Team:Paris/Production modeling
From 2009.igem.org
(→B. Modeling Vesicles creation.) |
Christophe.R (Talk | contribs) (→A more flexible design) |
||
| (224 intermediate revisions not shown) | |||
| Line 1: | Line 1: | ||
| - | <span/ id="bottom">[https://2009.igem.org/ iGEM ] > [[Team:Paris#top | Paris]] > [[Team:Paris/ | + | <span/ id="bottom">[https://2009.igem.org/ iGEM ] > [[Team:Paris#top | Paris]] > [[Team:Paris/DryLab#bottom | DryLab]] > [[Team:Paris/Production_modeling#bottom | Vesicle production Model (delay model)]] |
{{Template:Paris2009}} | {{Template:Paris2009}} | ||
{{Template:Paris2009_menu2}} | {{Template:Paris2009_menu2}} | ||
| - | == | + | |
| - | + | ||
| + | |||
| + | |||
| + | |||
| + | |||
| + | == DryLab - Delay model: Improving message quality== | ||
| + | |||
<html> | <html> | ||
| + | <style type="text/css"> | ||
| + | #left-side { | ||
| + | position: absolute; | ||
| + | height: 23px; | ||
| + | width: 30px; | ||
| + | top: 0px; | ||
| + | left: 120px; | ||
| + | margin-top:10px; | ||
| + | padding-top: 7px; | ||
| + | background: url(https://static.igem.org/mediawiki/2009/1/1b/Left_menu_pari.png); | ||
| + | z-index:4; | ||
| + | } | ||
| + | |||
| + | #middle-side { | ||
| + | height: 25px; | ||
| + | width: 400px; | ||
| + | position: absolute; | ||
| + | top: 0px; | ||
| + | left: 130px; | ||
| + | margin-top:10px; | ||
| + | padding-top: 5px; | ||
| + | background: #dadada; | ||
| + | z-index:5; | ||
| + | } | ||
| + | |||
| + | #right-side { | ||
| + | position: absolute; | ||
| + | height: 23px; | ||
| + | width: 30px; | ||
| + | margin-top:10px; | ||
| + | padding-top: 7px; | ||
| + | top: 0px; | ||
| + | left: 520px; | ||
| + | background: url(https://static.igem.org/mediawiki/2009/4/40/Right_menu_paris.png); | ||
| + | z-index:4; | ||
| + | } | ||
| + | #left-side2 { | ||
| + | position: absolute; | ||
| + | height: 23px; | ||
| + | width: 30px; | ||
| + | top: 35px; | ||
| + | left: 110px; | ||
| + | margin-top:10px; | ||
| + | padding-top: 7px; | ||
| + | background: url(https://static.igem.org/mediawiki/2009/4/40/Left_menu_paris2.png); | ||
| + | z-index:4; | ||
| + | } | ||
| + | |||
| + | #middle-side2 { | ||
| + | height: 25px; | ||
| + | width: 430px; | ||
| + | position: absolute; | ||
| + | top: 35px; | ||
| + | left: 120px; | ||
| + | margin-top:10px; | ||
| + | padding-top: 5px; | ||
| + | background: white; | ||
| + | z-index:5; | ||
| + | } | ||
| + | |||
| + | #right-side2 { | ||
| + | position: absolute; | ||
| + | height: 23px; | ||
| + | width: 30px; | ||
| + | margin-top:10px; | ||
| + | padding-top: 7px; | ||
| + | top: 35px; | ||
| + | left: 530px; | ||
| + | background: url(https://static.igem.org/mediawiki/2009/f/f8/Right_menu_paris2.png); | ||
| + | z-index:4; | ||
| + | } | ||
| + | |||
| + | a.menu_sub { | ||
| + | padding-left: 7px; | ||
| + | padding-right: 7px; | ||
| + | } | ||
| + | |||
| + | a.menu_sub_active { | ||
| + | padding-left: 7px; | ||
| + | padding-right: 7px; | ||
| + | color:#b0310e; | ||
| + | font-weight:bold; | ||
| + | } | ||
| + | </style> | ||
| + | <div id="left-side"></div> | ||
| + | <div id="middle-side"><center> | ||
| + | <a class="menu_sub"href="https://2009.igem.org/Team:Paris/DryLab#bottom"> Main </a>| | ||
| + | <a class="menu_sub_active"href="https://2009.igem.org/Team:Paris/Production_modeling#bottom"> Delay model</a>| | ||
| + | <a class="menu_sub"href="https://2009.igem.org/Team:Paris/Production_modeling2#bottom"> Vesicle model</a>| | ||
| + | <a class="menu_sub"href="https://2009.igem.org/Team:Paris/Transduction_modeling#bottom"> Fec simulation</a> | ||
| + | </center> | ||
</div> | </div> | ||
| - | <div id=" | + | <div id="right-side"></div> |
| + | <div id="left-side2"></div> | ||
| + | <div id="middle-side2"><center> | ||
| + | <a class="menu_sub_active"href="https://2009.igem.org/Team:Paris/Production_modeling#bottom"> Introduction </a>| | ||
| + | <a class="menu_sub"href="https://2009.igem.org/Team:Paris/Production_modeling#Implementing_the_delay_by_a_transcriptional_cascade"> Transcriptional Cascade</a>| | ||
| + | <a class="menu_sub"href="https://2009.igem.org/Team:Paris/Production_modeling#Implementing_the_delay_by_feed-forward_motifs"> The feed-forward motif</a> | ||
| + | </center> | ||
</div> | </div> | ||
| - | <div id=" | + | <div id="right-side2"></div> |
</html> | </html> | ||
| - | |||
| - | + | This part aimed at solving the first problem linked to the creation of messengers : | |
| + | |||
| + | |||
| + | |||
| + | <center>'''To improve the quality of signal sent, how can we get a good synchronization between the maximal vesicles production rate and the maximal concentration of proteins to encapsulate'''</center> | ||
| + | |||
| + | |||
| + | <span/ id="1"><span/ id="2"> | ||
| + | Obviously, it is crucial to send vesicles only when the concentration of proteins to encapsulate is at its maximum level in the periplasm. To optimize the quality of our message, we tried to make this peak of vesicles creation corresponding to maximum levels of protein concentration. We assumed that the number of vesicles produced is at its maximum when the concentration of TolRII has reached its peak. | ||
| + | |||
| + | |||
| + | In what follows, we consider two different designs for implementing the delay: using a transcriptional cascade and using feed-forward motifs. | ||
| + | We found that the '''design based on transcriptional cascade is sufficient for obtaining the desired delay'''<sup>[[https://2009.igem.org/Team:Paris/Production_modeling#References 1]]</sup>, whereas the '''design based on feed-forward motifs allows for more flexibility, at the cost of an increased complexity'''<sup>[[https://2009.igem.org/Team:Paris/Production_modeling#References 2]]</sup>. | ||
