Team:PKU Beijing/Project/Assemble
From 2009.igem.org
(→Result) |
(→Design) |
||
| (10 intermediate revisions not shown) | |||
| Line 60: | Line 60: | ||
At last, we have successfully constructed some of clonings in the first and second strategy, and the most of clonings in the third strategy (AND gate and Bistable on pSB4K5) and the last strategy (three plasmids). After testing the function of some cloning in the third strategy and last strategy, we got some promising results, and based on these results, we can continue to finish our whole project. | At last, we have successfully constructed some of clonings in the first and second strategy, and the most of clonings in the third strategy (AND gate and Bistable on pSB4K5) and the last strategy (three plasmids). After testing the function of some cloning in the third strategy and last strategy, we got some promising results, and based on these results, we can continue to finish our whole project. | ||
| - | Results of the third strategy: | + | '''Results of the third strategy:''' |
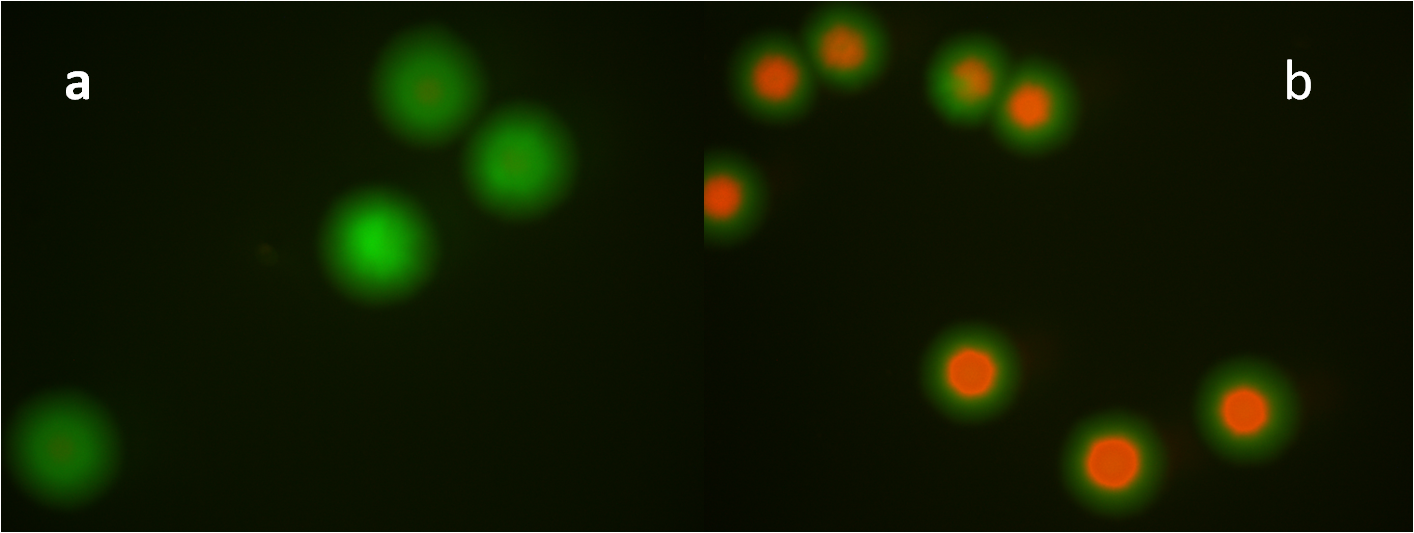
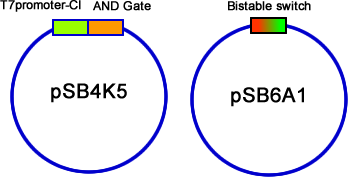
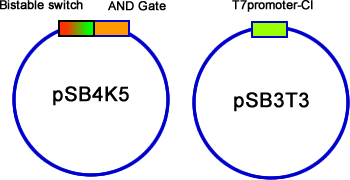
After constructed the AND gate and Bistable on pSB4K5, we then transformed T7-C1 (12 kinds in total: two plasmid, pSB3T3 & pSB6A1; six RBS) into the E.coli. Suspend the colonies contained two plasmids, and plate them with inducers or without inducers (control group). Use fluorescent stereoscope to test the expression of GFP and RFP of colonies. The best result we get is from strategy c (fig 14), Bistable is upstream of AND gate, and C1 with the RBS of B0030. The difference of control group and experiment group is distinguished (fig 18). | After constructed the AND gate and Bistable on pSB4K5, we then transformed T7-C1 (12 kinds in total: two plasmid, pSB3T3 & pSB6A1; six RBS) into the E.coli. Suspend the colonies contained two plasmids, and plate them with inducers or without inducers (control group). Use fluorescent stereoscope to test the expression of GFP and RFP of colonies. The best result we get is from strategy c (fig 14), Bistable is upstream of AND gate, and C1 with the RBS of B0030. The difference of control group and experiment group is distinguished (fig 18). | ||
| - | [[Image:PKU_assemble_SS.png|600px|center|thumb|Fig 18. The best result of the third strategy. Fig 18a: control group, without | + | [[Image:PKU_assemble_SS.png|600px|center|thumb|Fig 18. The best result of the third strategy. Fig 18a: control group, without induction. 18b: induced group: suspend the cells, add 5uL 1M salicylate and 5uL 1M arabinose, plate together with the cells. 18a and 18b are both merged pictures, for original individual figures, please follow [[Team:PKU_Beijing/Project/Assemble/Individual_Pictures|this link]]]] |
Discussion: | Discussion: | ||
| Line 72: | Line 72: | ||
<br> | <br> | ||
