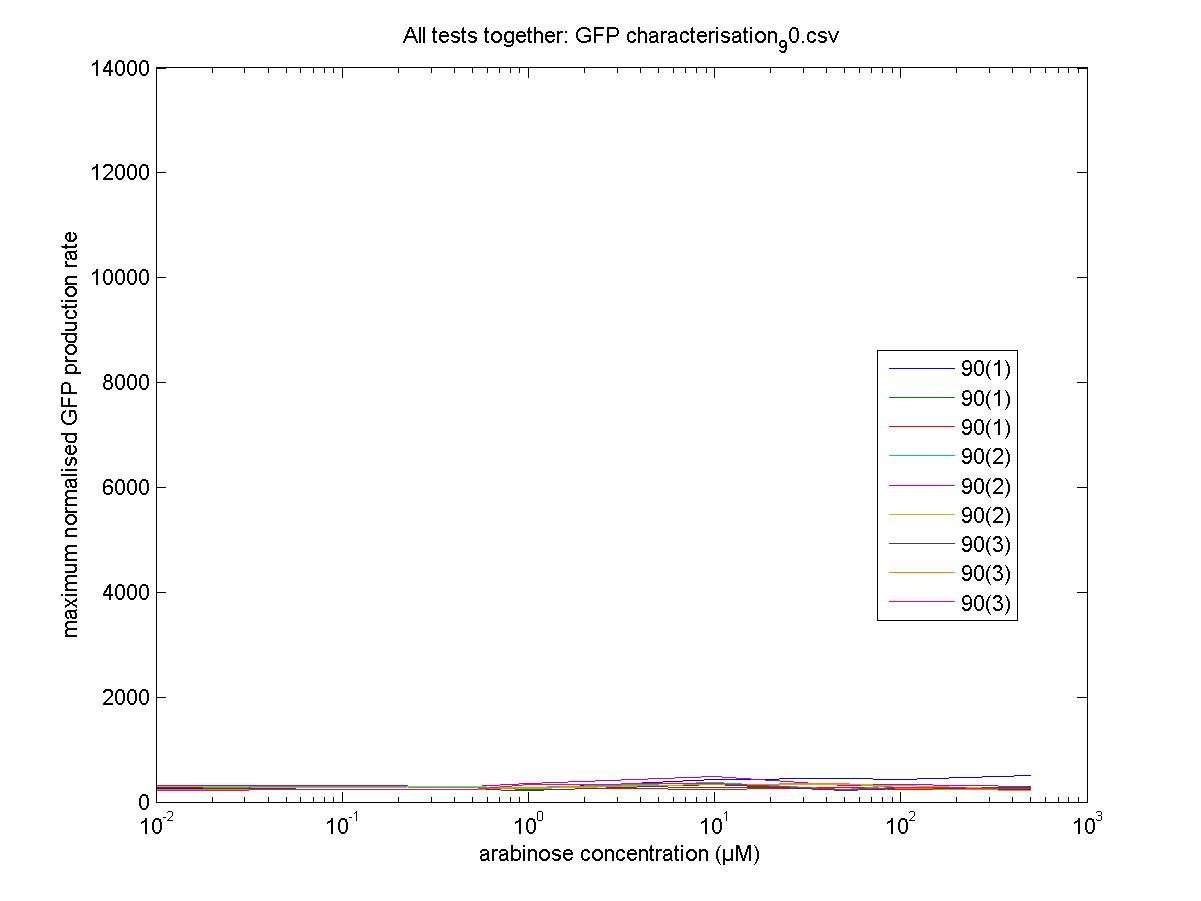
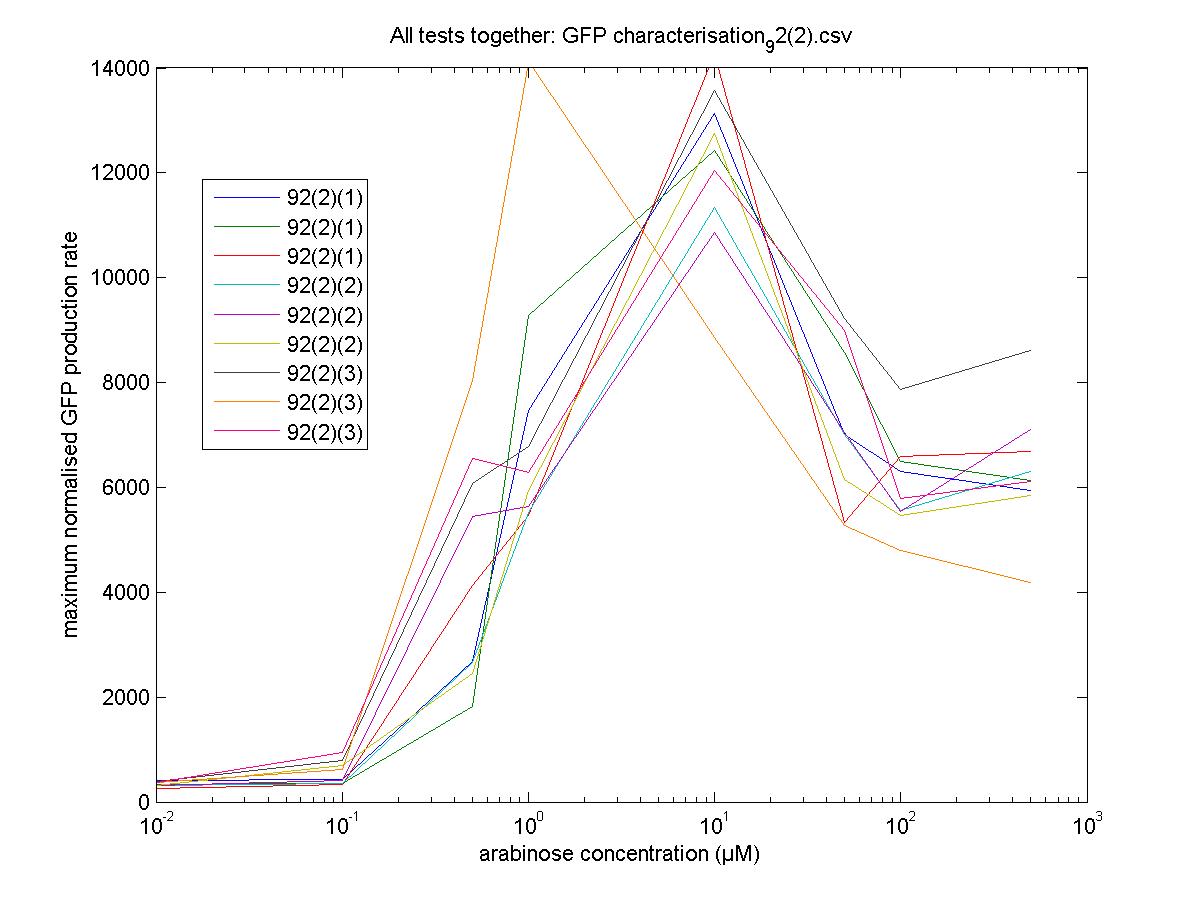
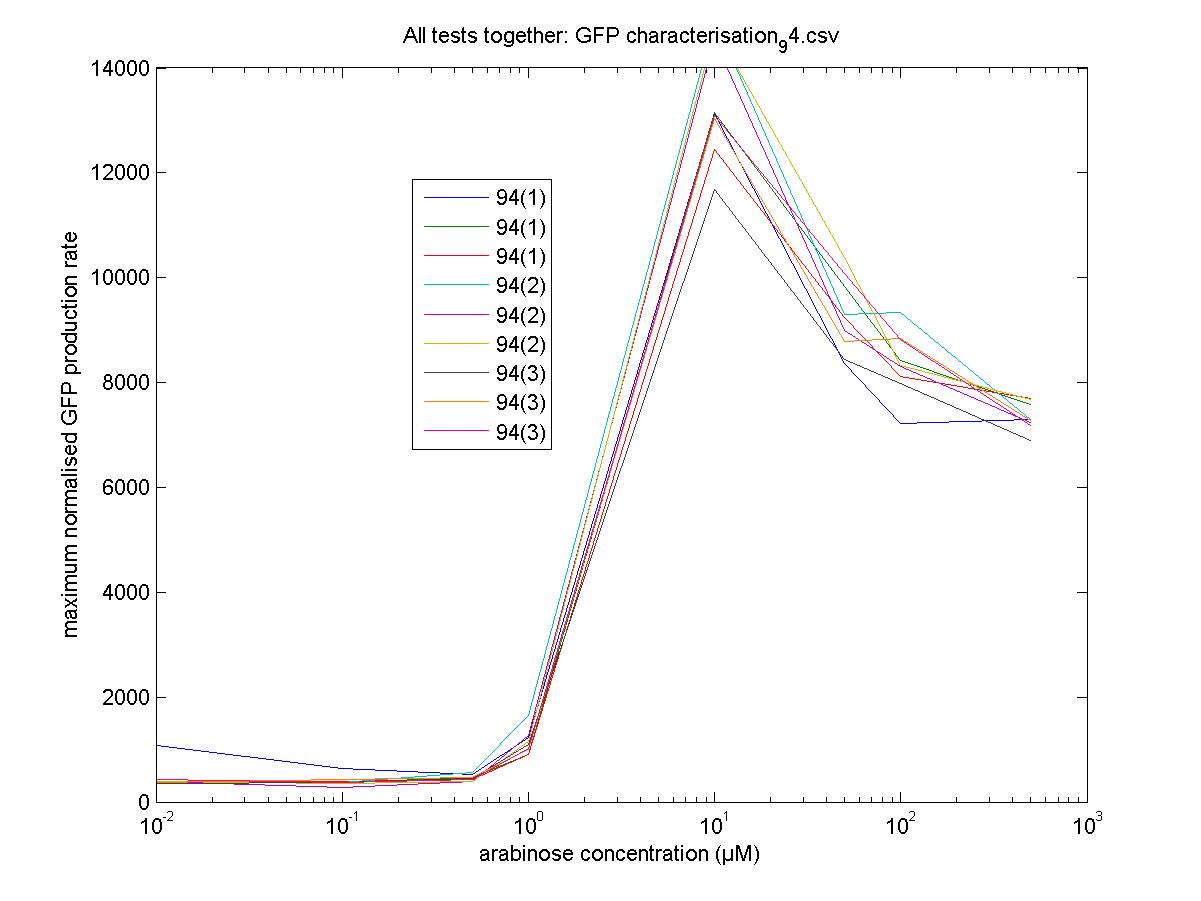
