Team:Paris/15 August 2009
From 2009.igem.org
Contents |
NoteBook
|
|
|
|
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
Brain work
edit please ^^
Lab work
Transformation
- For overnight (at 16°C) ligation solution of g3p in pSB1A3
- For overnight (at 16°C) ligation solution of ClyA in pSB1A3
Miniprep
- ptet
- pBad/araC
- Double terminater
Glycerol Stock
- New strains
- [S35] AA93 ΔfecA
- [S36] fecIR mutant
- [S37] fecIR wt
- [S38] fecA
Yesterday transformations results
- tatABC
- No bacteria
- pFecA
- Negative control = 0 bacteria
- 1:1 raport = 43 bacteria
- 1:7 raport = 2 bacteria
- 1:7 concentred = Too much
- g3p
- Negative control = 0 bacteria
- 1:1 raport = 1 bacteria
- 1:7 raport = 0 bacteria
- 1:7 concentred = 16 bacteria
PCR screening protocol
- Program named COLONIES, use with VF2 and VR oligos
- Lid 105°C
- 1: 99°C 2min
- 2: Sound
- 3: Pause until press "Enter" button
- 4: 95°C 5min
- 5: 95°C 30sec
- 6: 55°C 30sec
- 7: 72°C [45sec] (1kb/bp, caution the universal oligo add 156bp)
- 8: Goto 5 29x
- 9:72°C 7min
- 10: Hold at 4°C
- Preparation of the ON culture (for Glycerol stock and Miniprep of the positives clones)
- 7ml of LB+ Amp (because we use pSB1A3 as vector)
- Bacterial solution
- 100uL of H2O and few bacteria in a PCR tube
- Put the cone in LB solution for ON culture
- Put the PR tube in PCR and run the fist part of the program
- PCR mix
- Master Mix (2x) : 12,5ul
- Oligo VF2 (10uM) : 0,5ul
- Oligo VR (10uM) : 0,5ul
- H2O : 6,5ul
- Bacteria hoted at 99°C by PCR program : 5ul
- Put PCR tube withe the PCR mix in the PCR and run the second part of the program (just push the "Enter" button)
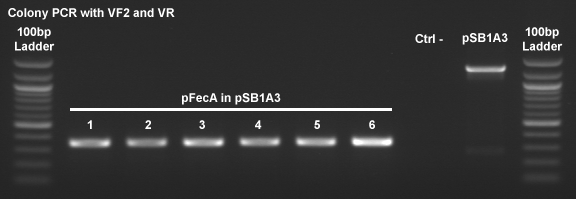
PCR screening
- For 6 clones
- Negative control without bacteria
- "Positive" control with pSB1A3 wich have the mRFP (1050bp)
Gel migration of the PCR screening
- All the clone have se same profil (more than 300bp and under 400bp) and it seem to be good...Confirmation tomorow by enzymes digestion.
- Negative control is...negative
- Positive control is...positif but have a second band. (Must be confirmed)
To do list
edit please ^^
 "
"