| + | These studies are described below. | ||
| + | |||
| + | |||
| + | ===Implementing the delay by a transcriptional cascade=== | ||
| + | <span/ id="3"><span/ id="4"> | ||
| + | |||
| + | We have decided to construct the following system. | ||
| + | The addition of arabinose to the medium leads to the activation of the pBad promoters: two proteins are produced: the protein to encapsulate and LacI-LVA<sup>[[https://2009.igem.org/Team:Paris/Production_modeling#References 3]]</sup><sup>[[https://2009.igem.org/Team:Paris/Production_modeling#References 4]]</sup>. The latter inhibits the expression of TetR and thus releases the inhibition of TolRII, responsible for vesicle formation. | ||
| - | + | [[Image:Global On.jpg|600px|center]] | |
| + | <span/ id="5"> | ||
| + | To study this system, we simply used differential equations. We used the method described by U. Alon in his book ''An Introduction to Systems Biology'', with notably Hill functions to describe the regulation of the promoters<sup>[[https://2009.igem.org/Team:Paris/Production_modeling#References 5]]</sup>. | ||
| + | The network behavior can be described by a system of differential equations: | ||
<center> | <center> | ||
| Line 29: | Line 154: | ||
|- style="background: #0d3e99; text-align: center; color:white;" | |- style="background: #0d3e99; text-align: center; color:white;" | ||
| - | |[[Image:Genetic network equations.png| | + | |[[Image:Genetic network equations.png|400px| Modeling Overview]] |
|} | |} | ||
</center> | </center> | ||
| - | + | <html> | |
| - | + | <div style="width: 400px; | |
| + | text-align: justify; | ||
| + | margin-left:200px;"> | ||
| + | These equations describe the evolution of the concentrations of the messenger protein P, LacI repressor, TetR repressor, and the TolRII protein responsible for vesiculation. The gammas are the degradation rates of the respective compounds, while the sigmoidal functions describe the non-linear production rates. The parameters are estimated according to typical values as discussed below</div> | ||
| + | </html> | ||
As a first approximation, we assumed that : | As a first approximation, we assumed that : | ||
| - | *When the pBad promoter is induced, the concentration of arabinose in the medium is very high and constant during he whole study ; as a consequence, we will consider that the creation rate of Protein and LacI* is constant during the experiment : | + | *When the pBad promoter is induced, the concentration of arabinose in the medium is very high and constant during he whole study; as a consequence, we will consider that the creation rate of Protein P and LacI* is constant during the experiment: |
<center> | <center> | ||
| Line 44: | Line 173: | ||
|- style="background: #0d3e99; text-align: center; color:white;" | |- style="background: #0d3e99; text-align: center; color:white;" | ||
| - | |[[Image:Constant rate copie.png| | + | |[[Image:Constant rate copie.png|360px|center| Arabinose constant]] |
|} | |} | ||
</center> | </center> | ||
| - | + | <span/ id="6"> | |
| - | *We considered that all the binding constants are identical and of an average of 40nM which correspond to approximately 40 monomers per cell ; we can write that : | + | *We considered that all the binding constants are identical<sup>[[https://2009.igem.org/Team:Paris/Production_modeling#References 6]]</sup> and of an average of 40nM which correspond to approximately 40 monomers per cell; we can write that: |
<center> | <center> | ||
| Line 56: | Line 185: | ||
|- style="background: #0d3e99; text-align: center; color:white;" | |- style="background: #0d3e99; text-align: center; color:white;" | ||
| - | |[[Image: | + | |[[Image:K equals.jpg|175px|center| Binding constants]] |
|} | |} | ||
| Line 63: | Line 192: | ||
| - | *We chose identical | + | *We chose identical intrinsic promoter activities, β, all equal to 4000 proteins/cell cycle: |
<center> | <center> | ||
| Line 77: | Line 206: | ||
| - | *All the time units were expressed in units of cell cycle (approximately half an hour) | + | *All the time units were expressed in units of cell cycle (approximately half an hour). As a consequence we chose a dilution rate γ of 1 for protein without special tags. For the LacI protein with a degradation tag (LacI-LVA), the dilution rate is 3 times higher<sup>[[https://2009.igem.org/Team:Paris/Production_modeling#References 3]]</sup>: |
<center> | <center> | ||
| Line 83: | Line 212: | ||
|- style="background: #0d3e99; text-align: center; color:white;" | |- style="background: #0d3e99; text-align: center; color:white;" | ||
| - | |[[Image:Dilution One.png| | + | |[[Image:Dilution One.png|200px|center| Binding constants]] |
|} | |} | ||
| Line 92: | Line 221: | ||
|- style="background: #0d3e99; text-align: center; color:white;" | |- style="background: #0d3e99; text-align: center; color:white;" | ||
| - | |[[Image:Dilution LVA.png| | + | |[[Image:Dilution LVA.png|80px|center| Binding constants]] |
|} | |} | ||
</center> | </center> | ||
| - | |||
| - | + | For simulation, we used the following initial quantities: | |
| + | *4000 units of TetR (steady state level at inactivation) | ||
| + | *no protein to be encapsulated | ||
| + | *no LacI | ||
| + | *no TolRII | ||
| - | + | We got the following dynamics using Matlab software, showing that '''this simple system is capable of generating the desired behavior'''. | |
| - | : | + | [[Image:Delay System.jpg|800px|center]] |
| - | + | <center> | |
| + | Plot showing that vesiculation happens only once the protein encapsulated has reached its maximal concentration. | ||
| + | </center> | ||
| - | |||
| + | ===Implementing the delay by feed-forward motifs=== | ||
| - | |||
| - | |||