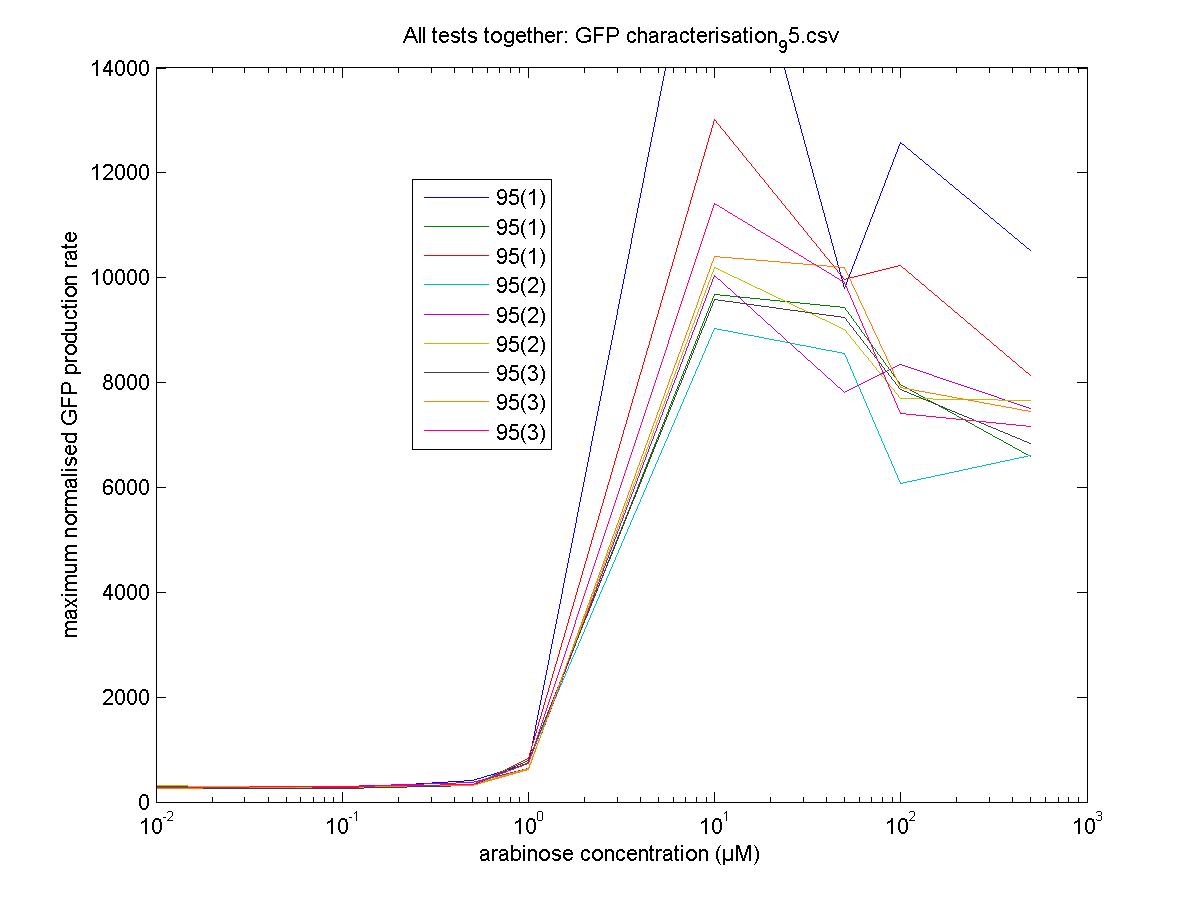
Maximum Rates against Arabinose Concentrations
80
81
82
84
85
90
92
94
95
Reconstruction
Below is a table listing the sensitivity tuners as they appear in the registry as biobricks:
 "
"