Team:Paris/Production testing
From 2009.igem.org
(→Part II : Functional testing of the part II) |
|||
| (104 intermediate revisions not shown) | |||
| Line 1: | Line 1: | ||
| - | <span/ id="bottom">[https://2009.igem.org/ iGEM ] > [[Team:Paris#top | Paris]] > [[ | + | <span/ id="bottom">[https://2009.igem.org/ iGEM ] > [[Team:Paris#top | Paris]] > [[team:Paris/WetLab#WetLab| WetLab]] > [[Team:Paris/Production_testing#bottom | Production]] |
{{Template:Paris2009}} | {{Template:Paris2009}} | ||
{{Template:Paris2009_menu2}} | {{Template:Paris2009_menu2}} | ||
| - | == WetLab == | + | |
| - | <center> [[ | + | == WetLab - Production == |
| - | + | <html> | |
| + | <style type="text/css"> | ||
| + | #left-side { | ||
| + | position: absolute; | ||
| + | height: 23px; | ||
| + | width: 30px; | ||
| + | top: 0px; | ||
| + | left: 180px; | ||
| + | margin-top:10px; | ||
| + | padding-top: 7px; | ||
| + | background: url(https://static.igem.org/mediawiki/2009/1/1b/Left_menu_pari.png); | ||
| + | z-index:4; | ||
| + | } | ||
| + | |||
| + | #middle-side { | ||
| + | height: 25px; | ||
| + | width: 320px; | ||
| + | position: absolute; | ||
| + | top: 0px; | ||
| + | left: 190px; | ||
| + | margin-top:10px; | ||
| + | padding-top: 5px; | ||
| + | background: #dadada; | ||
| + | z-index:5; | ||
| + | } | ||
| + | |||
| + | #right-side { | ||
| + | position: absolute; | ||
| + | height: 23px; | ||
| + | width: 30px; | ||
| + | margin-top:10px; | ||
| + | padding-top: 7px; | ||
| + | top: 0px; | ||
| + | left: 490px; | ||
| + | background: url(https://static.igem.org/mediawiki/2009/4/40/Right_menu_paris.png); | ||
| + | z-index:4; | ||
| + | } | ||
| + | |||
| + | a.menu_sub { | ||
| + | padding-left: 7px; | ||
| + | padding-right: 7px; | ||
| + | } | ||
| + | |||
| + | a.menu_sub_active { | ||
| + | padding-left: 7px; | ||
| + | padding-right: 7px; | ||
| + | color:#b0310e; | ||
| + | font-weight:bold; | ||
| + | } | ||
| + | </style> | ||
| + | <div id="left-side"></div> | ||
| + | <div id="middle-side"><center> | ||
| + | <a class="menu_sub"href="https://2009.igem.org/Team:Paris/WetLab#bottom"> Main </a>| | ||
| + | <a class="menu_sub" href="https://2009.igem.org/Team:Paris/Addressing_testing#bottom"> Addressing</a>| | ||
| + | <a class="menu_sub_active"href="https://2009.igem.org/Team:Paris/Production_testing#bottom"> Production</a>| | ||
| + | <a class="menu_sub"href="https://2009.igem.org/Team:Paris/Transduction_testing#bottom"> Reception</a> | ||
| + | </center> | ||
| + | </div> | ||
| + | <div id="right-side"></div> | ||
| + | </html> | ||
| + | |||
| + | #[[Team:Paris/Production_testing#The delay | The delay]] | ||
| + | ##[[Team:Paris/Production_testing#Part I : Testing delay phase I| Part I : Testing delay phase I]] | ||
| + | ##[[Team:Paris/Production_testing#Part I : Functional testing of the part I| Part I : Functional testing of the part I]] | ||
| + | ##[[Team:Paris/Production_testing#Part II : Testing delay phase II| Part II : Testing delay phase II]] | ||
| + | ##[[Team:Paris/Production_testing#Part II : Functional testing of the part II| Part II : Functional testing of the part II]] | ||
| + | #[[Team:Paris/Production_testing#The destabilization of the membrane| The destabilization of the membrane]] | ||
