Team:Paris/Transduction testing
From 2009.igem.org
(→Fusion) |
(→Fusion) |
||
| (97 intermediate revisions not shown) | |||
| Line 1: | Line 1: | ||
| - | <span/ id="bottom">[https://2009.igem.org/ iGEM ] > [[Team:Paris#top | Paris]] > [[ | + | <span/ id="bottom">[https://2009.igem.org/ iGEM ] > [[Team:Paris#top | Paris]] > [[team:Paris/WetLab#WetLab| WetLab]] > [[Team:Paris/Transduction_testing#bottom | Reception]] |
{{Template:Paris2009}} | {{Template:Paris2009}} | ||
{{Template:Paris2009_menu4}} | {{Template:Paris2009_menu4}} | ||
| - | == WetLab == | + | |
| - | <center> | + | == WetLab - Reception== |
| - | <center> | + | <html> |
| + | <style type="text/css"> | ||
| + | #left-side { | ||
| + | position: absolute; | ||
| + | height: 23px; | ||
| + | width: 30px; | ||
| + | top: 0px; | ||
| + | left: 180px; | ||
| + | margin-top:10px; | ||
| + | padding-top: 7px; | ||
| + | background: url(https://static.igem.org/mediawiki/2009/1/1b/Left_menu_pari.png); | ||
| + | z-index:4; | ||
| + | } | ||
| + | |||
| + | #middle-side { | ||
| + | height: 25px; | ||
| + | width: 320px; | ||
| + | position: absolute; | ||
| + | top: 0px; | ||
| + | left: 190px; | ||
| + | margin-top:10px; | ||
| + | padding-top: 5px; | ||
| + | background: #dadada; | ||
| + | z-index:5; | ||
| + | } | ||
| + | |||
| + | #right-side { | ||
| + | position: absolute; | ||
| + | height: 23px; | ||
| + | width: 30px; | ||
| + | margin-top:10px; | ||
| + | padding-top: 7px; | ||
| + | top: 0px; | ||
| + | left: 490px; | ||
| + | background: url(https://static.igem.org/mediawiki/2009/4/40/Right_menu_paris.png); | ||
| + | z-index:4; | ||
| + | } | ||
| + | |||
| + | a.menu_sub { | ||
| + | padding-left: 7px; | ||
| + | padding-right: 7px; | ||
| + | } | ||
| + | |||
| + | a.menu_sub_active { | ||
| + | padding-left: 7px; | ||
| + | padding-right: 7px; | ||
| + | color:#b0310e; | ||
| + | font-weight:bold; | ||
| + | } | ||
| + | </style> | ||
| + | <div id="left-side"></div> | ||
| + | <div id="middle-side"><center> | ||
| + | <a class="menu_sub"href="https://2009.igem.org/Team:Paris/WetLab#bottom"> Main </a>| | ||
| + | <a class="menu_sub" href="https://2009.igem.org/Team:Paris/Addressing_testing#bottom"> Addressing</a>| | ||
| + | <a class="menu_sub"href="https://2009.igem.org/Team:Paris/Production_testing#bottom"> Production</a>| | ||
| + | <a class="menu_sub_active"href="https://2009.igem.org/Team:Paris/Transduction_testing#bottom"> Reception</a> | ||
| + | </center> | ||
| + | </div> | ||
| + | <div id="right-side"></div> | ||
| + | </html> | ||
| + | |||
| + | In this section we have two kinds of experiments : | ||
| + | |||
| + | |||
| + | #[[Team:Paris/Transduction_testing#Fusion| Fusion :]] the construction required for the fusion of the vesicles that contain the message with the receiver. | ||
| + | #[[Team:Paris/Transduction_testing#Transduction| Transduction :]] the construction of the system that will be able de decypher the message. | ||
| + | |||
| + | |||
| + | The first part deals with G3P , OmpAL , Jun, Fos , and AIDA protein. the second with the transduction system of the fec operon (FecA, FecR, FecI). | ||
| + | |||
<html> | <html> | ||
</div> | </div> | ||
| Line 13: | Line 82: | ||
</html> | </html> | ||
| + | == Fusion == | ||
| - | + | Fusion : the construction required for the fusion of the vesicles that contain the message. | |
| - | + | G3P , OmpAL , Jun, Fos , and AIDA protein. | |
| - | |||
| - | |||
| - | |||
| + | <center><font style="color:green;font-weight:bold; font-size:16px">The green color means the experiment was a success</font></center> | ||
| + | |||
| + | |||
| + | [[Image:Paris_construc_7_aida.jpg |800px|center|plasmid = ]] | ||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | <font style="color:black;font-weight:bold; font-size:16px"><I>'''Experiments ran :'''</I></font> | ||
| + | |||
| + | |||
| + | {| | ||
| + | |- style="background: #0a3585; text-align: center; color:black; font-weight:bold;font-size:14px " | ||
| + | |width="300px" ; bgcolor="#F8F8F8"| Column 1 | ||
| + | |width="300px" ; bgcolor="#E0E3FE"| Column 2 | ||
| + | |width="300px" ; bgcolor="#CBD1FD"| Column 3 | ||
| + | |width="300px" ; bgcolor="#BCCDFD"| Column 4 | ||
| + | |||
| + | |- style="background: #bebebe; text-align: center;" | ||
| + | |bgcolor="#F8F8F8"|'''PCR :''' | ||
| + | |||
| + | AidA and Linker : Design, synthesize and order in two parts due to the seize. | ||
| + | |||
| + | |bgcolor="#E0E3FE"|- | ||
| + | |bgcolor="#CBD1FD"|- | ||
| + | |bgcolor="#BCCDFD"|- | ||
| + | |||