| - | Results of the last strategy (three plasmids): | + | '''Results of the last strategy (three plasmids): ''' |
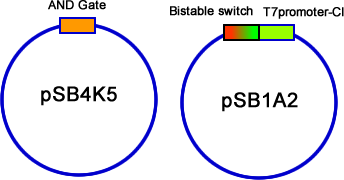
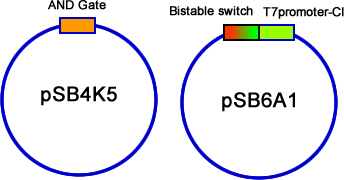
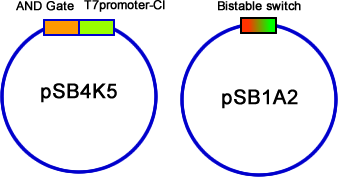
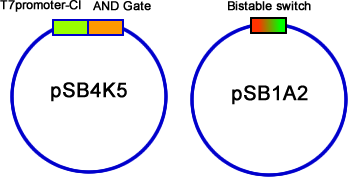
| - | We have constructed the three modules, AND gate, Bistable and T7-CI, on three different plasmids: pSB4K5, pSB1A2/pSB6A1 and pSB3T3. Then we transformed them into E.coli one by one. Then the colonies contained three plasmids were picked into 3 mL liquid LB and incubated until the OD600 value reached 0.4~0.6. At this time, we used different combinations of different concentration of arabinose and salicylate to induce. Every two hours we discarded 2mL LB culture, and added 2mL new LB into it, and the concentration of inducers should be kept. Repeating this step for 3 times, and then cultivated the culture overnight to ensure saturated induction and interaction between different modules. The results were tested by flowcytometry, | + | We have constructed the three modules, AND gate, Bistable and T7-CI, on three different plasmids: pSB4K5, pSB1A2/pSB6A1 and pSB3T3. Then we transformed them into E.coli one by one. Then the colonies contained three plasmids were picked into 3 mL liquid LB and incubated until the OD600 value reached 0.4~0.6. At this time, we used different combinations of different concentration of arabinose and salicylate to induce. Every two hours we discarded 2mL LB culture, and added 2mL new LB into it, and the concentration of inducers should be kept. Repeating this step for 3 times, and then cultivated the culture overnight to ensure saturated induction and interaction between different modules. The results were tested by flowcytometry, controls that constitutively expresses GFP and RFP are used to determine the red cell region and green cell region(fig 19). Flowcytometry data of the first stage system is analysis by counting the cells that falls in both of the regions. |
[[Image:PKU_assemble_control.png|600px|center|thumb|Fig 19. The RFP, GFP controls and definition of different states areas. The Y axis denotes the strength of RFP fluorescence, and the X axis means the strength of GFP fluorescence. Fig 19a is the result of RFP control, so we enclosed a polygon area as the state of RFP in Bistable Switch (CI side). Fig 19b is the result of GFP control, depended on which we defined the area as the state of GFP. ]] | [[Image:PKU_assemble_control.png|600px|center|thumb|Fig 19. The RFP, GFP controls and definition of different states areas. The Y axis denotes the strength of RFP fluorescence, and the X axis means the strength of GFP fluorescence. Fig 19a is the result of RFP control, so we enclosed a polygon area as the state of RFP in Bistable Switch (CI side). Fig 19b is the result of GFP control, depended on which we defined the area as the state of GFP. ]] | ||