| + | |||
<html> | <html> | ||
</div> | </div> | ||
| Line 13: | Line 80: | ||
</html> | </html> | ||
| - | == | + | ==The delay == |
| - | + | We separate the construction on two plasmids : Part I and Part II. | |
| + | ===Part I : Testing delay phase I=== | ||
| - | |||
| - | + | <center><font style="color:green;font-weight:bold; font-size:16px">The green color means the experiment was a success</font></center> | |
| + | [[Image:Paris_construc_3_delay1.jpg |800px|center|plasmid = PSB3T5]] | ||
| - | < | + | |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | <font style="color:black;font-weight:bold; font-size:18px"><I>'''Experiments ran :'''</I></font> | |
| - | + | ||
| - | + | ||
| - | + | {| | |
| - | </ | + | |- style="background: #0a3585; text-align: center; color:black; font-weight:bold;font-size:14px " |
| + | |width="300px" ; bgcolor="#F8F8F8"| Column 1 | ||
| + | |width="300px" ; bgcolor="#E0E3FE"| Column 2 | ||
| + | |width="300px" ; bgcolor="#CBD1FD"| Column 3 | ||
| + | |width="300px" ; bgcolor="#BCCDFD"| Column 4 | ||
| + | |||
| + | |- style="background: #bebebe; text-align: center;" | ||
| + | |bgcolor="#F8F8F8"|'''PCR :''' | ||
| + | |||
| + | pBAD: plate igem | ||
| + | |||
| + | |||
| + | LacI: plate igem | ||
| + | |||
| + | |||
| + | Terminator: plate igem | ||
| + | |||
| + | |||
| + | RBS: plate igem | ||
| + | |||
| + | |||
| + | YFP: plate igem | ||
| + | |||
| + | |bgcolor="#E0E3FE"|- | ||
| + | |bgcolor="#CBD1FD"|- | ||
| + | |bgcolor="#BCCDFD"|- | ||
| + | |||
| + | |- style="background: #d8d8d8; text-align: center;" | ||
| + | |bgcolor="#F8F8F8"|'''Verification on gel :''' | ||
| + | |||
| + | ok | ||
| + | |||
| + | |bgcolor="#E0E3FE"|- | ||
| + | |bgcolor="#CBD1FD"|- | ||
| + | |bgcolor="#BCCDFD"|- | ||
| + | |||
| + | |- style="background: #bebebe; text-align: center;" | ||
| + | |bgcolor="#F8F8F8"|'''Purification on gel :''' | ||
| + | |||
| + | ok | ||
| + | |||
| + | |bgcolor="#E0E3FE"|- | ||
| + | |bgcolor="#CBD1FD"|- | ||
| + | |bgcolor="#BCCDFD"|- | ||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |- style="background: #d8d8d8; text-align: center;" | ||
| + | |bgcolor="#F8F8F8"|'''Digestion:''' | ||
| + | |||
| + | pBAD S/P | ||
| + | |||
| + | |||
| + | lacI E/S | ||
| + | |||
| + | |||
| + | terminator E/X | ||
| + | |||
| + | |||
| + | RBS S/P | ||
| + | |||
| + | |||
| + | YFP X/P | ||
| + | |||
| + | |bgcolor="#E0E3FE"|'''Digestion:''' | ||
| + | pBAD-LacI E/S | ||
| + | |||
| + | |||
| + | RBS-YFP E/S | ||
| + | |||
| + | |bgcolor="#CBD1FD"|'''Digestion:''' | ||
| + | RBS-YFP-ter E/X | ||
| + | |||
| + | |bgcolor="#BCCDFD"|- | ||
| + | |||
| + | |- style="background: #bebebe; text-align: center;" | ||
| + | |bgcolor="#F8F8F8"|'''Verification digestion:''' | ||
| + | |||
| + | ok | ||
| + | |||
| + | |bgcolor="#E0E3FE"|'''Verification digestion:''' | ||
| + | |||
| + | ok | ||
| + | |||
| + | |bgcolor="#CBD1FD"|'''Verification digestion:''' | ||
| + | |||
| + | ok | ||
| + | |||
| + | |bgcolor="#BCCDFD"|- | ||
| + | |||
| + | |||
| + | |- style="background: #d8d8d8; text-align: center;" | ||
| + | |bgcolor="#F8F8F8"|'''Ligation:''' | ||
| + | pBAD-LacI | ||
| + | |||
| + | RBS-YFP | ||
| + | |||
| + | |bgcolor="#E0E3FE"|'''Ligation:''' | ||