| + | |||
| + | |- style="background: #d8d8d8; text-align: center;" | ||
| + | |bgcolor="#F8F8F8"|'''Verification on gel :''' | ||
| + | |||
| + | ok | ||
| + | |||
| + | |bgcolor="#E0E3FE"|- | ||
| + | |bgcolor="#CBD1FD"|- | ||
| + | |bgcolor="#BCCDFD"|- | ||
| + | |||
| + | |- style="background: #bebebe; text-align: center;" | ||
| + | |bgcolor="#F8F8F8"|'''Purification on gel :''' | ||
| + | |||
| + | ok | ||
| + | |||
| + | |bgcolor="#E0E3FE"|- | ||
| + | |||
| + | |bgcolor="#CBD1FD"|- | ||
| + | |bgcolor="#BCCDFD"|- | ||
| + | |||
| + | |||
| + | |||
| + | |- style="background: #d8d8d8; text-align: center;" | ||
| + | |bgcolor="#F8F8F8"|'''Digestion:''' | ||
| + | |||
| + | AidA CTerm Kpn1/HindIII | ||
| + | |||
| + | AidA NTerm Kpn1/HindIII | ||
| + | |||
| + | |bgcolor="#E0E3FE"|'''Digestion:''' | ||
| + | |||
| + | AidA complete E/P | ||
| + | |||
| + | PSB1A3 E/P | ||
| + | |||
| + | We did it to obtain a biobrick first | ||
| + | |||
| + | |bgcolor="#CBD1FD"|'''Digestion:''' | ||
| + | none | ||
| + | |bgcolor="#BCCDFD"|- | ||
| + | |||
| + | |||
| + | |- style="background: #bebebe; text-align: center;" | ||
| + | |bgcolor="#F8F8F8"|'''Digestion checking:''' | ||
| + | |||
| + | ok | ||
| + | |||
| + | |bgcolor="#E0E3FE"|'''Digestion checking:''' | ||
| + | ok | ||
| + | |||
| + | |||
| + | |bgcolor="#CBD1FD"|'''Digestion checking:''' | ||
| + | none | ||
| + | |||
| + | |||
| + | |bgcolor="#BCCDFD"| | ||
| + | |||
| + | - | ||
| + | |||
| + | |||
| + | |- style="background: #d8d8d8; text-align: center;" | ||
| + | |bgcolor="#F8F8F8"|'''Ligation:''' | ||
| + | |||
| + | AidA CTerm (Kpn1/HindIII):: aidA NTerm (Kpn1/HindIII) | ||
| + | |||
| + | |||
| + | |bgcolor="#E0E3FE"|'''Ligation:''' | ||
| + | |||
| + | AidA complete E/P :: PSB1A3 E/P | ||
| + | |||
| + | |bgcolor="#CBD1FD"|'''Ligation:''' | ||
| + | none | ||
| + | |bgcolor="#BCCDFD"|- | ||
| + | |||
| + | |- style="background: #bebebe; text-align: center;" | ||
| + | |bgcolor="#F8F8F8"|'''Colony pCR :''' | ||
| + | |||
| + | ok | ||
| + | |||
| + | |bgcolor="#E0E3FE"|'''Colony PCR:''' | ||
| + | |||
| + | ok | ||
| + | |||
| + | |bgcolor="#CBD1FD"|'''Colony PCR :''' | ||
| + | none | ||
| + | |||
| + | |bgcolor="#BCCDFD"|- | ||
| + | |||
| + | |- style="background: #d8d8d8; text-align: center;" | ||
| + | |bgcolor="#F8F8F8"|'''Miniprep:''' | ||
| + | |||
| + | done | ||
| + | |||
| + | |bgcolor="#E0E3FE"|'''Miniprep:''' | ||
| + | |||
| + | done | ||
| + | |||
| + | |bgcolor="#CBD1FD"|'''Miniprep:''' | ||
| + | none | ||
| + | |||
| + | |bgcolor="#BCCDFD"|- | ||
| + | |||
| + | |- style="background: #bebebe; text-align: center;" | ||
| + | |bgcolor="#F8F8F8"|'''Sequencing :''' | ||
| + | unecessary | ||
| + | |||
| + | |bgcolor="#E0E3FE"|'''Sequencing :''' | ||
| + | none | ||
| + | |||
| + | |bgcolor="#CBD1FD"|'''Sequencing :''' | ||
| + | none | ||
| + | |||
| + | |||
| + | |bgcolor="#BCCDFD"|- | ||
| + | |||
| + | |- style="background: #d8d8d8; text-align: center;" | ||
| + | |bgcolor="#F8F8F8"| '''Stock glycerol:''' | ||
| + | |||
| + | - | ||
| + | |bgcolor="#E0E3FE"|'''Stock glycerol''' | ||
| + | |||
| + | S75 | ||
| + | |||
| + | |bgcolor="#CBD1FD"|'''Stock glycerol''' | ||
| + | |||
| + | none | ||
| + | |bgcolor="#BCCDFD"|- | ||
| + | |||
| + | |} | ||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | <center><font style="color:green;font-weight:bold; font-size:16px">The green color means the experiment was a success</font></center> | ||
| + | |||
| + | |||
| + | |||
| + | [[Image:Paris_construc_8_junfos.jpg |800px|center|plasmid = ]] | ||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | <font style="color:black;font-weight:bold; font-size:16px"><I>'''Experiments ran :'''</I></font> | ||
| + | |||
| + | |||
| + | {| | ||
| + | |- style="background: #0a3585; text-align: center; color:black; font-weight:bold;font-size:14px " | ||
| + | |width="300px" ; bgcolor="#F8F8F8"| Column 1 | ||
| + | |width="300px" ; bgcolor="#E0E3FE"| Column 2 | ||
| + | |width="300px" ; bgcolor="#CBD1FD"| Column 3 | ||
| + | |width="300px" ; bgcolor="#BCCDFD"| Column 4 | ||
| + | |||
| + | |- style="background: #bebebe; text-align: center;" | ||
| + | |bgcolor="#F8F8F8"|'''PCR :''' | ||
| + | |||
| + | Jun *: design and order to be a a mutant incapable to form homodimere. | ||
| + | |||
| + | pLac : previously done | ||
| + | |||
| + | SSOmpA : previously done | ||
| + | |||
| + | Fos : previously done | ||
| + | |||
| + | |bgcolor="#E0E3FE"|- | ||
| + | |bgcolor="#CBD1FD"|- | ||
| + | |bgcolor="#BCCDFD"|- | ||
| + | |||
