Team:Paris/Tool DataBase
From 2009.igem.org
Christophe.R (Talk | contribs) (→DataBase overview) |
Cha.olivier (Talk | contribs) (→Biological Databases) |
||
| (41 intermediate revisions not shown) | |||
| Line 1: | Line 1: | ||
| + | <span/ id="bottom">[https://2009.igem.org/ iGEM ] > [[Team:Paris#top | Paris]] > [[Team:Paris/Tool_introduction#top | Tool]] > [[Team:Paris/Tool_DataBase#bottom | Databases]] | ||
{{Template:Paris2009}} | {{Template:Paris2009}} | ||
| - | {{Template: | + | {{Template:Paris2009_menu6}} |
| - | == | + | ==Databases== |
| - | At this point, we want to know how do we think our database. | + | At this point, we want to know how do we think our database. In fact we don't have only one database. We thought of separate the application database in several parts. In order to have an application which is as effective as we want, we discuss about the fact of being able to fit it for as many laboratory needs as we could. We had to keep in mind that laboratories aren't the same and some have already their own databases or even use LIMS |
| - | * theoretical DNA ( | + | ===How many ?=== |
| - | * physical DNA ( | + | The main idea was to separate the application database by uses. We know that we need to have a database witch will be use for protocols, and/or for the lab notebook and another one for the biological use. We know that some laboratories use some system for the stock management and need to be connected together, but that's not enough. |
| + | |||
| + | ====Protocols==== | ||
| + | The protocols database is obvious :). We need to have it separate from the rest of the data. It's the main part of our application, and we need to think it independently of the biological part. Knowing that many laboratories already use biological databases (but not too many protocols database), making the protocols apart is more easy to implement. | ||
| + | |||
| + | ====Biology==== | ||
| + | We already discuss about the stock management database generally already implement in laboratories (system implement in every LIMS software/freeware). But only these two databases aren't enough for the whole system.For some, the stock management database not only support biological use and isn't as specific as it should be. Stock doesn't mean every times previous constructions, or future ones. We need to have another one more specific to biology, one for all constructions, we call it the ''in silico'' database. | ||
| + | |||
| + | ====Schema==== | ||
| + | [[Image:Databases.png | 400px | center ]] | ||
| + | |||
| + | ===Evolution=== | ||
| + | The main idea of separate those databases is to be able to trade them by existing ones. The originals databases need to be fully adaptable. The software support natively the SQLite databases, but also will support mySQL, pgSQL, Oracle, and Microsoft SQL. | ||
| + | |||
| + | <html> | ||
| + | </div> | ||
| + | <div id="paris_content_boxtop"> | ||
| + | </div> | ||
| + | <div id="paris_content"> | ||
| + | </html> | ||
| + | |||
| + | ==Biological Databases== | ||
| + | |||
| + | We thought of integrating a modifiable software which can fit easily with databases. At a biological point of view, we've got to deal with the fact that we have to represent every DNA, those we have, those we had, and those we'll have. We also need a Database who can be use as a lab notebook, and a Protocols reference system. For the Biological part, we discuss about the principle to separate the theoretical DNA with the physical one: | ||
| + | |||
| + | * theoretical DNA (''in silico'' DNA : <sub>is</sub>DNA) represent every DNA in any form (circular, linear, double, single, length etc...). With this methodology, we don't need to discuss about the presence of it in the lab. We can also name a <sub>is</sub>DNA witch is compose of other <sub>is</sub>DNA, and then, form entity like plasmid or even more full genome. | ||
| + | * physical DNA (laboratory DNA : <sub>lab</sub>DNA) represent the working present DNA in the lab. It refers to the <sub>is</sub>DNA with some physical informations, like : quantities, freezer places and owners. | ||
| + | |||
| + | |||
| + | this biological core database is enveloped by owner database which support the LabBook and Protocol databases. | ||
| + | |||
| + | |||
| + | {{Template:Paris2009_guided|Tool_introduction#top|Tool_OSXSoft#top}} | ||
Latest revision as of 16:04, 21 October 2009
iGEM > Paris > Tool > Databases
Contents |
Databases
At this point, we want to know how do we think our database. In fact we don't have only one database. We thought of separate the application database in several parts. In order to have an application which is as effective as we want, we discuss about the fact of being able to fit it for as many laboratory needs as we could. We had to keep in mind that laboratories aren't the same and some have already their own databases or even use LIMS
How many ?
The main idea was to separate the application database by uses. We know that we need to have a database witch will be use for protocols, and/or for the lab notebook and another one for the biological use. We know that some laboratories use some system for the stock management and need to be connected together, but that's not enough.
Protocols
The protocols database is obvious :). We need to have it separate from the rest of the data. It's the main part of our application, and we need to think it independently of the biological part. Knowing that many laboratories already use biological databases (but not too many protocols database), making the protocols apart is more easy to implement.
Biology
We already discuss about the stock management database generally already implement in laboratories (system implement in every LIMS software/freeware). But only these two databases aren't enough for the whole system.For some, the stock management database not only support biological use and isn't as specific as it should be. Stock doesn't mean every times previous constructions, or future ones. We need to have another one more specific to biology, one for all constructions, we call it the in silico database.
