Team:Paris/Transduction modeling The Fec Operon as used in our system : chemical equations and kinetics
From 2009.igem.org
(→Chemical Equations) |
(→Chemical Equations) |
||
| Line 121: | Line 121: | ||
This trancriptional cacade can be described thanks to simple chemical equations traducing the chronological and chemical steps of our tranduction and activation system. | This trancriptional cacade can be described thanks to simple chemical equations traducing the chronological and chemical steps of our tranduction and activation system. | ||
| - | [Image:Fec Complex.png|500px|center] | + | [[Image:Fec Complex.png|500px|center]] |
Revision as of 20:46, 21 October 2009
iGEM > Paris > Reception > Modeling
Contents |
Fec operon in our system
To try and understand our system, we decided to divide the global in simple reactions traducing the chemical steps of the transduction cascade ; each one of these reactions is described with a cinetic law, thus allowing to run both deterministic and stochastic simulation (see here for further informations on these simulations).
Chemical Equations
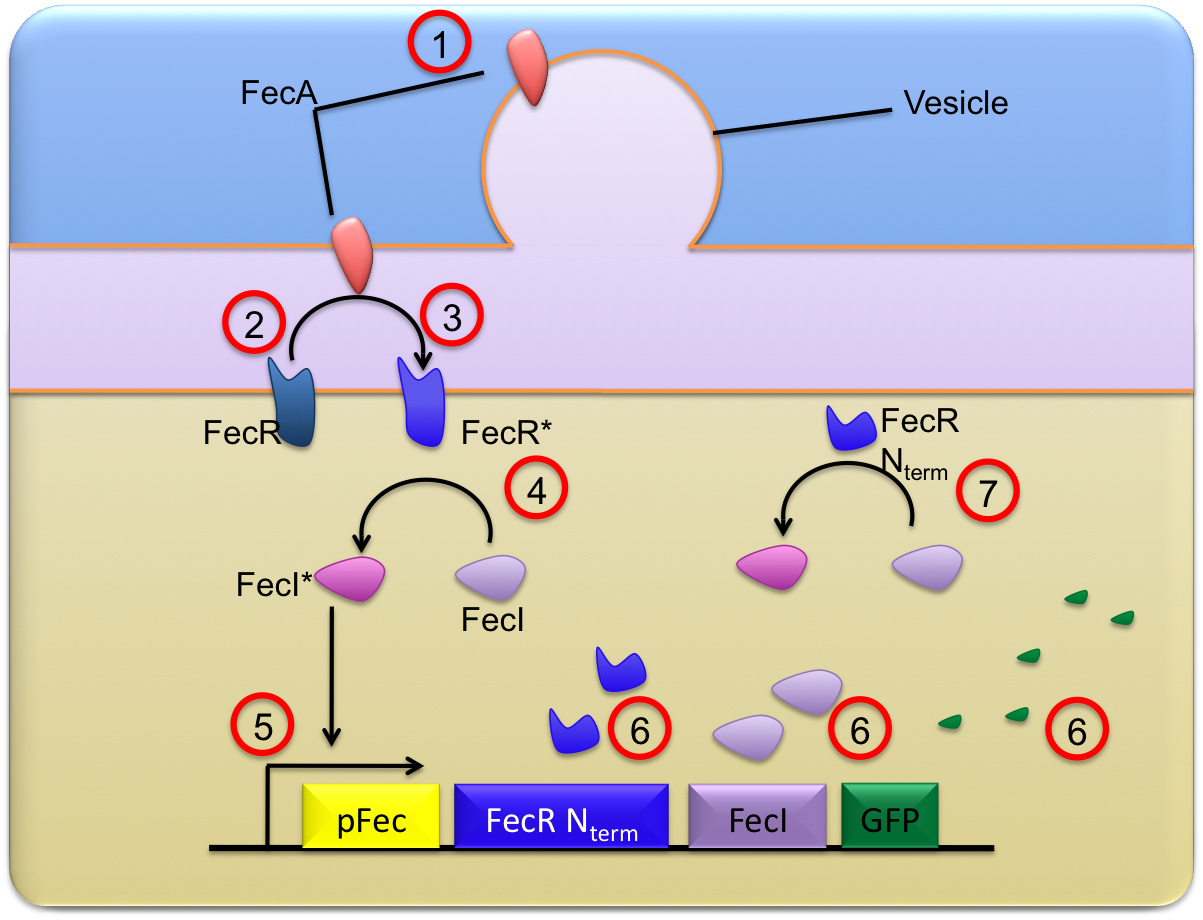
This trancriptional cacade can be described thanks to simple chemical equations traducing the chronological and chemical steps of our tranduction and activation system.
- The FecA molecules coming from the vesicules are clowly diffusing on the lipid bilayer of the outer membrane while vesicules are fusioning ; they finally reach the bacteria outer membrane where they are in proper conditions to be able to activate the FeR molecules. This "crossing" from OMV to the outer membrane is explicited as a first reaction :
- Then, once in the outer membrane, these FecA molecules are able to activate the FecR proteins constitutively present in the receiver. Here we have two possibilities to describe this step :
- the FecA molecule directly activates FecR, and the phnomenon is described in a single reaction :
- or FecA and FecR form a complex which is then destructed to release an activated FecR protein ; this mechanism is described through the 2 following reactions :
As an hypothesis, we considered that this crossing of the periplasm was the limiting step, we decided to examine these two different approaches in our simulations.