| + | |||
| + | |||
| + | |- style="background: #d8d8d8; text-align: center;" | ||
| + | |bgcolor="#F8F8F8"|'''Verification on gel :''' | ||
| + | |||
| + | ok | ||
| + | |||
| + | |bgcolor="#E0E3FE"| | ||
| + | |||
| + | - | ||
| + | |||
| + | |bgcolor="#CBD1FD"| | ||
| + | |||
| + | - | ||
| + | |bgcolor="#BCCDFD"|- | ||
| + | |||
| + | |- style="background: #bebebe; text-align: center;" | ||
| + | |bgcolor="#F8F8F8"|'''Purification on gel :''' | ||
| + | |||
| + | ok | ||
| + | |||
| + | |bgcolor="#E0E3FE"| | ||
| + | |||
| + | - | ||
| + | |||
| + | |bgcolor="#CBD1FD"| | ||
| + | |||
| + | - | ||
| + | |bgcolor="#BCCDFD"| | ||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |- style="background: #d8d8d8; text-align: center;" | ||
| + | |bgcolor="#F8F8F8"|'''Digestion:''' | ||
| + | |||
| + | pLac E/X | ||
| + | |||
| + | ssOmpA E/S | ||
| + | |||
| + | Jun E/X and E/P | ||
| + | |||
| + | PSB1A3 E/P | ||
| + | |||
| + | |bgcolor="#E0E3FE"|'''Digestion:''' | ||
| + | none | ||
| + | |bgcolor="#CBD1FD"|- | ||
| + | |bgcolor="#BCCDFD"|- | ||
| + | |||
| + | |||
| + | |||
| + | |- style="background: #bebebe; text-align: center;" | ||
| + | |bgcolor="#F8F8F8"|'''Digestion checking:''' | ||
| + | |||
| + | ok | ||
| + | |||
| + | |bgcolor="#E0E3FE"|'''Digestion checking:''' | ||
| + | |||
| + | none | ||
| + | |||
| + | |bgcolor="#CBD1FD"| | ||
| + | - | ||
| + | |bgcolor="#BCCDFD"| | ||
| + | - | ||
| + | |||
| + | |- style="background: #d8d8d8; text-align: center;" | ||
| + | |bgcolor="#F8F8F8"|'''Ligation:''' | ||
| + | |||
| + | pLac on PSB2K3 E/X :: ssOmpA E/S | ||
| + | |||
| + | Jun E/P :: PSB1A3 E/P | ||
| + | |||
| + | |bgcolor="#E0E3FE"|'''Ligation:''' | ||
| + | none | ||
| + | |bgcolor="#CBD1FD"|- | ||
| + | |bgcolor="#BCCDFD"|'- | ||
| + | |||
| + | |||
| + | |||
| + | |- style="background: #bebebe; text-align: center;" | ||
| + | |bgcolor="#F8F8F8"|'''Colony PCR:''' | ||
| + | |||
| + | ok | ||
| + | |||
| + | |bgcolor="#E0E3FE"|'''Colony PCR:''' | ||
| + | none | ||
| + | |||
| + | |bgcolor="#CBD1FD"|- | ||
| + | |||
| + | - | ||
| + | |bgcolor="#BCCDFD"|- | ||
| + | - | ||
| + | |||
| + | |- style="background: #d8d8d8; text-align: center;" | ||
| + | |bgcolor="#F8F8F8"|'''Miniprep:''' | ||
| + | |||
| + | ok | ||
| + | |bgcolor="#E0E3FE"|'''Miniprep:''' | ||
| + | |||
| + | none | ||
| + | |||
| + | |bgcolor="#CBD1FD"|- | ||
| + | |bgcolor="#BCCDFD"|- | ||
| + | |||
| + | |- style="background: #bebebe; text-align: center;" | ||
| + | |bgcolor="#F8F8F8"|'''Sequencing :''' | ||
| + | |||
| + | -ok | ||
| + | |||
| + | |bgcolor="#E0E3FE"|'''Sequencing :''' | ||
| + | |||
| + | -none | ||
| + | |||
| + | |bgcolor="#CBD1FD"|'- | ||
| + | |bgcolor="#BCCDFD"|- | ||
| + | |||
| + | |- style="background: #d8d8d8; text-align: center;" | ||
| + | |bgcolor="#F8F8F8"| '''Stock glycerol:''' | ||
| + | none | ||
| + | |||
| + | unecessary | ||
| + | |bgcolor="#E0E3FE"|'''Stock glycerol''' | ||
| + | |||
| + | none | ||
| + | |bgcolor="#CBD1FD"|- | ||
| + | |bgcolor="#BCCDFD"|- | ||
| + | |||
| + | - | ||
| + | |} | ||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | ===G3P=== | ||
| + | |||
| + | Even though it has not been realized, we planned to do these experiments to verify that G3P is expressed into the bacterial surface and induces the membranal fusion. | ||
| - | |||
| - | |||
| - | |||
*<b>Surface expression of G3P</b> | *<b>Surface expression of G3P</b> | ||
Surface expression of G3P could be seen with antibodies.<br> | Surface expression of G3P could be seen with antibodies.<br> | ||
Culture of bacteria expressing OmpA-linker-G3P construction on a membrane. Adding antibodies anti-G3P. Revelation should shows whether G3P is expressed at the surface of bacteria. | Culture of bacteria expressing OmpA-linker-G3P construction on a membrane. Adding antibodies anti-G3P. Revelation should shows whether G3P is expressed at the surface of bacteria. | ||
| + | |||
*<b>Floculation test</b> | *<b>Floculation test</b> | ||
| Line 38: | Line 441: | ||
Two separate bacterial population growth on media. A population (F-) expressing the OmpA-linker_G3P construction and a population (F +) which express sexual pilus (by expression of the episome F).<br> | Two separate bacterial population growth on media. A population (F-) expressing the OmpA-linker_G3P construction and a population (F +) which express sexual pilus (by expression of the episome F).<br> | ||