| + | |bgcolor="#CBD1FD"|'''Ligation:''' | ||
| + | |bgcolor="#BCCDFD"|'''Ligation:''' | ||
| + | |||
| + | |- style="background: #bebebe; text-align: center;" | ||
| + | |bgcolor="#F8F8F8"|'''Colony PCR :''' | ||
| + | ok | ||
| + | |bgcolor="#E0E3FE"|'''Colony PCR :''' | ||
| + | |||
| + | ok | ||
| + | |||
| + | |bgcolor="#CBD1FD"|'''Colony PCR :''' | ||
| + | |||
| + | ok | ||
| + | |bgcolor="#BCCDFD"|'''Colony PCR:''' | ||
| + | |||
| + | ok | ||
| + | |||
| + | |- style="background: #d8d8d8; text-align: center;" | ||
| + | |bgcolor="#F8F8F8"|'''Miniprep:''' | ||
| + | clone | ||
| + | |||
| + | |bgcolor="#E0E3FE"|'''Miniprep:''' | ||
| + | clone | ||
| + | |bgcolor="#CBD1FD"|'''Miniprep:''' | ||
| + | clone | ||
| + | |||
| + | |bgcolor="#BCCDFD"|'''Miniprep:''' | ||
| + | clone | ||
| + | |- style="background: #bebebe; text-align: center;" | ||
| + | |bgcolor="#F8F8F8"|'''Sequencing :''' | ||
| + | ok | ||
| + | |||
| + | |bgcolor="#E0E3FE"|'''Sequencing :''' | ||
| + | ok | ||
| + | |||
| + | |bgcolor="#CBD1FD"|'''Sequencing :''' | ||
| + | |||
| + | ok | ||
| + | |bgcolor="#BCCDFD"|'''Sequencing :''' | ||
| + | |||
| + | ok | ||
| + | |- style="background: #d8d8d8; text-align: center;" | ||
| + | |bgcolor="#F8F8F8"| '''Stock glycerol:''' | ||
| + | |||
| + | |||
| + | |bgcolor="#E0E3FE"|'''Stock glycerol''' | ||
| + | |||
| + | Cly A(Cter)-(Nter)RFP: S72(clone 8) | ||
| + | |||
| + | |bgcolor="#CBD1FD"|'''Stock glycerol''' | ||
| + | |||
| + | - | ||
| + | |bgcolor="#BCCDFD"|'''Stock glycerol''' | ||
| + | |||
| + | - | ||
| + | |} | ||
| + | |||
| + | ===Part I : Functional testing of the part I=== | ||
| + | |||
| + | PBAD LacI LVA- YFP was transforme into Top ten Bacteria and grew on medium : | ||
| + | |||
| + | |||
| + | 1% arabinose | ||
| + | |||
| + | 1% glucose | ||
| + | |||
| + | LB pure | ||
| + | |||
| + | |||
| + | To learn more, see the [[Team:Paris/Conclusion | Conclusion page]] | ||
| + | |||
| + | |||
| + | |||
| + | |||
| + | <html> | ||
| + | <a href="https://2009.igem.org/Team:Paris/Production_testing#bottom"><img style="width:40px; height:40px;" src="https://static.igem.org/mediawiki/2009/1/10/Paris_Up.png"/></a> | ||
| + | </html> | ||
| + | |||
| + | <html> | ||
| + | </div> | ||
| + | <div id="paris_content_boxtop"> | ||
| + | </div> | ||
| + | <div id="paris_content"> | ||
| + | </html> | ||
| + | |||
| + | ===Part II : Testing delay phase II=== | ||
| + | |||
| + | |||
| + | |||
| + | <center><font style="color:green;font-weight:bold; font-size:16px">The green color means the experiment was a success</font></center> | ||
| + | |||
| + | |||
| + | [[Image:Paris_construc_5_delay2.jpg |800px|center|plasmid = ]] | ||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | <font style="color:black;font-weight:bold; font-size:18px"><I>'''Experiments ran :'''</I></font> | ||
| + | |||
| + | |||
| + | |||
| + | {| | ||
| + | |- style="background: #0a3585; text-align: center; color:black; font-weight:bold;font-size:14px " | ||
| + | |width="300px" ; bgcolor="#F8F8F8"| Column 1 | ||
| + | |width="300px" ; bgcolor="#E0E3FE"| Column 2 | ||
| + | |width="300px" ; bgcolor="#CBD1FD"| Column 3 | ||
| + | |width="300px" ; bgcolor="#BCCDFD"| Column 4 | ||
| + | |||
