Team:Paris/17 August 2009
From 2009.igem.org
NoteBook
|
|
|
|
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
Lab work
Competent Bacteria
- Competent bacteria
- 1/50 ON culture of New DH5α
- OD600=0,6
- Transformation with puc19 (10pg/µl)
- 250µl of competent DH5α
- 1µl puc19
- Leave 20 min in ice
- 45s at 42°C
- Leave 2 min in ice
- Add 1ml of hot LB (42°C)
- 1h under agitation
- Put on plate (LB agar + Amp)
Vector Dephosphorylation
pSB1A3 digested by EcoRI/PstI, XbaI/PstI and EcoRI/SpeI have to be dephosphorylated before the ligation process.
Mix :
Vector : 30uL
Antarctic Phosphatase : 2uL
Buffer : 3,5 uL
Protocol
Put the solution at 37°C for 30 min
Add 2 uL of enzyme
37°C for 30 min
65°C for 20 min for heat inactivation
Ligation
D13 (X/P): 0,94 ug/mL
D14 (X/P): 1,2 ug/ml
pSB1A3 (X/P): 2,1 ug/ml
Ligation (2hrs at RT) of pSB1A3 w/ D13 (ompA signal) or D14 (TolRII)
Mix : total volume = 10 uL
vector (pSB1A3) : 2 uL
insert (A11 or A13) : 4uL
10X T4DNA Ligase buffer : 1 uL
T4 DNA Ligase : 0,5 uL
H2O: 2,5 uL
Negatif control : same mix as the previous one but without the insert. In these conditions the amount of water was increased until 6,5 uL.
Transformation
Transformation of (using the "heat choc" protocol) :
- BBa_B0034 (AmpR)
- BBa_E0030(KanR)
- pSB3C5 (CmR)
- pINTE3 (Col E3, AmpR)
- pTA (pTet-Col A, AmpR)
- pINTg3p (g3p Term, AmpR)
- pINK2(TolRII, AmpR)
- Ligation 1 ( set up on Friday 14th of august w/ D11 (RBS-Tet digested by Xba/Pst) and vecteur D12 (pSB2K3 w/ pLac digested by Spe/Pst) (KanR)
- previous ligation (w/ D13 and D14) (AmpR)
Transformation Check/PCR on colonies
Transformation check:
The x10mix ligation after centrifugation and put at 16°C overnight had work.
using less incubation time at RT seems to work better.
using less insert seems to work better
PCR on colonies:
using for 1 tube:
Master Mix 2X : 12,5µL
Oligo VF2 (10µM): 0,5µL
Oligo VR (10µM): 0,5µL
bacteria in H20 (see PCR on colonies protocol): 5µL
H20 : 6,5µL
PCR on colonies Check OK for colonie 3
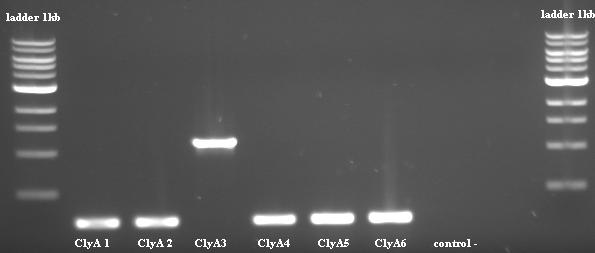
 "
"