Then we growth bacteria without agitation and we look (compared to culture controls) the speed of sedimentation. Faster sedimentation is due to an interaction between bacterial populations showing an increase of contact time. At vesicle level, it induce membrane fusion. | Then we growth bacteria without agitation and we look (compared to culture controls) the speed of sedimentation. Faster sedimentation is due to an interaction between bacterial populations showing an increase of contact time. At vesicle level, it induce membrane fusion. | ||
| + | |||
| + | |||
| + | <html> | ||
| + | <a href="https://2009.igem.org/Team:Paris/Transduction_testing#bottom"><img style="width:40px; height:40px;" src="https://static.igem.org/mediawiki/2009/1/10/Paris_Up.png"/></a> | ||
| + | </html> | ||
| + | |||
| + | <html> | ||
| + | </div> | ||
| + | <div id="paris_content_boxtop"> | ||
| + | </div> | ||
| + | <div id="paris_content"> | ||
| + | </html> | ||
| + | |||
| + | == Transduction == | ||
| + | |||
| + | |||
| + | Transduction : the construction of the system that will be able de decypher the message. | ||
| + | |||
| + | |||
| + | The transduction system of the fec operon (FecA, FecR, FecI). | ||
| + | |||
| + | |||
| + | |||
| + | |||
| + | <center><font style="color:green;font-weight:bold; font-size:16px">The green color means the experiment was a success</font></center> | ||
| + | |||
| + | |||
| + | |||
| + | [[Image:Paris_construct_9_pFec-RFP.jpg |800px|center|plasmid = ]] | ||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | <font style="color:black;font-weight:bold; font-size:16px"><I>'''Experiments ran :'''</I></font> | ||
| + | |||
| + | |||
| + | {| | ||
| + | |- style="background: #0a3585; text-align: center; color:black; font-weight:bold;font-size:14px " | ||
| + | |width="300px" ; bgcolor="#F8F8F8"| Column 1 | ||
| + | |width="300px" ; bgcolor="#E0E3FE"| Column 2 | ||
| + | |width="300px" ; bgcolor="#CBD1FD"| Column 3 | ||
| + | |width="300px" ; bgcolor="#BCCDFD"| Column 4 | ||
| + | |||
| + | |- style="background: #bebebe; text-align: center;" | ||
| + | |bgcolor="#F8F8F8"|'''PCR :''' | ||
| + | |||
| + | pFec matrix: K12 DNA | ||
| + | |||
| + | Oligo: TM | ||
| + | |||
| + | FecI matrix: K12 DNA | ||
| + | |||
| + | Oligo: TM | ||
| + | |||
| + | FecR 1-85 matrix: K12 DNA | ||
| + | |||
| + | Oligo: TM | ||
| + | |||
| + | mRFP matrix: PSB1A3 (previous IGEM plate) | ||
| + | |||
| + | Oligo: TM | ||
| + | |||
| + | |||
| + | |||
| + | |bgcolor="#E0E3FE"|- | ||
| + | |bgcolor="#CBD1FD"|- | ||
| + | |bgcolor="#BCCDFD"|- | ||
| + | |- style="background: #d8d8d8; text-align: center;" | ||
| + | |bgcolor="#F8F8F8"|'''Verification on gel :''' | ||
| + | |||
| + | ok | ||
| + | |||
| + | |bgcolor="#E0E3FE"|- | ||
| + | |bgcolor="#CBD1FD"|- | ||
| + | |bgcolor="#BCCDFD"|- | ||
| + | |||
| + | |- style="background: #bebebe; text-align: center;" | ||
| + | |bgcolor="#F8F8F8"|'''Purification on gel :''' | ||
| + | |||
| + | ok | ||
| + | |||
| + | |bgcolor="#E0E3FE"|- | ||
| + | |bgcolor="#CBD1FD"|- | ||
| + | |bgcolor="#BCCDFD"|- | ||
| + | |||
| + | |||
| + | |||
| + | |- style="background: #d8d8d8; text-align: center;" | ||
| + | |bgcolor="#F8F8F8"|'''Digestion:''' | ||
| + | |||
| + | Ter E/X | ||
| + | |||
| + | mRFP E/S | ||
| + | |||
| + | |bgcolor="#E0E3FE"| - | ||
| + | |bgcolor="#CBD1FD"| - | ||
| + | |bgcolor="#BCCDFD"| - | ||
| + | |||
| + | |- style="background: #bebebe; text-align: center;" | ||
| + | |bgcolor="#F8F8F8"|'''Digestion checking:''' | ||
| + | |||
| + | ok | ||
| + | |||
| + | |bgcolor="#E0E3FE"|- | ||
| + | |bgcolor="#CBD1FD"|- | ||
| + | |bgcolor="#BCCDFD"|- | ||
| + | |||
| + | |||
| + | |- style="background: #d8d8d8; text-align: center;" | ||
| + | |bgcolor="#F8F8F8"|'''Ligation:''' | ||
| + | |||
| + | mRFP E/S Ter E/X in Ter vector | ||
| + | |||
| + | |bgcolor="#E0E3FE"|- | ||
| + | |bgcolor="#CBD1FD"|- | ||
| + | |bgcolor="#BCCDFD"| - | ||
| + | |||
| + | |- style="background: #bebebe; text-align: center;" | ||
| + | |bgcolor="#F8F8F8"|'''Colony PCR:''' | ||
| + | |||
| + | ok | ||
| + | |||
| + | |bgcolor="#E0E3FE"|- | ||
| + | |bgcolor="#CBD1FD"|- | ||
| + | |bgcolor="#BCCDFD"|- | ||
| + | |||
| + | |- style="background: #d8d8d8; text-align: center;" | ||
| + | |bgcolor="#F8F8F8"|'''Miniprep:''' | ||
| + | |||
| + | ok | ||
| + | |||
| + | |bgcolor="#E0E3FE"|- | ||
| + | |bgcolor="#CBD1FD"|- | ||
| + | |bgcolor="#BCCDFD"|- | ||
| + | |||
| + | |- style="background: #bebebe; text-align: center;" | ||
| + | |bgcolor="#F8F8F8"|'''Sequencing :''' | ||
| + | ok | ||
| + | |||
| + | |bgcolor="#E0E3FE"| | ||
| + | - | ||
| + | |||
| + | |bgcolor="#CBD1FD"| | ||
| + | - | ||