| + | |- style="background: #bebebe; text-align: center;" | ||
| + | |bgcolor="#F8F8F8"|'''PCR :''' | ||
| + | |||
| + | terminator: igem plate | ||
| + | |||
| + | pLac: igem plate | ||
| + | |||
| + | |||
| + | TetR | ||
| + | |||
| + | |||
| + | pTet: | ||
| + | |||
| + | |||
| + | mRFP | ||
| + | |||
| + | |||
| + | |bgcolor="#E0E3FE"|- | ||
| + | |bgcolor="#CBD1FD"|- | ||
| + | |bgcolor="#BCCDFD"|- | ||
| + | |||
| + | |- style="background: #d8d8d8; text-align: center;" | ||
| + | |bgcolor="#F8F8F8"|'''Verification on gel :''' | ||
| + | |||
| + | ok | ||
| + | |||
| + | |bgcolor="#E0E3FE"|- | ||
| + | |||
| + | - | ||
| + | |||
| + | |bgcolor="#CBD1FD"|- | ||
| + | |||
| + | - | ||
| + | |bgcolor="#BCCDFD"|- | ||
| + | |||
| + | - | ||
| + | |||
| + | |- style="background: #bebebe; text-align: center;" | ||
| + | |bgcolor="#F8F8F8"|'''Purification on gel :''' | ||
| + | |||
| + | ok | ||
| + | |||
| + | |bgcolor="#E0E3FE"| | ||
| + | |||
| + | - | ||
| + | |||
| + | |bgcolor="#CBD1FD"| | ||
| + | |||
| + | - | ||
| + | |bgcolor="#BCCDFD"| | ||
| + | |||
| + | - | ||
| + | |||
| + | |||
| + | |||
| + | |- style="background: #d8d8d8; text-align: center;" | ||
| + | |bgcolor="#F8F8F8"|'''Digestion:''' | ||
| + | |||
| + | pLac S/P | ||
| + | |||
| + | |||
| + | TetR X/P | ||
| + | |||
| + | |||
| + | terminator E/X | ||
| + | |||
| + | |||
| + | pTet S/P | ||
| + | |||
| + | |||
| + | mRFP X/P | ||
| + | |||
| + | |||
| + | pSB3T5 E/P | ||
| + | |||
| + | |bgcolor="#E0E3FE"|'''Digestion:''' | ||
| + | plac-TetR E/S | ||
| + | |||
| + | pTet-mRFP X/P | ||
| + | |||
| + | |bgcolor="#CBD1FD"|'''Digestion:''' | ||
| + | pLac-tetR-ter S/P | ||
| + | |||
| + | |||
| + | |bgcolor="#BCCDFD"|'''Digestion:''' | ||
| + | pLac-tetR-ter-pTet-mRFP E/P | ||
| + | |||
| + | |||
| + | |- style="background: #bebebe; text-align: center;" | ||
| + | |bgcolor="#F8F8F8"|'''Digestion checking:''' | ||
| + | |||
| + | ok | ||
| + | |||
| + | |bgcolor="#E0E3FE"|'''Digestion checking:''' | ||
| + | |||
| + | ok | ||
| + | |||
| + | |bgcolor="#CBD1FD"|'''Digestion checking:''' | ||
| + | |||
| + | ok | ||
| + | |bgcolor="#BCCDFD"|'''Digestion checking:''' | ||
| + | |||
| + | ok | ||
| + | |||
| + | |||
| + | |- style="background: #d8d8d8; text-align: center;" | ||
| + | |bgcolor="#F8F8F8"|'''Ligation:''' | ||
| + | |||
| + | pLac-tetR | ||
| + | |||
| + | |||
| + | pTet-mRFP | ||
| + | |||
| + | |||
| + | |bgcolor="#E0E3FE"|'''Ligation:''' | ||
| + | |||
| + | pLac-TetR-ter | ||
| + | |bgcolor="#CBD1FD"|'''Ligation:''' | ||
| + | |||
| + | pLac-ter-pTet-mRFP | ||
| + | |||
| + | |bgcolor="#BCCDFD"|'''Ligation:''' | ||
| + | |||
| + | pLac-ter-pTet-mRFP-pSB3T5 | ||
| + | |||
| + | |||
| + | |- style="background: #bebebe; text-align: center;" | ||
| + | |bgcolor="#F8F8F8"|'''Colony PCR:''' | ||
| + | |||
| + | ok | ||
| + | |||
| + | |bgcolor="#E0E3FE"|'''Colony PCR :''' | ||
| + | |||
| + | ok | ||
| + | |||
| + | |bgcolor="#CBD1FD"|'''Colony PCR:''' | ||
| + | |||
| + | ok | ||
| + | |bgcolor="#BCCDFD"|'''Colony PCR :''' | ||
| + | |||
| + | ok | ||
| + | |||
| + | |- style="background: #d8d8d8; text-align: center;" | ||
| + | |bgcolor="#F8F8F8"|'''Miniprep:''' | ||
| + | |||
| + | ok | ||
| + | |||