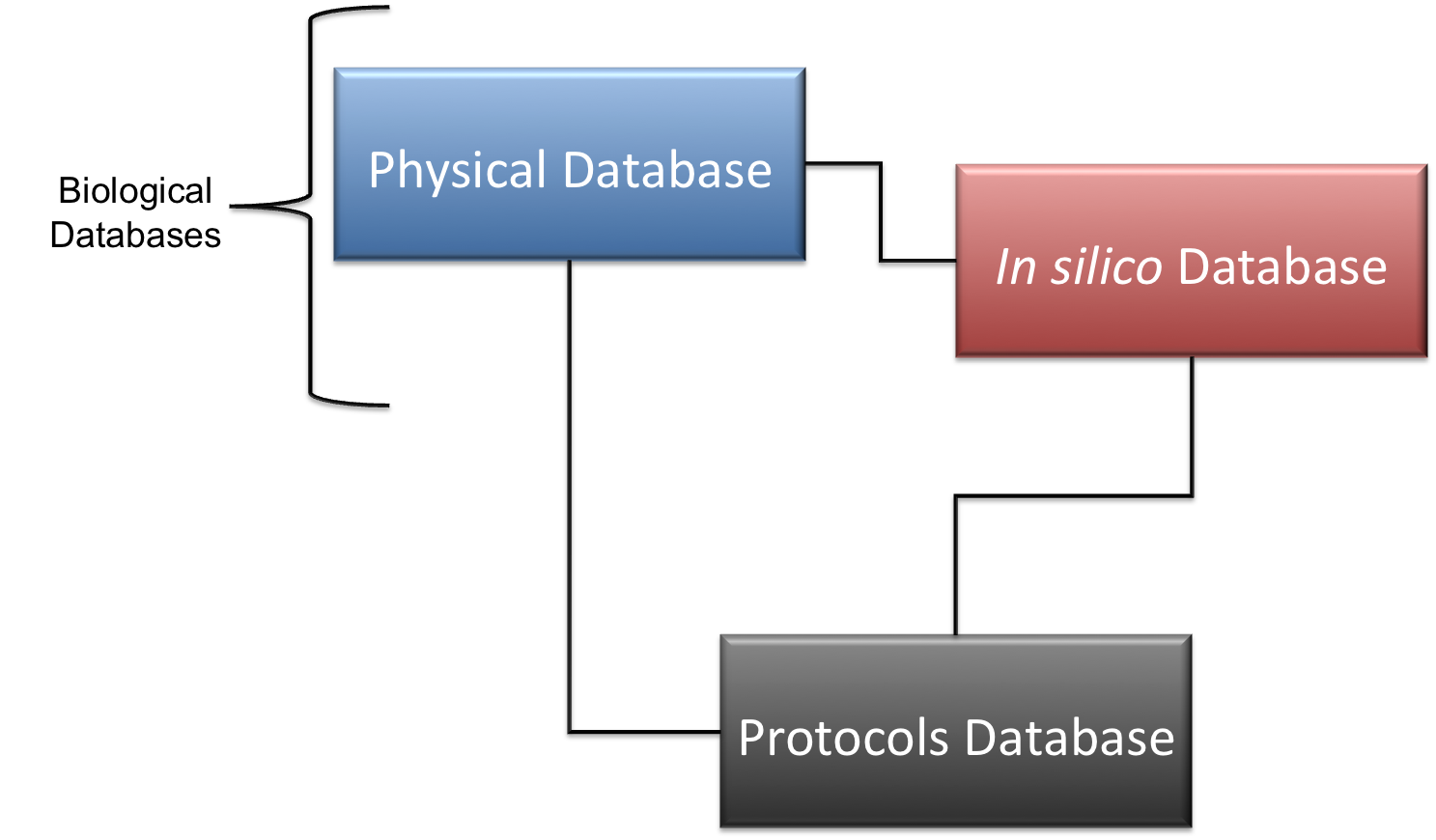
Schema
Evolution
The main idea of separate those databases is to be able to trade them by existing ones. The originals databases need to be fully adaptable. The software support natively the SQLite databases, but also will support mySQL, pgSQL, Oracle, and Microsoft SQL.
Biological Databases
We thought of integrating a modifiable software which can fit easily with databases. At a biological point of view, we've got to deal with the fact that we have to represent every DNA, those we have, those we had, and those we'll have. We also need a Database who can be use as a lab notebook, and a Protocols reference system. For the Biological part, we discuss about the principle to separate the theoretical DNA with the physical one:
- theoretical DNA (in silico DNA : isDNA) represent every DNA in any form (circular, linear, double, single, length etc...). With this methodology, we don't need to discuss about the presence of it in the lab. We can also name a isDNA witch is compose of other isDNA, and then, form entity like plasmid or even more full genome.
- physical DNA (laboratory DNA : labDNA) represent the working present DNA in the lab. It refers to the isDNA with some physical informations, like : quantities, freezer places and owners.
this biological core database is enveloped by owner database which support the LabBook and Protocol databases.
 "
"