| + | |bgcolor="#BCCDFD"| | ||
| + | - | ||
| + | |- style="background: #d8d8d8; text-align: center;" | ||
| + | |bgcolor="#F8F8F8"| '''Stock glycerol:''' | ||
| + | |||
| + | - | ||
| + | |bgcolor="#E0E3FE"|- | ||
| + | |bgcolor="#CBD1FD"|- | ||
| + | |bgcolor="#BCCDFD"|- | ||
| + | |} | ||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | <center><font style="color:green;font-weight:bold; font-size:16px">The green color means the experiment was a success</font></center> | ||
| + | |||
| + | |||
| + | |||
| + | [[Image:Paris_construc_6_pfec.jpg |800px|center|plasmid = ]] | ||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | <font style="color:black;font-weight:bold; font-size:16px"><I>'''Experiments ran :'''</I></font> | ||
| + | |||
| + | |||
| + | |||
| + | {| | ||
| + | |- style="background: #0a3585; text-align: center; color:black; font-weight:bold;font-size:14px " | ||
| + | |width="300px" ; bgcolor="#F8F8F8"| Column 1 | ||
| + | |width="300px" ; bgcolor="#E0E3FE"| Column 2 | ||
| + | |width="300px" ; bgcolor="#CBD1FD"| Column 3 | ||
| + | |width="300px" ; bgcolor="#BCCDFD"| Column 4 | ||
| + | |||
| + | |- style="background: #bebebe; text-align: center;" | ||
| + | |bgcolor="#F8F8F8"|'''PCR :''' | ||
| + | |||
| + | pFec matrix: K12 DNA | ||
| + | |||
| + | Oligo: TM | ||
| + | |||
| + | FecI matrix: K12 DNA | ||
| + | |||
| + | Oligo: TM | ||
| + | |||
| + | FecR 1-85 matrix: K12 DNA | ||
| + | |||
| + | Oligo: TM | ||
| + | |||
| + | mRFP matrix: PSB1A3 (previous IGEM plate) | ||
| + | |||
| + | Oligo: TM | ||
| + | |||
| + | |||
| + | |||
| + | |bgcolor="#E0E3FE"|- | ||
| + | |bgcolor="#CBD1FD"|- | ||
| + | |bgcolor="#BCCDFD"|- | ||
| + | |||
| + | |- style="background: #d8d8d8; text-align: center;" | ||
| + | |bgcolor="#F8F8F8"|'''Verification on gel :''' | ||
| + | |||
| + | ok | ||
| + | |||
| + | |bgcolor="#E0E3FE"| | ||
| + | |||
| + | - | ||
| + | |||
| + | |bgcolor="#CBD1FD"| | ||
| + | - | ||
| + | |bgcolor="#BCCDFD"| | ||
| + | - | ||
| + | |||
| + | |- style="background: #bebebe; text-align: center;" | ||
| + | |bgcolor="#F8F8F8"|'''Purification on gel :''' | ||
| + | |||
| + | ok | ||
| + | |||
| + | |bgcolor="#E0E3FE"| | ||
| + | |||
| + | - | ||
| + | |||
| + | |bgcolor="#CBD1FD"| | ||
| + | |||
| + | - | ||
| + | |bgcolor="#BCCDFD"| | ||
| + | |||
| + | - | ||
| + | |||
| + | |||
| + | |||
| + | |- style="background: #d8d8d8; text-align: center;" | ||
| + | |bgcolor="#F8F8F8"|'''Digestion:''' | ||
| + | |||
| + | Ter E/X | ||
| + | |||
| + | mRFP E/S | ||
| + | |||
| + | |bgcolor="#E0E3FE"|- | ||
| + | |bgcolor="#CBD1FD"|- | ||
| + | |bgcolor="#BCCDFD"|- | ||
| + | |||
| + | |- style="background: #bebebe; text-align: center;" | ||
| + | |bgcolor="#F8F8F8"|'''Verification digestion:''' | ||
| + | |||
| + | ok | ||
| + | |||
| + | |bgcolor="#E0E3FE"| | ||
| + | - | ||
| + | |||
| + | |bgcolor="#CBD1FD"| | ||
| + | |||
| + | - | ||
| + | |bgcolor="#BCCDFD"| | ||
| + | |||
| + | - | ||
| + | |||
| + | |||
| + | |- style="background: #d8d8d8; text-align: center;" | ||
| + | |bgcolor="#F8F8F8"|'''Ligation:''' | ||
| + | |||
| + | mRFP E/S Ter E/X in Ter vector | ||
| + | |||
| + | |bgcolor="#E0E3FE"|- | ||
| + | |bgcolor="#CBD1FD"|- | ||
| + | |bgcolor="#BCCDFD"| - | ||
| + | |||
| + | |- style="background: #bebebe; text-align: center;" | ||
| + | |bgcolor="#F8F8F8"|'''Colony PCR:''' | ||
| + | |||
| + | ok | ||
| + | |||
| + | |bgcolor="#E0E3FE"|- | ||
| + | |||
| + | - | ||
| + | |||
| + | |bgcolor="#CBD1FD"|- | ||
| + | |||
| + | - | ||
| + | |bgcolor="#BCCDFD"| | ||
| + | |||
| + | - | ||
| + | |||
| + | |- style="background: #d8d8d8; text-align: center;" | ||
| + | |bgcolor="#F8F8F8"|'''Miniprep:''' | ||
| + | |||
| + | ok | ||
| + | |||
| + | |bgcolor="#E0E3FE"| | ||
| + | |||
| + | - | ||
| + | |||
| + | |bgcolor="#CBD1FD"| | ||
| + | |||
| + | - | ||
| + | |||
| + | |bgcolor="#BCCDFD"| | ||
| + | - | ||
| + | |||
| + | |- style="background: #bebebe; text-align: center;" | ||
| + | |bgcolor="#F8F8F8"|'''Sequencing :''' | ||
| + | ok | ||
| + | |||
| + | |bgcolor="#E0E3FE"| | ||
| + | |||
| + | - | ||
| + | |||
| + | |bgcolor="#CBD1FD"| | ||
| + | - | ||
| + | |bgcolor="#BCCDFD"| | ||
| + | - | ||
| + | |- style="background: #d8d8d8; text-align: center;" | ||
| + | |bgcolor="#F8F8F8"| '''Stock glycerol:''' | ||
| + | |||
| + | - | ||
| + | |bgcolor="#E0E3FE"| | ||
| + | |||
| + | - | ||
| + | |||
| + | |bgcolor="#CBD1FD"| | ||
| + | - | ||
| + | |bgcolor="#BCCDFD"| | ||
| + | |||
| + | - | ||
