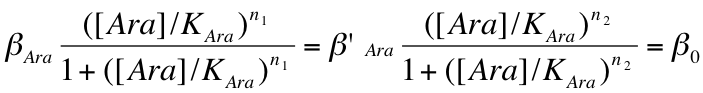Team:Paris/Production modeling
From 2009.igem.org
(→B. Modeling Vesicles creation.) |
(→A. Genetic Network Regulation) |
||
| Line 23: | Line 23: | ||
We had two hypothesis which led us to think over two different genetic regulatory network : | We had two hypothesis which led us to think over two different genetic regulatory network : | ||
| + | <font color=red> Je regrette mais les deux points ci-dessous ne sont clairs du tout... dans les deux cas vous supposez que le nombre de vésicules produites par unité de temps est maximal quand TolRII est maximal. | ||
| + | Vous ne discutez pas ci-dessous la raison du délai. Partie à retravailler</font> | ||
*first the number of vesicules produced is at its maximum when the concentration of TolRII has reached its peak ; this achieve this configuration, we tried to design a delay system based on a transcriptional cascade as described in the part “The delay sytem”. | *first the number of vesicules produced is at its maximum when the concentration of TolRII has reached its peak ; this achieve this configuration, we tried to design a delay system based on a transcriptional cascade as described in the part “The delay sytem”. | ||
| Line 33: | Line 35: | ||
===="The Delay system"==== | ===="The Delay system"==== | ||
| - | The first thing that we wanted to study in modeling was the efficiency of the construction chosen to create a delay. Our first approach was a deterministic analysis of the system using differential equations. The regulation of promoters was described using the Hill functions and the methods described by U. Alon in his book ''An Introduction to Systems Biology''. | + | The first thing that we wanted to study in modeling was the efficiency of the construction chosen to create a delay. Our first approach was a deterministic analysis of the system using differential equations. The regulation of promoters was described using the Hill functions and the methods described by U. Alon in his book ''An Introduction to Systems Biology'' <font color=red>Références complètes </font>. |
Revision as of 21:04, 14 October 2009
iGEM > Paris > Production > DryLab > Delay Model
Delay model est un mauvais titre du chapître.
A. Genetic Network Regulation
Changer ce titre aussi: on modélise ici la la production des protéines message et de la bouteille vésicule
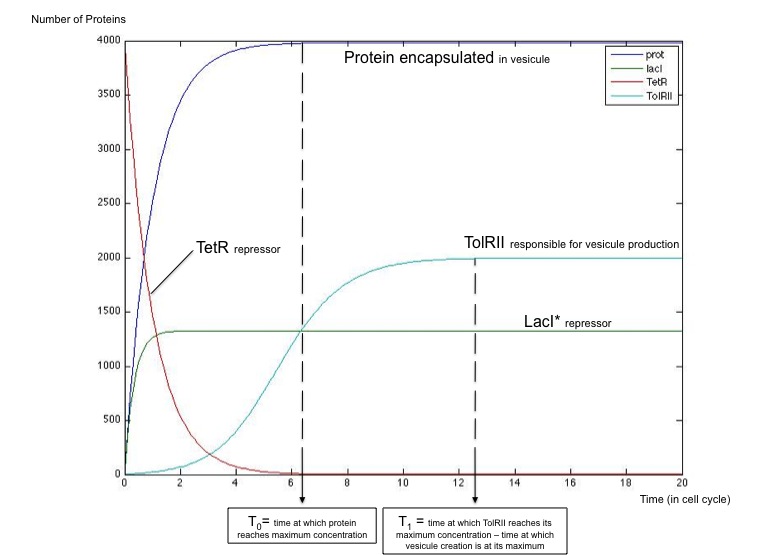
It is crucial to send vesicules only when the concentration of proteins to be sent is at its maximum level in the periplasm. To increase the power of our communication system, we also tried to make this peak of vesicules creation correspond with the peak of protein concentration.
We had two hypothesis which led us to think over two different genetic regulatory network :
Je regrette mais les deux points ci-dessous ne sont clairs du tout... dans les deux cas vous supposez que le nombre de vésicules produites par unité de temps est maximal quand TolRII est maximal.
Vous ne discutez pas ci-dessous la raison du délai. Partie à retravailler
- first the number of vesicules produced is at its maximum when the concentration of TolRII has reached its peak ; this achieve this configuration, we tried to design a delay system based on a transcriptional cascade as described in the part “The delay sytem”.
