Team:Paris/5 August 2009
From 2009.igem.org
(Difference between revisions)
Christophe.R (Talk | contribs) (→Microscope) |
(→Microscope) |
||
| Line 19: | Line 19: | ||
{| | {| | ||
|- style="background: #7B96B3; " | |- style="background: #7B96B3; " | ||
| - | ! colspan="11" style="background: # | + | ! colspan="11" style="background: #B40ECD;border:1px solid #333333 "|FM4-64 dying (wash then stain) strain MG4 |
|- style="background: #b3d0E4; text-align: center; color:black; " | |- style="background: #b3d0E4; text-align: center; color:black; " | ||
|width=10%| n°1 | |width=10%| n°1 | ||
| Line 72: | Line 72: | ||
{| | {| | ||
|- style="background: #7B96B3; " | |- style="background: #7B96B3; " | ||
| - | ! colspan="11" style="background: # | + | ! colspan="11" style="background: #B40ECD;border:1px solid #333333 "|FM4-64 dying (stain then wash) |
|- style="background: #b3d0E4; text-align: center; color:black; " | |- style="background: #b3d0E4; text-align: center; color:black; " | ||
|width=10%| n°5 | |width=10%| n°5 | ||
Revision as of 21:45, 7 August 2009
Contents |
NoteBook
|
|
|
|
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
Lab work
Microscope
| FM4-64 dying (wash then stain) strain MG4 | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| n°1 | n°2 | n°3 | n°4 | |||||||
| 500µl cells | - | - | - | |||||||
| centri 4000rpm 3min | - | - | - | |||||||
| resuspension 500µl MgSO4 | - | - | - | |||||||
| 1µl de colorant | 1µl de colorant | 2µl | 2µl | |||||||
| incubation 30min 37°C | incubation 60min 37°C | incubation 30min 37°C | incubation 60min 37°C | |||||||
| depot sur lame | - | - | - | |||||||
| nothing | all cell fluo | so much dye | so much dye | |||||||
| FM4-64 dying (stain then wash) | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| n°5 | n°6 | n°9 | n°10 | n°13 | n°14 | |||||
| 500µl cells | - | - | - | 500µl MgSO4 | 500µl Medium | |||||
| 1µl de colorant | 1µl de colorant | 1µl | 1µl | 1µl | 1µl | |||||
| incubation 30min 37°C | incubation 60min 37°C | incubation 30min 37°C | incubation 60min 37°C | incubation 60min 37°C | incubation 60min 37°C | |||||
| centri 4000rpm 3min | - | - | - | |||||||
| resuspension 500µl MgSO4 | resuspension 500µl MgSO4 | + filtrage | + filtrage | |||||||
| depot sur lame | - | - | - | - | - | |||||
| nothing | some cell fluo | noise | noise | noise | noise | |||||
FM4-64 dying with Kayo
protocol like n°2 = all cell fluo, no vesicle
protocol like n°6 = some cell fluo, noise, no vesicle
Molecular biology
Glycerol Stock
- [S20] JW4251, [S21] JN0729, [S22] JN0728, [S23] JN0727, [S24] JN0940, [S24] JN0940, [S25] JW5100, [S25] JW5100, [S26] JN5181, [S27] FF64, [S28] NEC280.
Miniprep
- Miniprep for P1/2/3/4/5/7/8
- Same protocol as yesterday. Storage at -20°C.
Purification
- Purification (using 1%agarose at 30min migration) of [P2]=BBa-J61002 with XbaI PstI (put pTet forward).
- D6=2053pb=[P2]Digested
- [ladder 1kb/[P2]/nothing/ladder 100pb]
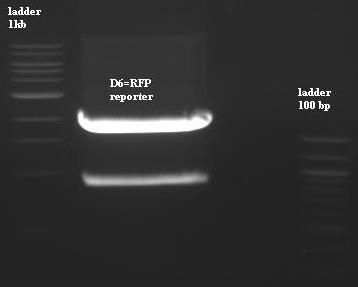