| + | |} | ||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | <center><font style="color:green;font-weight:bold; font-size:16px">The green color means the experiment was a success</font></center> | ||
| + | |||
| + | |||
| + | [[Image:Paris_construc_2_feca.jpg |800px|center|plasmid = ]] | ||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | <font style="color:black;font-weight:bold; font-size:16px"><I>'''Experiments ran :'''</I></font> | ||
| + | |||
| + | |||
| + | |||
| + | {| | ||
| + | |- style="background: #0a3585; text-align: center; color:black; font-weight:bold;font-size:14px " | ||
| + | |width="300px" ; bgcolor="#F8F8F8"| Column 1 | ||
| + | |width="300px" ; bgcolor="#E0E3FE"| Column 2 | ||
| + | |width="300px" ; bgcolor="#CBD1FD"| Column 3 | ||
| + | |width="300px" ; bgcolor="#BCCDFD"| Column 4 | ||
| + | |||
| + | |- style="background: #bebebe; text-align: center;" | ||
| + | |bgcolor="#F8F8F8"|'''PCR :''' | ||
| + | |||
| + | FecA matrix: K12 | ||
| + | |||
| + | pBAD: | ||
| + | |||
| + | OmpAs: pINR2 | ||
| + | |||
| + | |bgcolor="#E0E3FE"|'- | ||
| + | |bgcolor="#CBD1FD"|- | ||
| + | |bgcolor="#BCCDFD"|- | ||
| + | |||
| + | |- style="background: #d8d8d8; text-align: center;" | ||
| + | |bgcolor="#F8F8F8"|'''Verification on gel :''' | ||
| + | |||
| + | ok | ||
| + | |||
| + | |bgcolor="#E0E3FE"| | ||
| + | |||
| + | - | ||
| + | |||
| + | |bgcolor="#CBD1FD"| | ||
| + | |||
| + | - | ||
| + | |bgcolor="#BCCDFD"|- | ||
| + | |||
| + | - | ||
| + | |||
| + | |- style="background: #bebebe; text-align: center;" | ||
| + | |bgcolor="#F8F8F8"|'''Purification on gel :''' | ||
| + | |||
| + | ok | ||
| + | |||
| + | |bgcolor="#E0E3FE"|- | ||
| + | |||
| + | - | ||
| + | |||
| + | |bgcolor="#CBD1FD"|- | ||
| + | |||
| + | - | ||
| + | |bgcolor="#BCCDFD"|- | ||
| + | |||
| + | - | ||
| + | |||
| + | |||
| + | |||
| + | |- style="background: #d8d8d8; text-align: center;" | ||
| + | |bgcolor="#F8F8F8"|'''Digestion:''' | ||
| + | |||
| + | pBAD E/S | ||
| + | |||
| + | |bgcolor="#E0E3FE"|'''Digestion:''' | ||
| + | none | ||
| + | |bgcolor="#CBD1FD"|- | ||
| + | |bgcolor="#BCCDFD"|- | ||
| + | |||
| + | |- style="background: #bebebe; text-align: center;" | ||
| + | |bgcolor="#F8F8F8"|'''Digestion checking''' | ||
| + | |||
| + | ok | ||
| + | |||
| + | |bgcolor="#E0E3FE"|'''Digestion checking''' | ||
| + | |||
| + | none | ||
| + | |||
| + | |bgcolor="#CBD1FD"| | ||
| + | |||
| + | - | ||
| + | |bgcolor="#BCCDFD"| | ||
| + | - | ||
| + | |||
| + | |||
| + | |- style="background: #d8d8d8; text-align: center;" | ||
| + | |bgcolor="#F8F8F8"|'''Ligation:''' | ||
| + | |||
| + | none | ||
| + | |||
| + | |bgcolor="#E0E3FE"|'''Ligation:''' | ||
| + | none | ||
| + | |bgcolor="#CBD1FD"|- | ||
| + | |bgcolor="#BCCDFD"|- | ||
| + | |||
| + | |- style="background: #bebebe; text-align: center;" | ||
| + | |bgcolor="#F8F8F8"|'''Colony PCR :''' | ||
| + | |||
| + | none | ||
| + | |||
| + | |bgcolor="#E0E3FE"|'''Colony PCR :''' | ||
| + | none | ||
| + | |||
| + | |bgcolor="#CBD1FD"| | ||
| + | |||
| + | - | ||
| + | |bgcolor="#BCCDFD"| | ||
| + | - | ||
| + | |||
| + | |- style="background: #d8d8d8; text-align: center;" | ||
| + | |bgcolor="#F8F8F8"|'''Miniprep''' | ||
| + | |||
| + | none | ||
| + | |||
| + | |bgcolor="#E0E3FE"|'''Miniprep''' | ||
| + | |||
| + | none | ||
| + | |||
| + | |bgcolor="#CBD1FD"| | ||
| + | - | ||
| + | |||
| + | |bgcolor="#BCCDFD"| | ||
| + | |||
| + | - | ||
| + | |||
| + | |- style="background: #bebebe; text-align: center;" | ||
| + | |bgcolor="#F8F8F8"|'''Sequencing''' | ||
| + | none | ||
| + | |||
| + | |bgcolor="#E0E3FE"|'''Sequencing''' | ||
| + | none | ||
| + | |||
| + | |bgcolor="#CBD1FD"| | ||
| + | |||
| + | - | ||
| + | |bgcolor="#BCCDFD"| | ||
| + | - | ||
| + | |- style="background: #d8d8d8; text-align: center;" | ||
| + | |bgcolor="#F8F8F8"| '''Stock glycerol:''' | ||
| + | none | ||
| + | |||
| + | |bgcolor="#E0E3FE"| '''Stock glycerol:''' | ||
| + | none | ||
| + | |||
| + | |bgcolor="#CBD1FD"|- | ||
| + | |bgcolor="#BCCDFD"| | ||
| + | - | ||
| + | |} | ||
| + | |||
| + | |||
| + | |||
| + | <html> | ||
| + | <a href="https://2009.igem.org/Team:Paris/Transduction_testing#bottom"><img style="width:40px; height:40px;" src="https://static.igem.org/mediawiki/2009/1/10/Paris_Up.png"/></a> | ||
| + | </html> | ||
Latest revision as of 03:38, 22 October 2009
iGEM > Paris > WetLab > Reception
Contents |
WetLab - Reception
In this section we have two kinds of experiments :
- Fusion : the construction required for the fusion of the vesicles that contain the message with the receiver.