| - | + | Taking advantages of the defined enclosed areas, we can count the cell number in either states of Bistable switch. And this is a criterion to assess the quality of assembling. We have tested a number of assemblies and different combinations of various concentrations of arabinose and salicylate. | |
| + | |||
| + | At last, the best results are from the assembling system, which is made of AND gate on pSB4A5, Bistable on pSB1A2 and CI on plasmid pSB3T3, with the RBS of J44001. The concentrations of inducers are 10^-4M arabinose and 10^-6M salicylate. | ||
| + | |||
| + | |||
| + | [[Image:PKU_assemble_WSK.jpg|600px|center|thumb|Fig 20. The best result of the three plasmids strategy. The Y axis denotes the number of cells in the defined enclosed areas (figs on the left side for GFP area and figs on the right side for RFP area). The X axis means the strength of G(R)FP fluorescence. Fig 20a: control group, without inducing. Fig 20b: arabinose single inducing group, the concentration is 10^-4M. Fig 20c: salicylate single inducing group, the concentration is 10^-6M. Fig 20d: arabinose and salicylate double inducing group, the concentration of arabinose is 10^-4M and of salicylate is 10^-6M]] | ||
| + | |||
| + | We calculated red to green cell number ratio. The histogram of the Red/Green ratio under four conditions (no induction, 2 x singal induction, double induction) are shown in Fig21: | ||
| + | |||
| + | [[Image:PKU_Red_Region_divide_Green_Region.png|600px|center|thumb|Fig21. The columns are the red to green cell number ratio values. From left to right, it is the no induction data, arabinose singal induction data, salicylate singal induction data and the double induction data. Double induction data shows significant difference from that of singal induction and no induction. Although the arabinose induced is to some extent leaky, it shows great difference form the double induction group.]] | ||
| + | |||
| + | <br><br> | ||
==='''Second Stage Assembly (In Process)'''=== | ==='''Second Stage Assembly (In Process)'''=== | ||
| Line 92: | Line 103: | ||
The final assembly introduced another GFP reporter (the OR Gate - GFP set), so we want to delete the GFP gene from the bistable device, while the RFP gene is kept to track the change of memory state. | The final assembly introduced another GFP reporter (the OR Gate - GFP set), so we want to delete the GFP gene from the bistable device, while the RFP gene is kept to track the change of memory state. | ||
| - | Actually, our second stage assembly can be viewed as the construction of an AND Gate: | + | Actually, our second stage assembly can be viewed as the construction of an AND Gate: what we need to adjust is the rbs of PhiR73(amber mutation). Input to the PhiR73 is from the bistable module, and the input to supD is the same as the first AND Gate. |
| - | + | ||
{{PKU_Beijing/Foot}} | {{PKU_Beijing/Foot}} | ||
Latest revision as of 17:18, 21 October 2009
|
|||||||||||||
|
|||||||||||||
 "
"