- then, we also made the hypothesis that the number of vesicules by time units would be more important when the production of TolRII by time units is at its maximum ; this would allow a more powerful signal provided a good concentration of proteins. That is what we tried to put into place thanks to the use of feed-forwards ; see here for more explanations.
"The Delay system"
The first thing that we wanted to study in modeling was the efficiency of the construction chosen to create a delay. Our first approach was a deterministic analysis of the system using differential equations. The regulation of promoters was described using the Hill functions and the methods described by U. Alon in his book An Introduction to Systems Biology Références complètes .
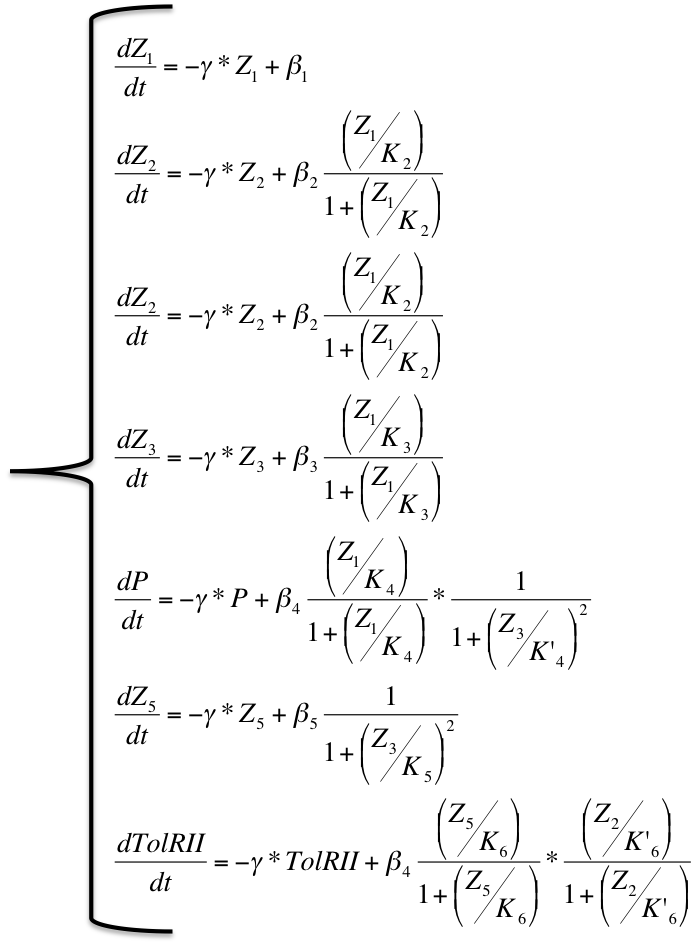
When the system is activated, there is Arabinose in the medium and the pBad promoters are activated. And the system can be described by this system of differential equations :
| Differential System |
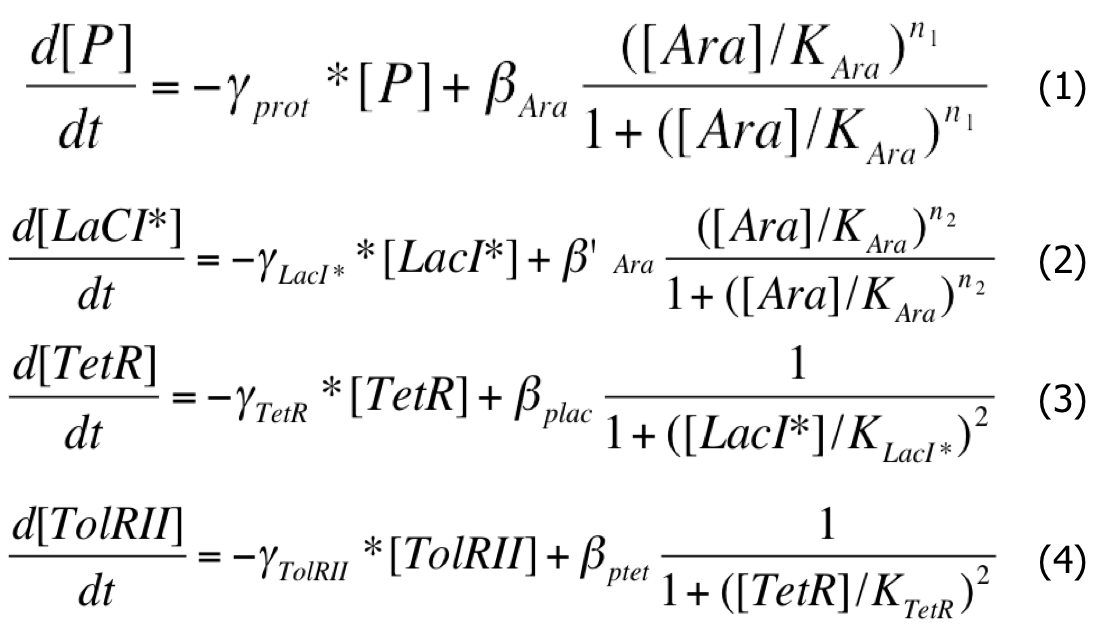
|
As a first approximation, we assumed that :
- When the pBad promoter is induced, the concentration of arabinose in the medium is very high and constant during he whole study ; as a consequence, we will consider that the creation rate of Protein and LacI* is constant during the experiment :
- We considered that all the binding constants are identical and of an average of 40nM which correspond to approximately 40 monomers per cell ; we can write that :
- We chose identical intrisinc promoter activities, β , all equal to 4000 proteins/cell cycle :
- All the time units were expressed in units of cell cycle (approximately half an hour) ; as a consequence we chose a dilution rate γ of 1 for protein without special tags. For the Laci protein with a LVA tag, the dilution rate is multiplied by 3:
The Feedforwards
Using the delay system in our genetic regulatory network, we were able to find a way to get a maximum quantity of TolRII when the concentration of exported proteins is at its maximum.
We also have thought of another device designed to have a reaction rate of TolRII production at its higher level when the concentration of exported is at its maximum. To this end, we decided to combine two feed-forward loops to create a "burst" in the production of each protein.