| - | + | Using the previous design, we were able to find a way to get the maximal vesicle formation (ie maximal TolRII quantity) when the concentration of proteins to encapsulate is at its maximum. This system satisfies our requirements. | |
| - | + | However, it is not very flexible. In particular, the concentration of the protein to encapsulate remains always high after induction. This could be detrimental for the cell if this protein is toxic. Consequently, we '''considered an alternative design allowing for more flexibility''', in which the expression of the protein to encapsulate can be '''transient'''. | |
| - | |||
| - | |||
| + | ====The Feed Forward Motif==== | ||
| + | <span/ id="7"> | ||
| - | + | The feed forward motif was studied and described by U. Alon in his book "An Introduction to Synthetic Biology" where he describes the different forms of existing feed forward; they can be either coherent or incoherent<sup>[[https://2009.igem.org/Team:Paris/Production_modeling#References 7]]</sup>. | |
| - | + | In each case, there exist a different network organization to create a feed-forward loop as shown on the picture | |
| - | + | ||
| + | [[Image:FFd global.jpg|250px|left]] | ||
| - | + | The X and Y inputs can be combined either using a logical AND or OR gate thus giving different dynamics properties in each case. | |
| - | + | The coherent feed forward loop allow the creation of a delay either on activation or on deactivation depending on the logical function between the two input X and Y. | |
| - | + | When the output logical gate is an AND function, the delay appears on activation, while when the output gate is an OR function, the delay appears on deactivation. | |
| + | An example of characterized feed-forward loop is the fliA system which was used by the [https://2008.igem.org/Team:Paris| 2008 Paris team] in order to create a oscillator. | ||
| + | |||
| + | In an incoherent feed forward loop, the two inputs of the last gate function have two different logical levels; one molecule (molecule X for example) is an activator while the other molecule is a repressor of the promoter controlling the expression of the output. | ||
| + | This way, when the input signal is on, the expression of molecule X starts activating both the promoters controlling the expression of Y and Z ; transcription starts and the amount of Y and Z molecules increases. Nevertheless, when the concentration of Y has reached its repression threshold, repression starts and the concentration of Z molecule decrease to its steady state. | ||
| + | |||
| + | |||
| + | |||
| + | |||
| + | [[Image:Double IFFD.jpg|450px|right]] | ||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | Two elementary incoherent feed forward motifs can be combined in order to create two time delayed bursts as described by U.Alon in its description of the development of ''Bacillus subtilis'' spore. Anyway, it seems quite difficult to find a molecule which would be both an activator and a repressor; we decided to try and think about another combination of these two feed forwards so as to get the dynamics we wanted as shown on the following part. | ||
| + | |||
| + | ====Our design proposition==== | ||
| + | |||
| + | |||
| + | The two feed-forward loops used are incoherent and organized in a way that each molecule is either an activator or a repressor (and not both). The following scheme gives the functioning of the system. | ||
| + | |||
| + | [[Image:Feed forward.png|300px|center]] | ||
| + | |||
| + | |||
| + | |||
| + | {| | ||
|- style="background: #0d3e99; text-align: center; color:white;" | |- style="background: #0d3e99; text-align: center; color:white;" | ||
| - | | | + | |width=10%| Steps |
| + | |width=70%| Description | ||
| + | |width=20%| Changes in concentration | ||
| + | |||
| + | |- style="background: #bebebe; text-align: center; color:black;" | ||
| + | |1 | ||
| + | |When the input signal is ON, molecule1 is created and activates the promoter upstream the coding sequence for molecules Z2 and Z3 as well as Z4 since there is no molecule Z3 in the medium (see the AND gate function) at the beginning ; there is an initial amount of Z5 molecule but no Z2 so the no Z6 molecule is produced. | ||
| + | |Z1 +++ | ||
| + | Z2+ | ||
| + | Z3+ | ||
| + | Z4++ | ||
| + | Z5~ | ||
| + | Z6~ | ||
| + | |||
| + | |- style="background: #d8d8d8; text-align: center; color:black;" | ||
| + | |2 | ||
| + | |After a while, there is enough Z3 to start the repression of Z4 promoter and there is also enough Z2 to start producing Z6 signficantly. | ||
| + | |Z1+++ | ||
| + | Z2+++ | ||
| + | Z3+++ | ||
| + | Z4- | ||
| + | Z5~ | ||
| + | Z6+ | ||
| + | |||
| + | |- style="background: #bebebe; text-align: center; color:black;" | ||
| + | |3 | ||
| + | |Then, because of the repression of Z3, the Z4 molecule reaches its steady level ; in the same time, Z3 starts repressing the production of Z5 and the amount of Z6 starts decreasing. | ||
| + | |Z1+ | ||
| + | Z2+ | ||
| + | Z3+ | ||
| + | Z4~ | ||
| + | Z5- | ||
| + | Z6+ | ||
| + | |||
| + | |- style="background: #d8d8d8; text-align: center; color:black;" | ||
| + | |4 | ||
| + | |Finally, Z3 represses the promoter allowing the creation of Z5 and every molecules reaches its steady level concentration. | ||
| + | |Z1~ | ||
| + | Z2~ | ||
| + | Z3~ | ||
| + | Z4~ | ||
| + | Z5- | ||
| + | Z6~ | ||
|} | |} | ||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| + | To study the dynamics of our model, we decided to use a classical approach using Michaelis-Menten equations to model the activation and repression of the various promoters, including a cooperativity of order 2 for the repressor which often the case for 'classical' repressors such as LacI. We obtained the following system of equations: | ||
| - | |||
| - | |||
| + | [[Image:Equation feed.png|300px|center]] | ||
