Team:Paris/Addressing testing
From 2009.igem.org
(→Adressing system) |
(→Adressing system) |
||
| Line 38: | Line 38: | ||
|- style="background: #bebebe; text-align: center;" | |- style="background: #bebebe; text-align: center;" | ||
| - | |PCR | + | |PCR : |
| - | |PCR | + | |PCR : |
| - | |PCR | + | |PCR : |
|- style="background: #d8d8d8; text-align: center;" | |- style="background: #d8d8d8; text-align: center;" | ||
| - | | | + | |Verification on gel : |
| - | | | + | |Verification on gel : |
| - | | | + | |Verification on gel : |
|- style="background: #bebebe; text-align: center;" | |- style="background: #bebebe; text-align: center;" | ||
| - | | | + | |Purification on gel : |
| - | | | + | |Purification on gel : |
| - | | | + | |Purification on gel : |
|- style="background: #d8d8d8; text-align: center;" | |- style="background: #d8d8d8; text-align: center;" | ||
| - | | | + | |digestion: |
| - | | | + | |digestion: |
| - | | | + | |digestion: |
|- style="background: #bebebe; text-align: center;" | |- style="background: #bebebe; text-align: center;" | ||
| - | | | + | |verification digestion: |
| - | | | + | |verification digestion: |
| - | | | + | |verification digestion: |
|- style="background: #d8d8d8; text-align: center;" | |- style="background: #d8d8d8; text-align: center;" | ||
| - | | | + | |ligation: |
| - | | | + | |ligation: |
| - | | | + | |ligation: |
|- style="background: #bebebe; text-align: center;" | |- style="background: #bebebe; text-align: center;" | ||
| - | | | + | |PCR colo : |
| - | | | + | |PCR colo : |
| - | | | + | |PCR colo : |
| + | |||
| + | |- style="background: #d8d8d8; text-align: center;" | ||
| + | |miniprep: | ||
| + | |miniprep: | ||
| + | |miniprep: | ||
| + | |||
| + | |||
| + | |- style="background: #bebebe; text-align: center;" | ||
| + | |sequencing : | ||
| + | |sequencing : | ||
| + | |sequencing : | ||
| + | |||
|} | |} | ||
| Line 105: | Line 117: | ||
PBAD E/S | PBAD E/S | ||
| - | + | verification digestion: | |
Revision as of 11:34, 17 October 2009
iGEM > Paris > WetLab > Addressing
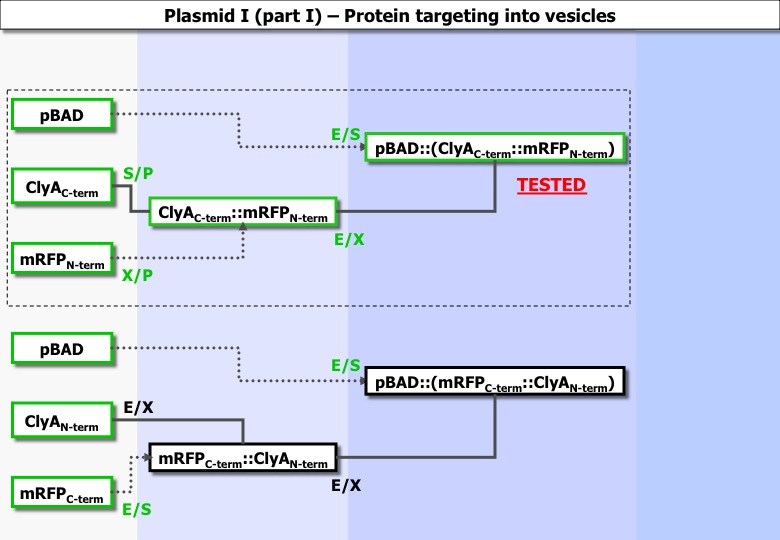
Adressing system
global constructions :
Time required : A lOT !!!!!
Experiments ran :
| row 1 | row 2 | row 3 |
| PCR : | PCR : | PCR :
|
| Verification on gel : | Verification on gel : | Verification on gel :
|
| Purification on gel : | Purification on gel : | Purification on gel : |
| digestion: | digestion: | digestion: |
| verification digestion: | verification digestion: | verification digestion:
|
| ligation: | ligation: | ligation:
|
| PCR colo : | PCR colo : | PCR colo : |
| miniprep: | miniprep: | miniprep:
|
| sequencing : | sequencing : | sequencing : |
From plate: mRFP
PCR :
ClyA Cterm plasmid : ? Oligo : TM :
ClyA Nterm plasmid : ? Oligo : TM :
mRFP Cterm plasmid : ? Oligo : TM :
mRFP Nterm plasmid : ? Oligo : TM :
Verification on gel
ok
Purification on gel
ok
digestion:
ClyA Cterm S/P RFP Nterm X/P PBAD E/S
verification digestion:
ligation:
ClyA Cter-RFP Nter vector PSB1A3
PCR colo :
miniprep:
verifivation of the miniprep by digestion
sequencing [[ClyA]]
-ClyA Nterm= biobrick+sequencage+stock glycerol Done
-ClyA Cterm= biobrick+sequencage+stock glycerol Done
[[-RFP-]]
-RFP Nterm= biobrick+sequencage+stock glycerol Done
-RFP Cterm= biobrick+sequencage+stock glycerol Done
[[-ClyA Cter-RFP Nter-]]
-ClyA Cter-RFP Nter= biobrick+stock glycerol Done
[[-RFP Cter-ClyA Nter-]]
-miss ligation between 2 parts (j'ai fait plusieurs ligation le 7ou8/09, mais pas marché le 9/09 apres culture overnight, voir si les autres ont continué)
[[PBad-ClyA Cter-RFP Nter-]]
-failed many time with:
Pbad cut in S/P and ClyA Cter-RFP Nter in X/P
Pbad cut in E/S and ClyA Cter-RFP Nter in E/X
 "
"