The Feed Forward Motif
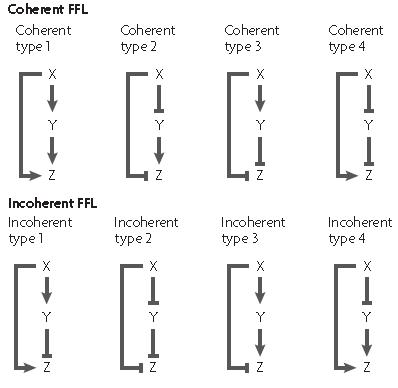
The feed forward motif was studied and described by U. Alon in his book "An Introduction to Synthetic Biology" where he describes the different form of existing feed forward ; they can be either coherent or incoherent.
In each case, there exist a different network organisation to create a feed-forward loop as shown on the picture
The X and Y inputs can be combined either using a logical AND or OR gate thus giving different dynamics properties in each case.
The coherent feed forward loop allow the creation of a delay either on activation or on desactivation depending on the logical function between the two input X and Y. When the output logical gate is an AND function, the delay appears on activation, while when the output gate is an OR function, the delay appears on desactivation. An example of caracterised feed-forward loop is the fliA system which was used by the 2008 Paris team in order to create a oscillator.
In an incoherent feed forward loop, the two inputs of the last gate function have two different logical levels; one molecule (molecule X for example) is an activator while the other molecule is a repressor of the promoter controlling the expression of the output.
This way, when the input signal is on, the expression of molecule X starts activating both the promoters controlling the expression of Y and Z ; transcription starts and the amount of Y and Z molecules increases. Nevertheless, when the concentration of Y has reached its repression threshold, repression starts and the concentration of Z molecule decrease to its steady state.
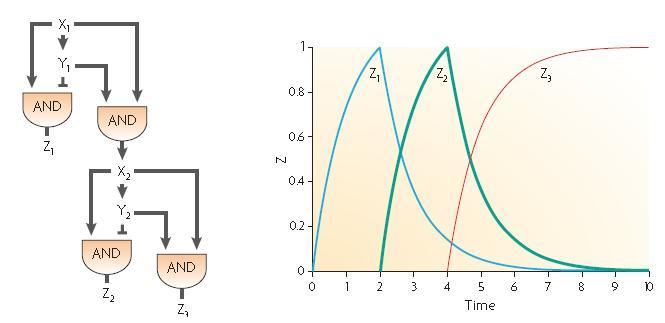
Two elementary incoherent feed forward motifs can be combined in order to create two time delayed bursts as described by U.Alon in its description of the development of Bacillus Subtilis spore. Anyway, it seems quite difficult to find a molecule which would be both an activator and a repressor ; we decided to try and think about another combination of these two feed forwards so as to get the dynamics we wanted as shown on the following part.