| - | |||
| - | |||
| - | |||
| - | + | It is possible to obtain a broad spectrum of different dynamical behaviors by playing with the value of the maximum production rate of Z3, called here β . This maximal production rate impacts deeply both the evolution of the exported protein concentration and the evolution of TolRII concentration. This rate is initially chosen equal to 40 and 400 molecules by cell cycle and we have plotted the different responses for multiples of these initial values. | |
| + | In each case, the initial conditions are the same : | ||
| + | *no proteins encapsulated | ||
| + | *no TolRII | ||
| + | *no Z1 molecules | ||
| + | *no Z2 molecules | ||
| + | *no Z3 molecules | ||
| + | *a weak amount of Z4 molecules (around a hundred) | ||
| + | |||
| + | |||
| + | <center>Weak maximal Z3 production rate</center> | ||
| + | <center>''An example of dynamics for a range of weak β values; for each plot, the maximal Z3 production rate is given by: β=k*40''</center> | ||
| + | [[Image:Simu ff.png|650px|center]] | ||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | <center>Strong maximal Z3 production rate</center> | ||
| + | <center>''An example of dynamics for a range of strong β values; for each plot, the maximal Z3 production rate is given by: β=k*400''</center> | ||
| + | [[Image:Feed forward bis.png|650px|center]] | ||
| + | |||
| + | ====A more flexible design==== | ||
| + | |||
| + | |||
| + | Regardless of the steady state concentrations of the molecules, the use of two incoherent feed-forward loops allows for '''more flexibility in the behaviors of our system'''. With a simple delay system, we just get one possible dynamic behavior whereas using a variable range of production rates, we can obtain two behaviors: | ||
| + | |||
| + | |||
| + | |||
| + | *When β is weak (β = 40), the number of vesicles (ie of TolRII) produced slowly increase before returning to zero, whereas the protein to be exported inside the vesicles is created very quickly and remains at a high concentration compared to the TolRII protein. We can therefore send vesicles containing an important amount of interesting proteins and, after a while, stop expressing dangerous proteins for the cell. All of this can be performed with a single input signal. | ||
| + | |||
| + | |||
| + | *When β is stronger, the behavior is different and we notice a change of concavity of the TolRII curve when the amount of protein exported is at its maximum. During such a change of concavity, the derivative reaches its maximum - in other words, the amount of TolRII produced by time units is at its highest level. Considering the hypothesis according to which the production rate of vesicles is proportional to TolRII production rate, we get a peak in vesicle production when the concentration of the proteins to exported is at its maximum. | ||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | Unfortunately, we did not have the time to try to construct this system in the lab and only the first design was implemented. Nevertheless, we thought that '''this analysis was very useful to get a good insight of the diversity of the possible behaviors of the sender cells'''. | ||
| + | |||
| + | |||
| + | {{Template:Paris2009_guided|Production_overview|Production_modeling2}} | ||
| + | |||
| + | <html> | ||
| + | </div> | ||
| + | <div id="paris_content_boxtop"> | ||
| + | </div> | ||
| + | <div id="paris_content"> | ||
| + | </html> | ||
| - | + | ====References==== | |
| - | + | <ol class="References"> | |
| + | <li> [[Team:Paris/Production_modeling#1 | ^]]Ultrasensitivity and noise propagation in a synthetic transcriptional cascade S.Hooshangi & R.Weiss | ||
| + | 2005 - [http://www.ncbi.nlm.nih.gov/pubmed/15738412 15738412]</li> | ||
| + | <li> [[Team:Paris/Production_modeling#1 | ^]]Network motifs : theory and experimental approaches U.Alon | ||
| + | 2007 - [http://www.ncbi.nlm.nih.gov/pubmed/17510665 117510665]</li> | ||
| + | <li> [[Team:Paris/Production_modeling#3 | ^]]New Unstable Variants of Green Fluorescent Protein for Studies of Transient Gene Expression in Bacteria Jens Bo Andersen & Søren Molin 1998 - [http://www.ncbi.nlm.nih.gov:80/pmc/articles/PMC106306/ 106306]</li> | ||
| + | <li> [[Team:Paris/Production_modeling#4 | ^]]Spatiotemporal control of gene expression with pulse-generating networks S.Basu & R.Weiss | ||
| + | 2003 - [http://www.ncbi.nlm.nih.gov/pubmed/15096621 15096621]</li> | ||
| + | <li> [[Team:Paris/Production_modeling#5 | ^]]Negative Autoregulation Speeds The Response Times of Transcription Network N.Rosenfold & U.Alon | ||
| + | 2002 - [http://www.ncbi.nlm.nih.gov/pubmed/12417193 2417193]</li> | ||
| + | <li> [[Team:Paris/Production_modeling#6 | ^]]A Synthetic oscillatory network of transcriptionnal regulators M.Ellowitz & S.Leibler 1999 - [http://www.ncbi.nlm.nih.gov/pubmed/10659856 10659856]</li> | ||
| + | <li> [[Team:Paris/Production_modeling#7 | ^]]Structure and function ot the feed-forward Loop Network Motif S.Mangan & U.Alon | ||
| + | 2003 - [http://www.ncbi.nlm.nih.gov/pubmed/14530388 14530388]</li> | ||
| + | </ol> | ||
Latest revision as of 23:54, 21 October 2009
iGEM > Paris > DryLab > Vesicle production Model (delay model)
Contents |
DryLab - Delay model: Improving message quality
This part aimed at solving the first problem linked to the creation of messengers :
Obviously, it is crucial to send vesicles only when the concentration of proteins to encapsulate is at its maximum level in the periplasm. To optimize the quality of our message, we tried to make this peak of vesicles creation corresponding to maximum levels of protein concentration. We assumed that the number of vesicles produced is at its maximum when the concentration of TolRII has reached its peak.
