Colony PCR of LuxPQ-B0015-R0040-LuxOU-B0015 construct in psB1AC3 using BBK CP F/R and LuxPQ F/LuxOU R primers
Colony PCR of LuxPQ-B0015-R0040-LuxOU-B0015 construct in psB1AC3 using BBK CP F/R and LuxPQ F/LuxOU R primers
Purpose: To verify the presence of LuxPQ-B0015-R0040-LuxOU-B0015 construct in psB1AC3.
Protocol
| |
MM1 (6x) (μL) |
MM2 (6x) (μL) |
| 10X PCR buffer minus MgCl2 |
30 |
30 |
| 10mM dNTPs |
6 |
6 |
| 50mM MgCl2 |
9 |
9 |
| F primer |
6 (BBK CP F) |
6 (LuxPQ F) |
| R primer |
6 (BBK CP R) |
6 (LuxOU R) |
| ddH2O |
241.8 |
241.8 |
| pTaq |
1.2 |
1.2 |
|
|
|
Positive control = LuxPQ-B0015-R0040-LuxOU-B0015 in AK3
PCR conditions
| # of cycles |
Temp (ºC) |
Time |
| 1 |
94 |
6 min |
| 36 |
94 |
30sec |
| 55 |
45sec |
| 72 |
6min 20sec |
| 1 |
72 |
10min |
| |
4 |
hold |
|
|
|
Result: the only bands that appeared on the gel were those of the ladders. Therefore, start new construction.
Colony PCR of ∆LuxPQ in psB1AC3 using BBK CP F/R and LuxPQ F/R primers
Purpose: To verify the presence of ∆LuxPQ in psB1AC3 (to verify a successful plasmid switch)
Protocol
| |
MM1 (8x) (μL) |
MM2 (6x) (μL) |
| 10X PCR buffer minus MgCl2 |
40 |
40 |
| 10mM dNTPs |
8 |
8 |
| 50mM MgCl2 |
12 |
12 |
| F primer |
8 (LuxPQ F) |
8 (BBK CP F) |
| R primer |
8 (LuxPQ R) |
8 (BBK CP R) |
| ddH2O |
322.4 |
322.4 |
| pTaq |
1.6 |
1.6 |
|
|
|
Positive control = ∆LuxPQ in psB1AK3
PCR conditions
| # of cycles |
Temp (ºC) |
Time |
|
| 1 |
94 |
6 min |
|
| 36 |
94 |
30sec |
|
| 55 |
45sec |
|
| 72 |
6min 20sec |
|
| 1 |
72 |
10min |
|
| |
4 |
hold |
|
|
|
|
|
|
|
|
|
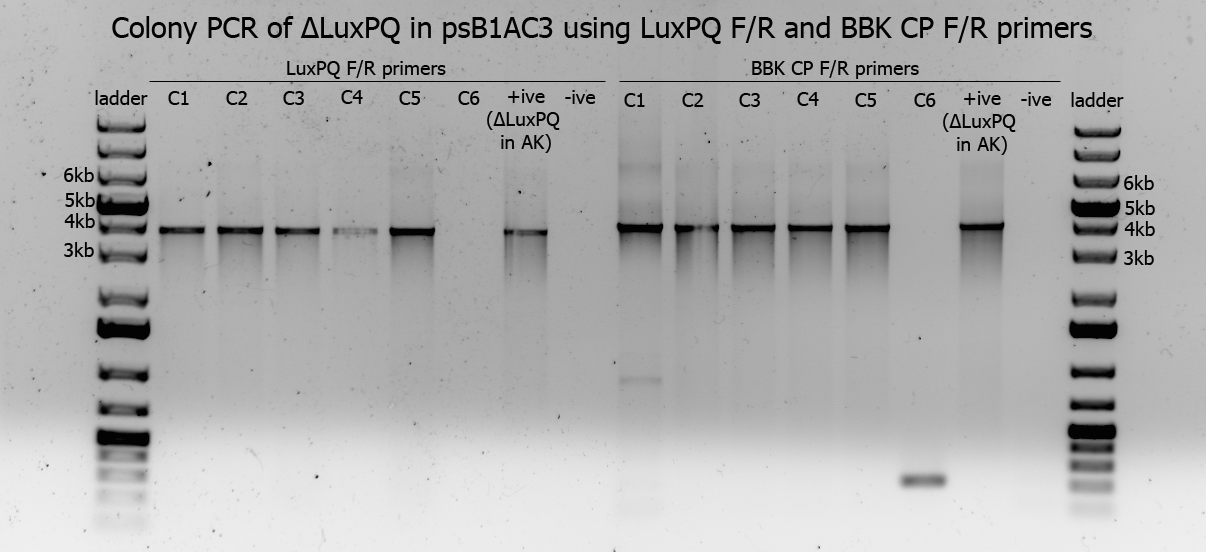
Isolating plasmid from LuxPQ-B0015-R0040-LuxOU-B0015 (AC) and LuxPQ (in AC)
Purpose: isolate and measure concentrations of pure plasmids.
| DNA |
260/280 |
260/230 |
Conc. [ng/μL] |
| ∆LuxPQ
in AC3 C1 |
1.83 |
2.23 |
146.8 |
| ∆LuxPQ
in AC3 C2 |
1.81 |
2.19 |
112.6 |
| ∆LuxPQ
in AC3 C3 |
1.8 |
2.27 |
145.8 |
| ∆LuxPQ
in AC3 C4 |
1.78 |
2.09 |
96.6 |
| ∆LuxPQ
in AC3 C5 |
1.81 |
2.25 |
115.3 |
| ∆LuxPQ
in AC3 C6 |
1.79 |
2.17 |
44.8 |
| PQ-B-R-OU-B
in AC3 C1 |
1.76 |
1.86 |
29.4 |
| PQ-B-R-OU-B
in AC3 C2 |
1.59 |
1.44 |
20.1 |
| PQ-B-R-OU-B
in AC3 C3 |
1.7 |
2.27 |
16.3 |
| PQ-B-R-OU-B
in AC3 C4 |
1.71 |
1.79 |
17 |
|
|
|
|
Construction of LuxPQ-B0015-R0040-LuxOU-B0015 by inserting B0015-R0040-LuxOU-B0015 (AK) into LuxPQ (in AC)
Purpose: To construct LuxPQ-B0015-R0040-LuxOU-B0015 by inserting B0015-R0040-LuxOU-B0015 (AK) into LuxPQ (in AC)
Protocol
Recipient
| Recipient
1 |
Recipient 2 |
|
|
| 2μL
REact 4 |
2μL REact 4 |
|
| 0.75
μL SpeI |
0.75 μL SpeI |
|
| 0.75
μL PstI |
0.75 μL PstI |
|
| 2
μL of ∆LuxPQ in AK3 C2 [112.6ng/μL] |
2 μL of ∆LuxPQ in AK3
C5 [115.3ng/μL] |
|
| 14.5
μL ddH2O |
14.5 μL ddH2O |
|
|
|
|
|
|
|
|
|
|
Insert
| 2μL
REact 2 |
|
|
|
| 0.75
μL XbaI |
|
|
| 0.75
μL PstI |
|
|
| 2
μL of LuxPQ-B0015-R0040-LuxOU-B0015 in AK3 C1 [156.3ng/μL] |
|
|
| 12.5
μL ddH2O |
|
|
|
|
|
|
|
|
|
|
|
|
|
Put in the incubator at 37ºC overnight.