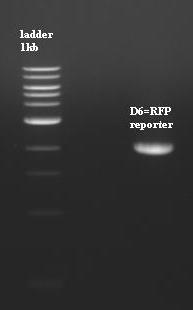
- The purification is ok but Ddon't use because it's doesn't have pTet, so we have to cut [P2] with EcorI and SpeI next time.
PCR
- Dilution oligo 1/10 (100µM to 10µM)
- 2µl Oligo
- 18µL H20 RNAse/DNAse free (Gibco)
- Dilution DNA 1/10
- 2µl Genomic DNA of K12 MG1655 ΔrecA::Kan [F4]
- 18µL H20 RNAse/DNAse free (Gibco)
- Mix x1 Vf=50µl in PCR tube (in ice) (Add with this order)
- 32,5µl H20 RNAse/DNAse free (Gibco)
- 10µl Buffer Polymerase fusion 5x
- 1µl dNTP
- 2,5µl Oligo F 10µM
- 2,5µl Oligo R 10µM
- 1µl Genomic DNA of K12 1/10
- 0,5µl Polymerase Fusion
- Mix x6,5
- 211,25µl H20 RNAse/DNAse free (Gibco)
- 65µl Buffer Polymerase fusion 5x
- 6,5µl dNTP
- 6,5µl Genomic DNA of K12 1/10
- Put 44,5µl of Mix x6,5 in a PCR tube
- Add 2,5µl Oligo F 10µM
- Add 2,5µl Oligo R 10µM
- Add 0,5µl Polymerase Fusion
- For A1/2/3/4/5/6, Program PHUSION:
- 1: 98°C, 1min
- 2: 98°C, 10sec
- 3: 65°C, 30sec
- 4: 72°C, 1min
- Goto 2 29x
- 72°C, 10min
- 4°C, -
Gel migration
*Gel 1%: 5µl of [A1|A2|100bp|A3|A4|1kb|A5|A6]


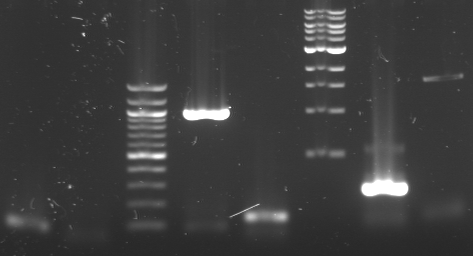
- A1 = Amplification of N-term domain of M13's g3p = 274bp (Useless because we don't use g3p plasmide as matrice => Ctrl-)
- A2 = Amplification of N-term domain of colicin E3 = 1036bp (Useless because we don't use the right matrice => Ctrl-)
- A3 = Amplification of clyA with RBS in the primer = 987bp
- A4 = Amplification of OmpA signal for export to periplasm = 135bp
- A5 = Amplification of tolR = 279bp (Try again because of the second band)
- A6 = Amplification of tatA with RBS = 1634bp (Try again because of the second band)



ON culture
- [S8] DH5α(pSB2K3): LB Kan
- [29] DH5α(pSB1A3): LB Amp
- [30] S113 iGEM08 = DH5α(BBa_K136050): LB Amp
To do list
| Matricule | TODO |
| Luc | bacteria coloration microscope protocol |
| Romain | Look for his feet |
| Charlotte | bibliography on SNAREs/Autotransporters/Jun and Fos complex |
| Stoff | merge algo protocol/ Wiki |
| Chris | modeling: TeTR |
| Lisa | WTF !!! |
| Caroline | bacteria coloration microscope protocol |
| Souf | if(!oligo){return Wiki;} return PCR; |
| Vicard | Lab : Gel / purification / insers |
| Pierre | Alice / Tol/Pal modelisation / geophysics |
| Sylvain | if(!oligo){return Maltose;} return PCR; |
| Guillaume | Miniprep for P(1/2/3/4/5/7/8) / New P7 digestion? / Glycerol Stock (DH5α/Kaio strains) |
 "
"