In what follows, we consider two different designs for implementing the delay: using a transcriptional cascade and using feed-forward motifs.
We found that the design based on transcriptional cascade is sufficient for obtaining the desired delay[1], whereas the design based on feed-forward motifs allows for more flexibility, at the cost of an increased complexity[2].
These studies are described below.
Implementing the delay by a transcriptional cascade
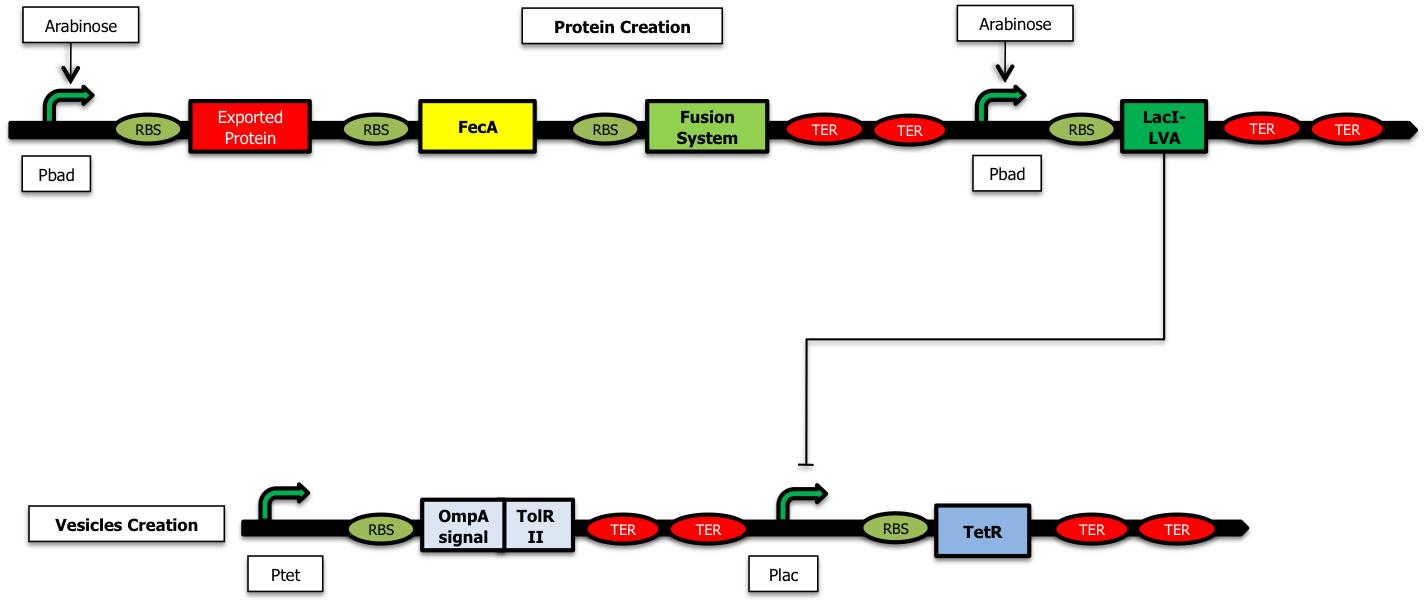
We have decided to construct the following system. The addition of arabinose to the medium leads to the activation of the pBad promoters: two proteins are produced: the protein to encapsulate and LacI-LVA[3][4]. The latter inhibits the expression of TetR and thus releases the inhibition of TolRII, responsible for vesicle formation.
To study this system, we simply used differential equations. We used the method described by U. Alon in his book An Introduction to Systems Biology, with notably Hill functions to describe the regulation of the promoters[5].