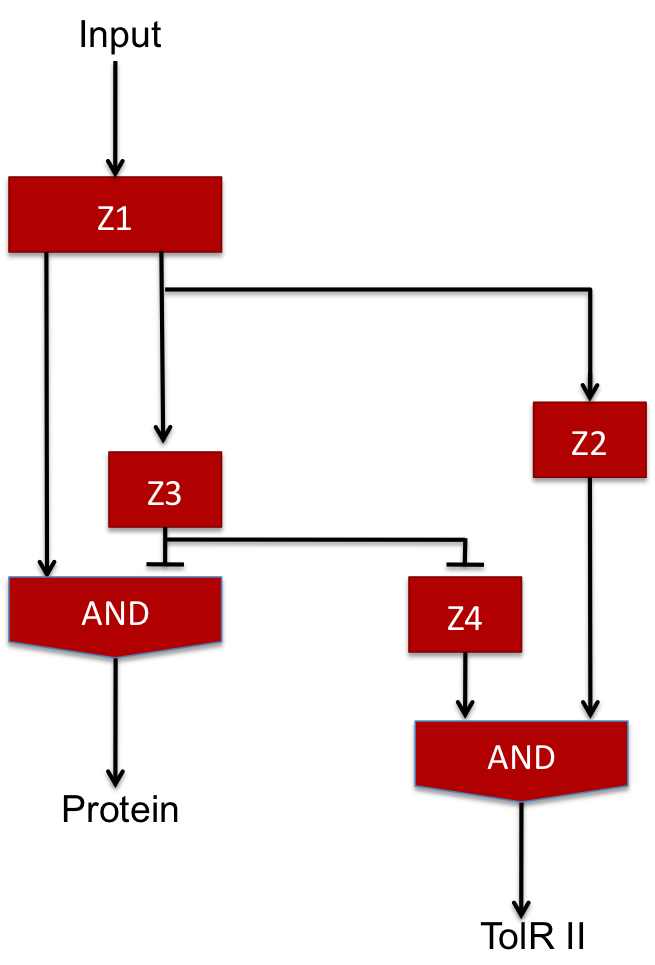
Our design proposition
The two feed-forward loops used are incoherent and agenced in a way that each molecule is either an activator or a repressor (and not both). The following scheme gives the functionning of the system.
| Steps | Description | Changes in concentration |
| 1 | When the input signal is ON, molecule1 is created and activates the promoter upstream the coding sequence for molecules Z2 and Z3 as well as Z4 since there is no molecule Z3 in the medium (see the AND gate function) at the beginning ; there is an initial amount of Z5 molecule but no Z2 so the no Z6 molecule is produced. | Z1 +++
Z2+ Z3+ Z4++ Z5~ Z6~ |
| 2 | After a while, there is enough Z3 to start the repression of Z4 promoter and there is also enough Z2 to start producing Z6 signficantly. | Z1+++
Z2+++ Z3+++ Z4- Z5~ Z6+ |
| 3 | Then, because of the repression of Z3, the Z4 molecule reaches its steady level ; in the same time, Z3 starts repressing the production of Z5 and the amount of Z6 starts decreasing. | Z1+
Z2+ Z3+ Z4~ Z5- Z6+ |
| 4 | Finally, Z3 repressesthe promoter allowing the creation of Z5 and every molecules reaches its steady level concentration. | Z1~
Z2~ Z3~ Z4~ Z5- Z6~ |
To study the dynamics of our model, we decided to use a classical approach using Michaelis-Menten equations to medel the activation and repression of the various promoters, including a cooperativity of order 2 for the repressor which often the case for 'classical' repressors such as LacI. We obtained the following system of equations :
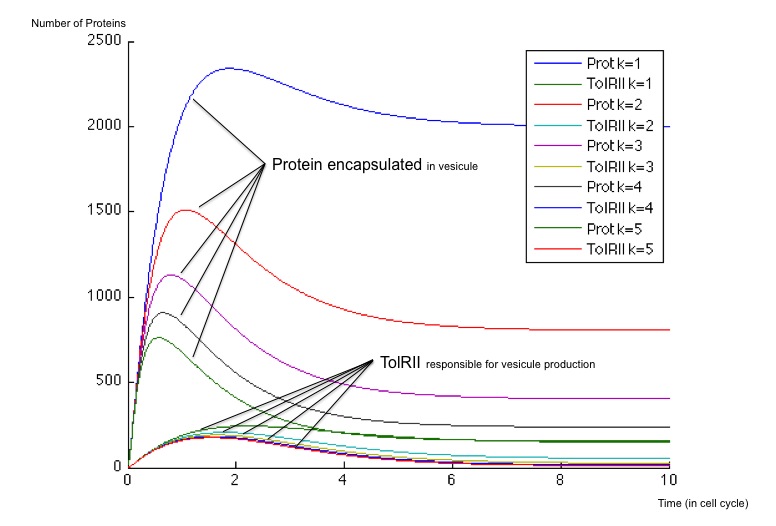
It is possible to obtain a important scale of values according to the choice of an optimized β for the production of molecule Z3 whose steady concentration and production rates impact on the steady state concentrations of the protein exported as well as the TolRII expressed. The β is chosen initially equal to 40 and 400 molecules by cell cycle and we have plotted the different response for multiples of this initial value.