| + | |bgcolor="#E0E3FE"|'''Miniprep:''' | ||
| + | |||
| + | ok | ||
| + | |||
| + | |bgcolor="#CBD1FD"|'''Miniprep:''' | ||
| + | |||
| + | ok | ||
| + | |||
| + | |bgcolor="#BCCDFD"|'''Miniprep:''' | ||
| + | |||
| + | ok | ||
| + | |||
| + | |- style="background: #bebebe; text-align: center;" | ||
| + | |bgcolor="#F8F8F8"|'''Sequencing :''' | ||
| + | |||
| + | ok | ||
| + | |||
| + | |bgcolor="#E0E3FE"|'''Sequencing :''' | ||
| + | |||
| + | ok | ||
| + | |||
| + | |bgcolor="#CBD1FD"|'''Sequencing :''' | ||
| + | |||
| + | ok | ||
| + | |bgcolor="#BCCDFD"|'''Sequencing :''' | ||
| + | |||
| + | ok | ||
| + | |- style="background: #d8d8d8; text-align: center;" | ||
| + | |bgcolor="#F8F8F8"| '''Stock glycerol:''' | ||
| + | |||
| + | - | ||
| + | |bgcolor="#E0E3FE"|'''Stock glycerol''' | ||
| + | |||
| + | - | ||
| + | |bgcolor="#CBD1FD"|'''Stock glycerol''' | ||
| + | |||
| + | - | ||
| + | |bgcolor="#BCCDFD"|'''Stock glycerol''' | ||
| + | |||
| + | - | ||
| + | |} | ||
| + | |||
| + | ===Part II : Functional testing of the part II=== | ||
| + | |||
| + | Ptet RFP Plac TetR was tranforme into Top10 bacteria and grew on media : | ||
| + | |||
| + | |||
| + | 0,1% IPTG . | ||
| + | |||
| + | 1% glucose. | ||
| + | |||
| + | LB pure. | ||
| + | |||
| + | |||
| + | To learn more, see the [[Team:Paris/Conclusion | Conclusion page]] | ||
| + | |||
| + | <html> | ||
| + | <a href="https://2009.igem.org/Team:Paris/Production_testing#bottom"><img style="width:40px; height:40px;" src="https://static.igem.org/mediawiki/2009/1/10/Paris_Up.png"/></a> | ||
| + | </html> | ||
| + | |||
| + | |||
| + | <html> | ||
| + | </div> | ||
| + | <div id="paris_content_boxtop"> | ||
| + | </div> | ||
| + | <div id="paris_content"> | ||
| + | </html> | ||
| + | |||
| + | ==The destabilization of the membrane== | ||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | <center><font style="color:green;font-weight:bold; font-size:16px">The green color means the experiment was a success</font></center> | ||
| + | |||
| + | |||
| + | [[Image:Paris_construc_4_tr2.jpg |800px|center|plasmid = PSB3T5]] | ||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | <font style="color:black;font-weight:bold; font-size:18px"><I>'''Experiments ran :'''</I></font> | ||
| + | |||
| + | |||
| + | {| | ||
| + | |- style="background: #0a3585; text-align: center; color:black; font-weight:bold;font-size:14px " | ||
| + | |width="300px" ; bgcolor="#F8F8F8"| Column 1 | ||
| + | |width="300px" ; bgcolor="#E0E3FE"| Column 2 | ||
| + | |width="300px" ; bgcolor="#CBD1FD"| Column 3 | ||
| + | |width="300px" ; bgcolor="#BCCDFD"| Column 4 | ||
| + | |||
| + | |- style="background: #bebebe; text-align: center;" | ||
| + | |bgcolor="#F8F8F8"|'''PCR :''' | ||
| + | |||
| + | pTet: igem plate | ||
| + | |||
| + | OmpAs: | ||
| + | ToRII: pINK2 | ||
| + | g3p: pINTg3p | ||
| + | |||
| + | |||
| + | |bgcolor="#E0E3FE"| | ||
| + | |bgcolor="#CBD1FD"|' | ||
| + | |bgcolor="#BCCDFD"| | ||
| + | |||
| + | |- style="background: #d8d8d8; text-align: center;" | ||
| + | |bgcolor="#F8F8F8"|'''Verification on gel :''' | ||
| + | |||
| + | ok | ||
| + | |||
| + | |bgcolor="#E0E3FE"|- | ||
| + | |bgcolor="#CBD1FD"|- | ||
| + | |bgcolor="#BCCDFD"|- | ||
| + | |||
| + | |||