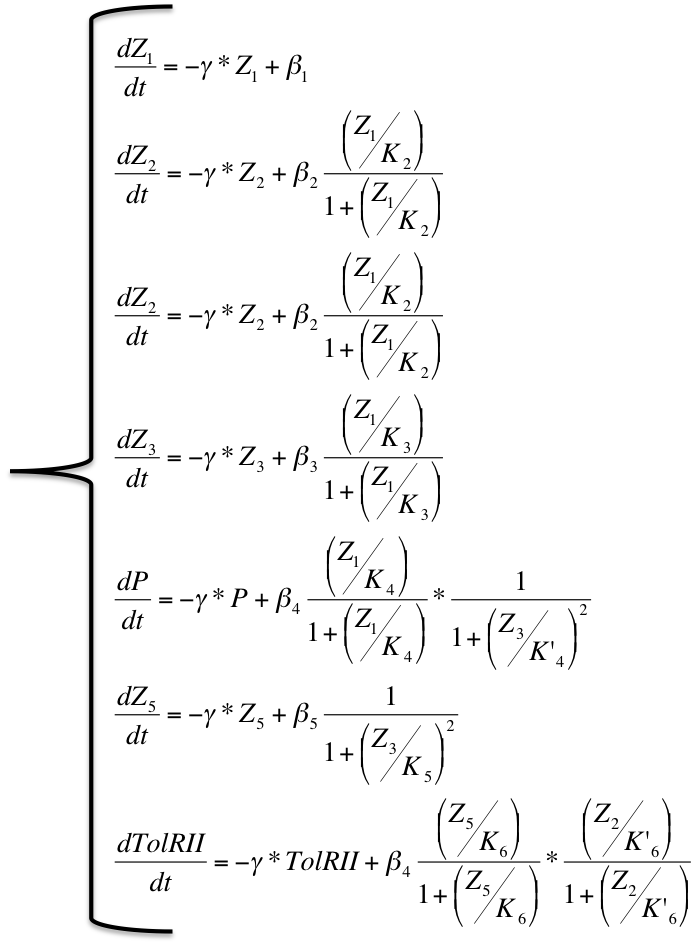
The network behavior can be described by a system of differential equations:
| Differential System |
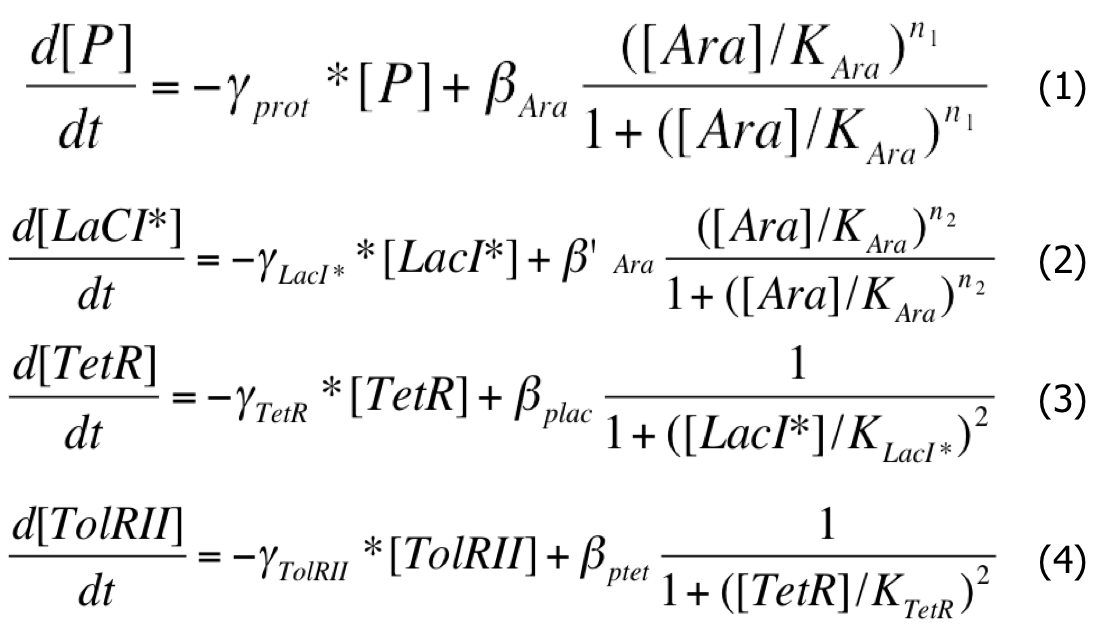
|
As a first approximation, we assumed that :
- When the pBad promoter is induced, the concentration of arabinose in the medium is very high and constant during he whole study; as a consequence, we will consider that the creation rate of Protein P and LacI* is constant during the experiment:
- We considered that all the binding constants are identical[6] and of an average of 40nM which correspond to approximately 40 monomers per cell; we can write that:
- We chose identical intrinsic promoter activities, β, all equal to 4000 proteins/cell cycle:
- All the time units were expressed in units of cell cycle (approximately half an hour). As a consequence we chose a dilution rate γ of 1 for protein without special tags. For the LacI protein with a degradation tag (LacI-LVA), the dilution rate is 3 times higher[3]:
For simulation, we used the following initial quantities:
- 4000 units of TetR (steady state level at inactivation)
- no protein to be encapsulated
- no LacI
- no TolRII
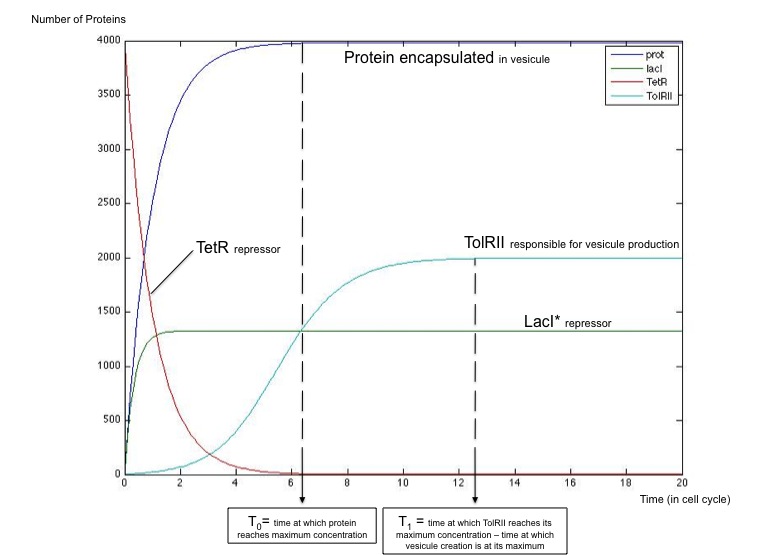
We got the following dynamics using Matlab software, showing that this simple system is capable of generating the desired behavior.