A flexibility of Behaviour
Regardless of the steady state concentrations of the molecules, the use of two incohenrent feed forward loops allow a more important flexibility in the design of our system. With a simple delay system, we just get one possible dynamic behavior while using a variable range of production rates, we can obtain two behavior :
- First, when the β is weak (β = 40), the number of TolRII produced slowly increase before returning to zero without another input signal of any kind ; on the other hand, the protein exported inside the vesicles is created very quickly and remains at a high concentration compared to the TolRII protein ; we can therefore send vesicles containing an important amount of interesting proteins and, after a while, stop expressing dangerous proteins for the cell. All of this can be performed with and single input signal.
- When the β is stronger, the behavioris different and we can notice a change of concavity of the TolRII curve when the amount of protein exported is at its maximum. During such a change of concavity, the derivative reaches its maximum - in other words, the amount of TolRII produced by time units is at its higher level. Considering the hypothesis according to which the number of vesicules by time units would be more important when the production of TolRII by time units is at its maximum, we get a peak in vesicule production in this case.
- In any case, the number of TolRII proteins is less important than the amount of exported protein, thus protecting the cells from lysis.
B. Modeling Vesicles creation.
Our model is based on three different physical phenomenon:
- The lipid surface conformation, Tol-Pal proteins diffusion and the increase of the osmotic pressure in the periplasm
- The Lipid conformation of the outer membrane is a well known problem: at 35°c the lipid bilayer behaves like a liquid which conformation character is ruled by an energy called the bending energy. This energy represents the fact that the lipid bilayer will search a special conformation depending on the shape and the chemical properties of its constituents.
- The conformation of the lippopolysaccharides and the normal phospholipids can be seen has a lipid bilayer which conformed itself as a liquid crystal at the growth culture. The conformation of lipid bilayer has been well studied and a lot of theoretical results have been shown. Some of them are of great interest for our project. This conformation is something which can enable us to understand the way vesicle can be produced and how they will be received. In our project we first notice that from a basic calculus we can obtain very interesting results on the way outer membrane vesicles can be produced.
- We first use the bending energy has a rough shape for our model and its understanding:
- This formula give use the abilities to explain the way lipids will organize together. E is the energy of a whole lipid bilayer (or monolayer).Kb and Kg are Bending and Gaussian modulii which can be obtained by experiments. γ0 is the intrinsic curvature of the outer membrane which describes the local form of a lipid bilayer when it is at is lowest state of energy ,the more stable. γm and Hg are the mean curvature and the gaussian curvature. Our first task was to calculate the energy of two different shapes of membranes.
- We compute the E.coli Shape before the division and then the vesicles' shape.
- We consider E.coli's shape as a cylinder of rayon r =0.3 μm and of infinite length. The aim of this first representation was to estimate this energy in the division region of E.coli before division. With this approximation γm is equal to 2/r and γg=0. In this case:
- For the vesicles we consider their basic shape as a sphere of rayon r’ so the bending energy by lipid area units is:
- Thus as the area of a sphere is known and is independent of the location on the sphere we can write: In the same way we can write for an area of E.coli lipids that the bending energy is:
- The considerations aforementioned can provide a basic vision of the statistical repartition of vesicles in case of absence of integrity control system in the outer membrane.