- Transduction : the construction of the system that will be able de decypher the message.
The first part deals with G3P , OmpAL , Jun, Fos , and AIDA protein. the second with the transduction system of the fec operon (FecA, FecR, FecI).
Fusion
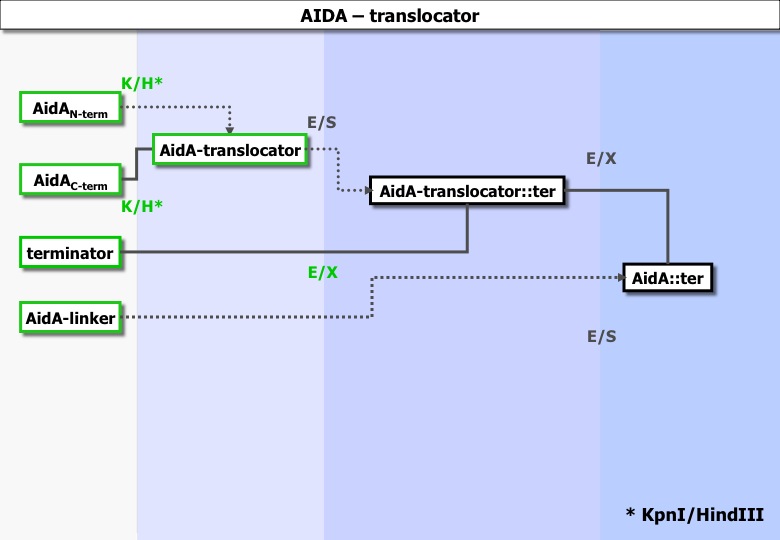
Fusion : the construction required for the fusion of the vesicles that contain the message.
G3P , OmpAL , Jun, Fos , and AIDA protein.
Experiments ran :
| Column 1 | Column 2 | Column 3 | Column 4 |
| PCR :
AidA and Linker : Design, synthesize and order in two parts due to the seize. | - | - | -
|
| Verification on gel :
ok | - | - | - |
| Purification on gel :
ok | - | - | -
|
| Digestion:
AidA CTerm Kpn1/HindIII AidA NTerm Kpn1/HindIII | Digestion:
AidA complete E/P PSB1A3 E/P We did it to obtain a biobrick first | Digestion:
none | -
|
| Digestion checking:
ok | Digestion checking:
ok
| Digestion checking:
none
|
-
|
| Ligation:
AidA CTerm (Kpn1/HindIII):: aidA NTerm (Kpn1/HindIII)
| Ligation:
AidA complete E/P :: PSB1A3 E/P | Ligation:
none | - |
| Colony pCR :
ok | Colony PCR:
ok | Colony PCR :
none | - |
| Miniprep:
done | Miniprep:
done | Miniprep:
none | - |
| Sequencing :
unecessary | Sequencing :
none | Sequencing :
none
| - |
| Stock glycerol:
- | Stock glycerol
S75 | Stock glycerol
none | - |
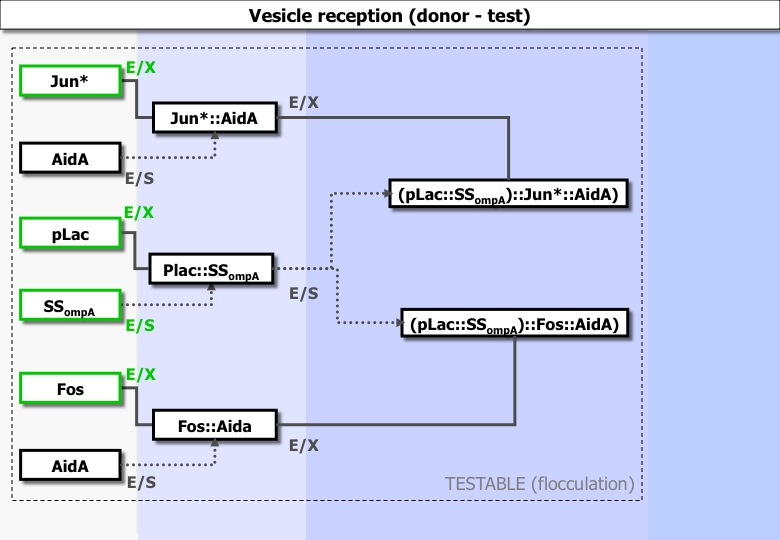
Experiments ran :
| Column 1 | Column 2 | Column 3 | Column 4 |
| PCR :
Jun *: design and order to be a a mutant incapable to form homodimere. pLac : previously done SSOmpA : previously done Fos : previously done | - | - | -
|
| Verification on gel :
ok |
- |
- | - |
| Purification on gel :
ok |
- |
- |
|
| Digestion:
pLac E/X ssOmpA E/S Jun E/X and E/P PSB1A3 E/P | Digestion:
none | - | -
|
| Digestion checking:
ok | Digestion checking:
none |
- |
- |
| Ligation:
pLac on PSB2K3 E/X :: ssOmpA E/S Jun E/P :: PSB1A3 E/P | Ligation:
none | - | '-
|
| Colony PCR:
ok | Colony PCR:
none | -
- | -
- |
| Miniprep:
ok | Miniprep:
none | - | - |
| Sequencing :
-ok | Sequencing :
-none | '- | - |
| Stock glycerol:
none unecessary | Stock glycerol
none | - | -
- |
G3P
Even though it has not been realized, we planned to do these experiments to verify that G3P is expressed into the bacterial surface and induces the membranal fusion.