Plot showing that vesiculation happens only once the protein encapsulated has reached its maximal concentration.
Implementing the delay by feed-forward motifs
Using the previous design, we were able to find a way to get the maximal vesicle formation (ie maximal TolRII quantity) when the concentration of proteins to encapsulate is at its maximum. This system satisfies our requirements. However, it is not very flexible. In particular, the concentration of the protein to encapsulate remains always high after induction. This could be detrimental for the cell if this protein is toxic. Consequently, we considered an alternative design allowing for more flexibility, in which the expression of the protein to encapsulate can be transient.
The Feed Forward Motif
The feed forward motif was studied and described by U. Alon in his book "An Introduction to Synthetic Biology" where he describes the different forms of existing feed forward; they can be either coherent or incoherent[7].
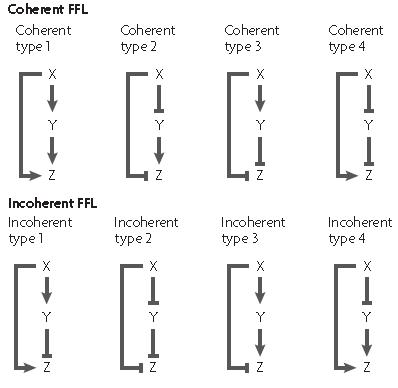
In each case, there exist a different network organization to create a feed-forward loop as shown on the picture
The X and Y inputs can be combined either using a logical AND or OR gate thus giving different dynamics properties in each case.
The coherent feed forward loop allow the creation of a delay either on activation or on deactivation depending on the logical function between the two input X and Y.
When the output logical gate is an AND function, the delay appears on activation, while when the output gate is an OR function, the delay appears on deactivation.
An example of characterized feed-forward loop is the fliA system which was used by the 2008 Paris team in order to create a oscillator.
In an incoherent feed forward loop, the two inputs of the last gate function have two different logical levels; one molecule (molecule X for example) is an activator while the other molecule is a repressor of the promoter controlling the expression of the output.
This way, when the input signal is on, the expression of molecule X starts activating both the promoters controlling the expression of Y and Z ; transcription starts and the amount of Y and Z molecules increases. Nevertheless, when the concentration of Y has reached its repression threshold, repression starts and the concentration of Z molecule decrease to its steady state.
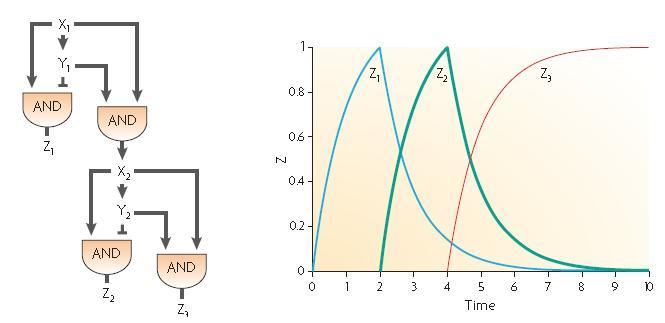
Two elementary incoherent feed forward motifs can be combined in order to create two time delayed bursts as described by U.Alon in its description of the development of Bacillus subtilis spore. Anyway, it seems quite difficult to find a molecule which would be both an activator and a repressor; we decided to try and think about another combination of these two feed forwards so as to get the dynamics we wanted as shown on the following part.
Our design proposition
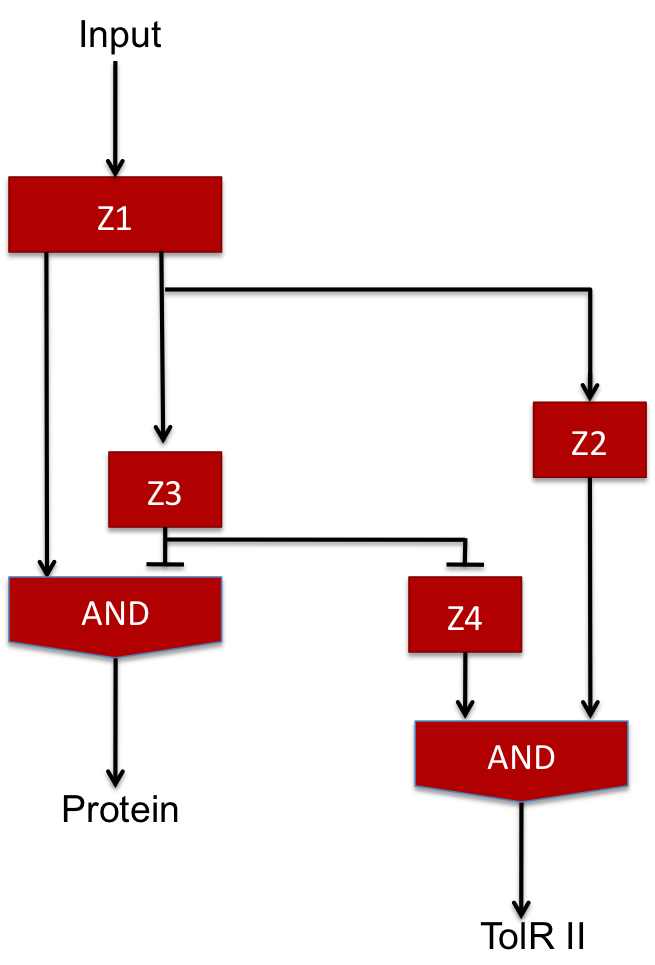
The two feed-forward loops used are incoherent and organized in a way that each molecule is either an activator or a repressor (and not both). The following scheme gives the functioning of the system.