| + | |- style="background: #bebebe; text-align: center;" | ||
| + | |bgcolor="#F8F8F8"|'''Purification on gel :''' | ||
| + | ok | ||
| + | |bgcolor="#E0E3FE"|- | ||
| + | |bgcolor="#CBD1FD"|- | ||
| + | |bgcolor="#BCCDFD"|- | ||
| + | |||
| + | |||
| + | |||
| + | |- style="background: #d8d8d8; text-align: center;" | ||
| + | |bgcolor="#F8F8F8"|'''Digestion:''' | ||
| + | |||
| + | pTet S/P | ||
| + | |||
| + | OmpAs X/P | ||
| + | |||
| + | |||
| + | TolRII X/P | ||
| + | |||
| + | |||
| + | g3p X/P | ||
| + | |||
| + | |||
| + | |||
| + | |bgcolor="#E0E3FE"|'''Digestion:''' | ||
| + | pTet-OmpAs S/P | ||
| + | |bgcolor="#CBD1FD"|'''Digestion:''' | ||
| + | none | ||
| + | |||
| + | |bgcolor="#BCCDFD"|- | ||
| + | |||
| + | |||
| + | |- style="background: #bebebe; text-align: center;" | ||
| + | |bgcolor="#F8F8F8"|'''Digestion checking:''' | ||
| + | |||
| + | ok | ||
| + | |||
| + | |bgcolor="#E0E3FE"|'''Digestion checking:''' | ||
| + | |||
| + | ok | ||
| + | |||
| + | |bgcolor="#CBD1FD"|'''Digestion checking:''' | ||
| + | |||
| + | none | ||
| + | |bgcolor="#BCCDFD"|- | ||
| + | |||
| + | |||
| + | |- style="background: #d8d8d8; text-align: center;" | ||
| + | |bgcolor="#F8F8F8"|'''Ligation:''' | ||
| + | pTet-OmpAs | ||
| + | |||
| + | |bgcolor="#E0E3FE"|'''Ligation:''' | ||
| + | none | ||
| + | |||
| + | |||
| + | |bgcolor="#CBD1FD"|'''Ligation:''' | ||
| + | none | ||
| + | |bgcolor="#BCCDFD"|- | ||
| + | |||
| + | |- style="background: #bebebe; text-align: center;" | ||
| + | |bgcolor="#F8F8F8"|'''Colony PCR :''' | ||
| + | |||
| + | ok | ||
| + | |||
| + | |bgcolor="#E0E3FE"|'''Colony PCR:''' | ||
| + | |||
| + | none | ||
| + | |||
| + | |bgcolor="#CBD1FD"|'''Colony PCR:''' | ||
| + | |||
| + | none | ||
| + | |bgcolor="#BCCDFD"|- | ||
| + | |||
| + | |- style="background: #d8d8d8; text-align: center;" | ||
| + | |bgcolor="#F8F8F8"|'''Miniprep:''' | ||
| + | |||
| + | ok | ||
| + | |||
| + | |bgcolor="#E0E3FE"|'''Miniprep:''' | ||
| + | |||
| + | none | ||
| + | |||
| + | |bgcolor="#CBD1FD"|'''Miniprep:''' | ||
| + | |||
| + | none | ||
| + | |||
| + | |bgcolor="#BCCDFD"|- | ||
| + | |||
| + | |- style="background: #bebebe; text-align: center;" | ||
| + | |bgcolor="#F8F8F8"|'''Sequencing :''' | ||
| + | |||
| + | ok | ||
| + | |||
| + | |bgcolor="#E0E3FE"|'''Sequencing :''' | ||
| + | |||
| + | none | ||
| + | |||
| + | |bgcolor="#CBD1FD"|'''Sequencing :''' | ||
| + | |||
| + | none | ||
| + | |bgcolor="#BCCDFD"|- | ||
| + | |||
| + | |||
| + | |- style="background: #d8d8d8; text-align: center;" | ||
| + | |bgcolor="#F8F8F8"| '''Stock glycerol:''' | ||
| + | |||
| + | - | ||
| + | |bgcolor="#E0E3FE"|'''Stock glycerol''' | ||
| + | |||
| + | none | ||
| + | |||
| + | |bgcolor="#CBD1FD"|'''Stock glycerol''' | ||
| + | |||
| + | none | ||
| + | |bgcolor="#BCCDFD"|- | ||
| + | |} | ||
| + | |||
| + | |||
| + | |||
| + | |||
| + | <html> | ||
| + | <a href="https://2009.igem.org/Team:Paris/Production_testing#bottom"><img style="width:40px; height:40px;" src="https://static.igem.org/mediawiki/2009/1/10/Paris_Up.png"/></a> | ||
| + | </html> | ||
Latest revision as of 03:58, 22 October 2009
iGEM > Paris > WetLab > Production
The delay
We separate the construction on two plasmids : Part I and Part II.