- Surface expression of G3P
Surface expression of G3P could be seen with antibodies.
Culture of bacteria expressing OmpA-linker-G3P construction on a membrane. Adding antibodies anti-G3P. Revelation should shows whether G3P is expressed at the surface of bacteria.
- Floculation test
"Fusion" could be approached by a flocculation test.
Two separate bacterial population growth on media. A population (F-) expressing the OmpA-linker_G3P construction and a population (F +) which express sexual pilus (by expression of the episome F).
Then we growth bacteria without agitation and we look (compared to culture controls) the speed of sedimentation. Faster sedimentation is due to an interaction between bacterial populations showing an increase of contact time. At vesicle level, it induce membrane fusion.
Transduction
Transduction : the construction of the system that will be able de decypher the message.
The transduction system of the fec operon (FecA, FecR, FecI).
Experiments ran :
| Column 1 | Column 2 | Column 3 | Column 4 |
| PCR :
pFec matrix: K12 DNA Oligo: TM FecI matrix: K12 DNA Oligo: TM FecR 1-85 matrix: K12 DNA Oligo: TM mRFP matrix: PSB1A3 (previous IGEM plate) Oligo: TM
| - | - | - |
| Verification on gel :
ok | - | - | - |
| Purification on gel :
ok | - | - | -
|
| Digestion:
Ter E/X mRFP E/S | - | - | - |
| Digestion checking:
ok | - | - | -
|
| Ligation:
mRFP E/S Ter E/X in Ter vector | - | - | - |
| Colony PCR:
ok | - | - | - |
| Miniprep:
ok | - | - | - |
| Sequencing :
ok |
- |
- |
- |
| Stock glycerol:
- | - | - | - |
Experiments ran :
| Column 1 | Column 2 | Column 3 | Column 4 |
| PCR :
pFec matrix: K12 DNA Oligo: TM FecI matrix: K12 DNA Oligo: TM FecR 1-85 matrix: K12 DNA Oligo: TM mRFP matrix: PSB1A3 (previous IGEM plate) Oligo: TM
| - | - | - |
| Verification on gel :
ok |
- |
- |
- |
| Purification on gel :
ok |
- |
- |
-
|
| Digestion:
Ter E/X mRFP E/S | - | - | - |
| Verification digestion:
ok |
- |
- |
-
|
| Ligation:
mRFP E/S Ter E/X in Ter vector | - | - | - |
| Colony PCR:
ok | -
- | -
- |
- |
| Miniprep:
ok |
- |
- |
- |
| Sequencing :
ok |
- |
- |
- |
| Stock glycerol:
- |
- |
- |
- |
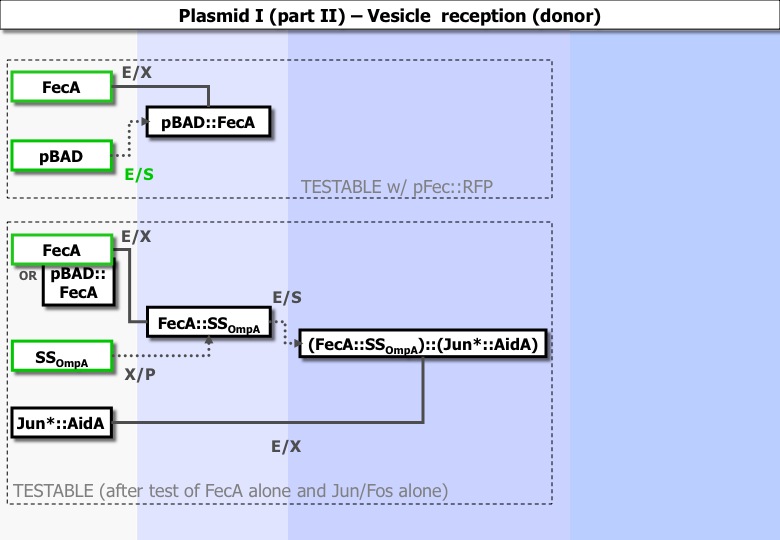
Experiments ran :
| Column 1 | Column 2 | Column 3 | Column 4 |
| PCR :
FecA matrix: K12 pBAD: OmpAs: pINR2 | '- | - | - |
| Verification on gel :
ok |
- |
- | -
- |
| Purification on gel :
ok | -
- | -
- | -
-
|
| Digestion:
pBAD E/S | Digestion:
none | - | - |
| Digestion checking
ok | Digestion checking
none |
- |
-
|
| Ligation:
none | Ligation:
none | - | - |
| Colony PCR :
none | Colony PCR :
none |
- |
- |
| Miniprep
none | Miniprep
none |
- |
- |
| Sequencing
none | Sequencing
none |
- |
- |
| Stock glycerol:
none | Stock glycerol:
none | - |
- |

 "
"