| Steps | Description | Changes in concentration |
| 1 | When the input signal is ON, molecule1 is created and activates the promoter upstream the coding sequence for molecules Z2 and Z3 as well as Z4 since there is no molecule Z3 in the medium (see the AND gate function) at the beginning ; there is an initial amount of Z5 molecule but no Z2 so the no Z6 molecule is produced. | Z1 +++
Z2+ Z3+ Z4++ Z5~ Z6~ |
| 2 | After a while, there is enough Z3 to start the repression of Z4 promoter and there is also enough Z2 to start producing Z6 signficantly. | Z1+++
Z2+++ Z3+++ Z4- Z5~ Z6+ |
| 3 | Then, because of the repression of Z3, the Z4 molecule reaches its steady level ; in the same time, Z3 starts repressing the production of Z5 and the amount of Z6 starts decreasing. | Z1+
Z2+ Z3+ Z4~ Z5- Z6+ |
| 4 | Finally, Z3 represses the promoter allowing the creation of Z5 and every molecules reaches its steady level concentration. | Z1~
Z2~ Z3~ Z4~ Z5- Z6~ |
To study the dynamics of our model, we decided to use a classical approach using Michaelis-Menten equations to model the activation and repression of the various promoters, including a cooperativity of order 2 for the repressor which often the case for 'classical' repressors such as LacI. We obtained the following system of equations:
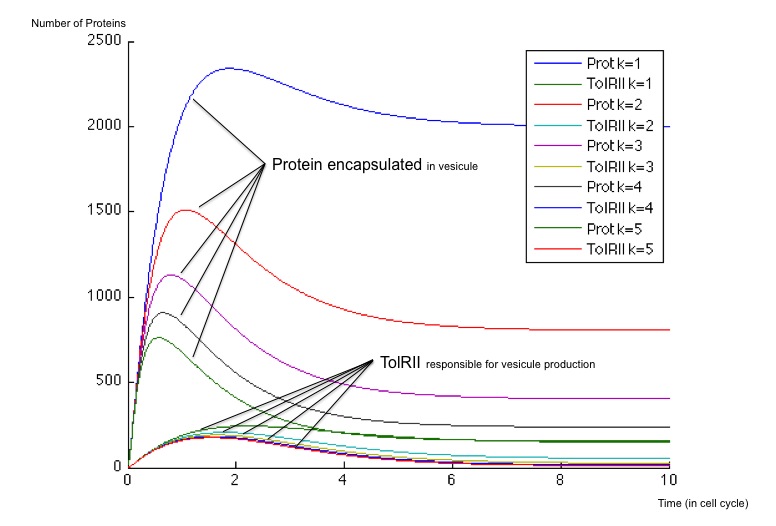
It is possible to obtain a broad spectrum of different dynamical behaviors by playing with the value of the maximum production rate of Z3, called here β . This maximal production rate impacts deeply both the evolution of the exported protein concentration and the evolution of TolRII concentration. This rate is initially chosen equal to 40 and 400 molecules by cell cycle and we have plotted the different responses for multiples of these initial values.
In each case, the initial conditions are the same :
- no proteins encapsulated
- no TolRII
- no Z1 molecules
- no Z2 molecules
- no Z3 molecules
- a weak amount of Z4 molecules (around a hundred)
A more flexible design
Regardless of the steady state concentrations of the molecules, the use of two incoherent feed-forward loops allows for more flexibility in the behaviors of our system. With a simple delay system, we just get one possible dynamic behavior whereas using a variable range of production rates, we can obtain two behaviors:
- When β is weak (β = 40), the number of vesicles (ie of TolRII) produced slowly increase before returning to zero, whereas the protein to be exported inside the vesicles is created very quickly and remains at a high concentration compared to the TolRII protein. We can therefore send vesicles containing an important amount of interesting proteins and, after a while, stop expressing dangerous proteins for the cell. All of this can be performed with a single input signal.
- When β is stronger, the behavior is different and we notice a change of concavity of the TolRII curve when the amount of protein exported is at its maximum. During such a change of concavity, the derivative reaches its maximum - in other words, the amount of TolRII produced by time units is at its highest level. Considering the hypothesis according to which the production rate of vesicles is proportional to TolRII production rate, we get a peak in vesicle production when the concentration of the proteins to exported is at its maximum.
Unfortunately, we did not have the time to try to construct this system in the lab and only the first design was implemented. Nevertheless, we thought that this analysis was very useful to get a good insight of the diversity of the possible behaviors of the sender cells.
References
- ^Ultrasensitivity and noise propagation in a synthetic transcriptional cascade S.Hooshangi & R.Weiss 2005 - [http://www.ncbi.nlm.nih.gov/pubmed/15738412 15738412]
- ^Network motifs : theory and experimental approaches U.Alon 2007 - [http://www.ncbi.nlm.nih.gov/pubmed/17510665 117510665]
- ^New Unstable Variants of Green Fluorescent Protein for Studies of Transient Gene Expression in Bacteria Jens Bo Andersen & Søren Molin 1998 - [http://www.ncbi.nlm.nih.gov:80/pmc/articles/PMC106306/ 106306]
- ^Spatiotemporal control of gene expression with pulse-generating networks S.Basu & R.Weiss 2003 - [http://www.ncbi.nlm.nih.gov/pubmed/15096621 15096621]
- ^Negative Autoregulation Speeds The Response Times of Transcription Network N.Rosenfold & U.Alon 2002 - [http://www.ncbi.nlm.nih.gov/pubmed/12417193 2417193]
- ^A Synthetic oscillatory network of transcriptionnal regulators M.Ellowitz & S.Leibler 1999 - [http://www.ncbi.nlm.nih.gov/pubmed/10659856 10659856]
- ^Structure and function ot the feed-forward Loop Network Motif S.Mangan & U.Alon 2003 - [http://www.ncbi.nlm.nih.gov/pubmed/14530388 14530388]
 "
"