- Once FecR is activated, it can activate the FecI molecule ; to avoid an to much variability in the approaches, we decided to consider that this reaction occured as a single step (with no intermediate complex formation) :
- Then, FecI* can activate the pfec promoter in a reversible reaction :
- After the pfec activation, transcription and translation can start ; the creation of proteins is materialised by the single reaction :
where FecR_Nterm correspond to the residues ??-??? of FecR which make this molecule constitutively active (the FecR_Nterm is able to activate FecI without needing to be in presence of FecA)
- Finally, the constitutively active FecR_Nterm can activate the FecI proteins :
- It is also compulsory to consider some dilution and degradation reactions for the molecules FecA_OM, FecI, FecI*, FecR_Nterm, FecR* and GFP. The natural FecR is considered as a molecule having reached a steady state level ; as a consequence the only noticeable changes are the results of a reaction of activation.
Kinetics Equations
As we do not have much information on the kinetic laws, we have decided to chose a mass action law for each reaction. Each reaction has a kinetic constant k determining the reaction rate given the concentratin of reactants. This will lead us to write a differential system used for a deterministic resolution.
The first thing to do is to write the reaction rate of each chemical step for the two systems of reactions :
- A system without an intermediary step in the FecR activation
| Reaction Number | Kinetics Constants | Reaction | Reaction Rate |
| 1 | kf_1 | FecA_OMV ---> FecA_OM | kf_1*[FecA_OMV] |
| 2 | kf_2 | FecA_OM + FecR ---> FecA_OM + FecR* | kf_2*[FecA_OM]*[FecR] |
| 3 | kf_3 | FecR* + FecI ---> FecI* + FecR* | kf_3*[FecI*]*[FecR*] |
| 4 | kf_4 | FecI* + pfec <---> pfec* | kf_4*[FecI*]*[pfec]-kr_4[pfec*] |
| 5 | kf_5 | pfec* ---> FecI + FecR_Nterm + GFP | kf_5*[pfec*] |
| 6 | kf_6 | FecR_Nterm + FecI ---> FecI* + FecR_Nterm | kf_6*[FecR_Nterm]*[FecI] |
| 7 | kf_7 | FecA_OM ---> ∅ | kf_7*[FecA_OM] |
| 8 | kf_8 | FecI ---> ∅ | kf_8*[FecI] |
| 9 | kf_9 | FecI* ---> ∅ | kf_9*[FecI*] |
| 10 | kf_10 | FecR_Nterm ---> ∅ | kf_10*[FecR_Nterm] |
| 11 | kf_11 | FecR* ---> ∅ | kf_11*[FecR*] |
| 12 | kf_12 | GFP ---> ∅ | kf_12*[GFP] |
- A system of equations with an intermediary reaction in the activation of FecR
| Reaction Number | Kinetics Constants | Reaction | Reaction Rate |
| 1 | kf_1 | FecA_OMV ---> FecA_OM | kf_1*[FecA_OMV] |
| 2 | kf_2 & kr_2 | FecA_OM + FecR <---> FecA_OM-FecR | kf_2*[FecA_OM]*[Fec_R] - kr_2*[FecA_OM-FecR] |
| 3 | kf_3 | FecA_OM-FecR ---> FecA_OM + FecR* | kf_3*[FecA_OM-FecR] |
| 4 | kf_4 | FecR* + FecI ---> FecI* + FecR* | kf_4*[FecR*]*[FecI] |
| 5 | kf_5 & kr_5 | FecI* + pfec <---> pfec* | kf_5*[FecI*]*[pfec] - kr_5*[pfec*] |
| 6 | kf_6 | pfec* ---> FecI + FecR_Nterm + GFP | kf_6*[pfec*] |
| 7 | kf_7 | FecR_Nterm + FecI ---> FecI* + FecR_Nterm | kf_7*[FecR_Nterm]*[FecI] |
| 8 | kf_8 | FecA_OM ---> ∅ | kf_8*[FecA_OM] |
| 9 | kf_9 | FecI ---> ∅ | kf_9*[FecI] |
| 10 | kf_10 | FecI* ---> ∅ | kf_10*[FecI*] |
| 11 | kf_11 | FecR_Nterm ---> ∅ | kf_11*[FecR_Nterm] |
| 12 | kf_12 | FecR* ---> ∅ | kf_12*[FecR*] |
| 13 | kf_13 | GFP ---> ∅ | kf_13*[GFP] |
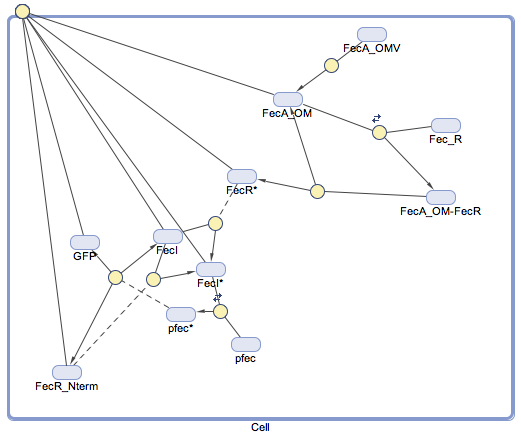
These notations were used in the Simbiology toolbox of Matlab to perform all the stochastic simulations ; the diagram of reactions is presented below (for a complexaion between FecA and FecR) :
Result Plots
Here are presented some results from the stochastic simulations
Draft
For a given component A , which takes part in reations R1, R2, ...Rn, we can write that :
where the δi are equal to the stoechiometric coefficients of the species A in the reaction i and Vi is the reaction rate of reaction i. Given this consideratin and using the tables of reactions, we can start writing down the differential equations ruling the evolution of the concentrations :
 "
"