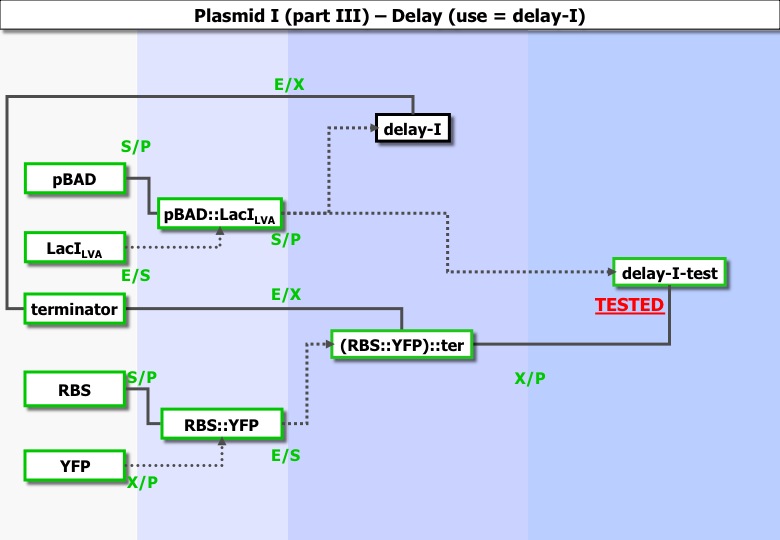
Part I : Testing delay phase I
Experiments ran :
| Column 1 | Column 2 | Column 3 | Column 4 |
| PCR :
pBAD: plate igem
| - | - | - |
| Verification on gel :
ok | - | - | - |
| Purification on gel :
ok | - | - | -
|
| Digestion:
pBAD S/P
| Digestion:
pBAD-LacI E/S
| Digestion:
RBS-YFP-ter E/X | - |
| Verification digestion:
ok | Verification digestion:
ok | Verification digestion:
ok | -
|
| Ligation:
pBAD-LacI RBS-YFP | Ligation: | Ligation: | Ligation: |
| Colony PCR :
ok | Colony PCR :
ok | Colony PCR :
ok | Colony PCR:
ok |
| Miniprep:
clone | Miniprep:
clone | Miniprep:
clone | Miniprep:
clone |
| Sequencing :
ok | Sequencing :
ok | Sequencing :
ok | Sequencing :
ok |
| Stock glycerol:
| Stock glycerol
Cly A(Cter)-(Nter)RFP: S72(clone 8) | Stock glycerol
- | Stock glycerol
- |
Part I : Functional testing of the part I
PBAD LacI LVA- YFP was transforme into Top ten Bacteria and grew on medium :
1% arabinose
1% glucose
LB pure
To learn more, see the Conclusion page
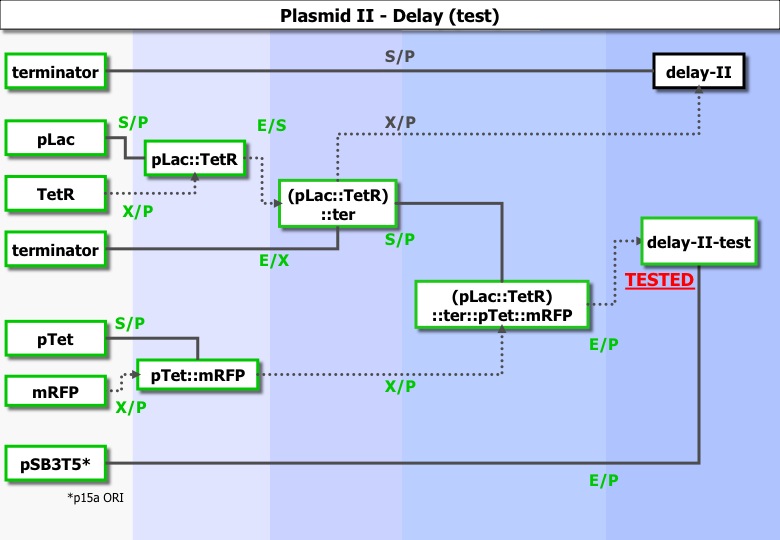
Part II : Testing delay phase II
Experiments ran :
| Column 1 | Column 2 | Column 3 | Column 4 |
| PCR :
terminator: igem plate pLac: igem plate
| - | - | - |
| Verification on gel :
ok | -
- | -
- | -
- |
| Purification on gel :
ok |
- |
- |
-
|
| Digestion:
pLac S/P
| Digestion:
plac-TetR E/S pTet-mRFP X/P | Digestion:
pLac-tetR-ter S/P
| Digestion:
pLac-tetR-ter-pTet-mRFP E/P
|
| Digestion checking:
ok | Digestion checking:
ok | Digestion checking:
ok | Digestion checking:
ok
|
| Ligation:
pLac-tetR
| Ligation:
pLac-TetR-ter | Ligation:
pLac-ter-pTet-mRFP | Ligation:
pLac-ter-pTet-mRFP-pSB3T5
|
| Colony PCR:
ok | Colony PCR :
ok | Colony PCR:
ok | Colony PCR :
ok |
| Miniprep:
ok | Miniprep:
ok | Miniprep:
ok | Miniprep:
ok |
| Sequencing :
ok | Sequencing :
ok | Sequencing :
ok | Sequencing :
ok |
| Stock glycerol:
- | Stock glycerol
- | Stock glycerol
- | Stock glycerol
- |
Part II : Functional testing of the part II
Ptet RFP Plac TetR was tranforme into Top10 bacteria and grew on media :
0,1% IPTG .
1% glucose.
LB pure.
To learn more, see the Conclusion page
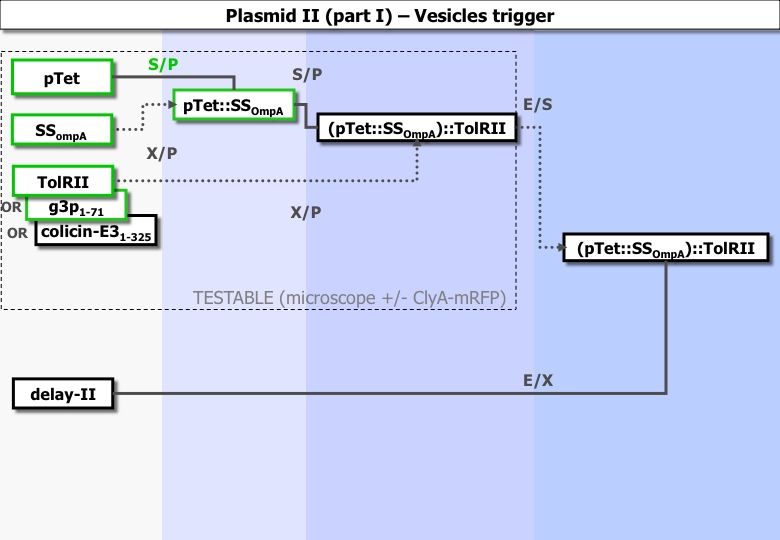
The destabilization of the membrane
Experiments ran :
| Column 1 | Column 2 | Column 3 | Column 4 |
| PCR :
pTet: igem plate OmpAs: ToRII: pINK2 g3p: pINTg3p
| ' | ||
| Verification on gel :
ok | - | - | -
|
| Purification on gel :
ok | - | - | -
|
| Digestion:
pTet S/P OmpAs X/P
| Digestion:
pTet-OmpAs S/P | Digestion:
none | -
|
| Digestion checking:
ok | Digestion checking:
ok | Digestion checking:
none | -
|
| Ligation:
pTet-OmpAs | Ligation:
none
| Ligation:
none | - |
| Colony PCR :
ok | Colony PCR:
none | Colony PCR:
none | - |
| Miniprep:
ok | Miniprep:
none | Miniprep:
none | - |
| Sequencing :
ok | Sequencing :
none | Sequencing :
none | -
|
| Stock glycerol:
- | Stock glycerol
none | Stock glycerol
none | - |

 "
"