- The first energy is the potential energy of the lipid area in E.coli outer membrane necessary to the construction of a vesicle and the second one the potential energy of the same lipid area but in the conformation of a vesicle shape. So the energy which must be given to the whole system to create a vesicle is:
- We can suppose that the most easily created vesicles will be the ones which requires a minimum energy . By derivation we find that the minimum his obtain for:
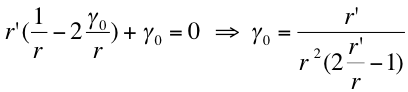
|
- Hence as we know that the range of created vesicles radii is 25 nm to 175nm we can suppose that the r’ is somehow about 100 nm and so:
- We can consider the orders of magnitude as really realistic. In fact we know that the E.coli lipid bilayer is built of distinct types of lipids: Lippopolysacharides and simple phospholipids. LPS are located in the exterior lipid layer of the outer membrane. The others are located in the interior lipid bilayer. Moreover, those LPS present a sugar extension toward the medium. Those sugars can bind to each other. So we can assume that they are going to create clusters and to curve the membrane toward the exterior of E.coli.
- Tol and Pal are membrane proteins which are located respectively in the outer and the inner membrane. The diffusion of proteins in those lipid bilayers can be modelled by a probabilistic Brownian movement. This diffusion model gives us the law of probability for the location of Tol and Pal on the membranes. It has been observed that the Tol and Pal proteins interact with each other, which is linked to the membrane stability: indeed the Tol and Pal will bind inner and outer membrane and furthermore stabilize the outer membrane using the peptidoglycan rigidity.
- The osmotic pressure in the periplasm is the same that the medium pressure in normal time. But during the division period of the bacteria the peptidoglycan is degraded to be recycled in a new cell wall. During this phenomena of turn-over a part of the peptidoglycan is released in the periplasm which increased its osmotic pressure.
- Thus, we can consider that if there is not enough Tol-Pal linked proteins the outer membrane will distort to create a beginning of vesicle. But in this part of the membrane the tol pal proteins will not have the possibility to bind themselves and they will be free to diffuse in other parts of the membranes. The surface shape will guide the proteins to the border of the vesicle and stabilized the shape of the vesicle.
- To describe the time variation of the vesicles creation we first have to use a more complicated approach: we need to use an equation which is obtained by derivation of the bending energy:
Ou-Yang and Helfrich [Phys. Rev. Lett. 59 (1987) 2486]
- Our first model which didn't use the brownian movement :
- We first conciders that the membrane equation can be first simplified to a one dimentional model :in a first approach we use a one dimensional model based on a polar curvature simplification approximation :
- the curvature in polar is:
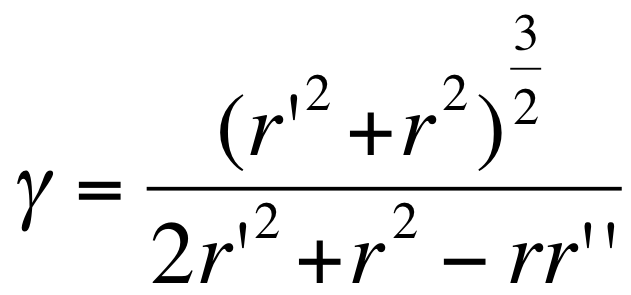
|
- With the hypothesis of a cylinder form the mean and gaussian curvatures can be written as:
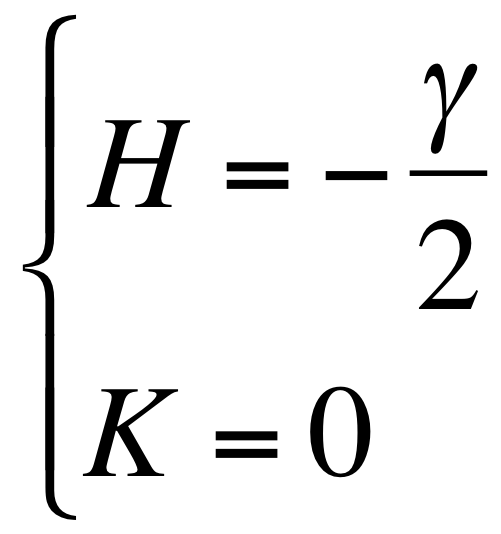
|
- indeed we have in those hypothesis:
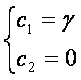
|
- Thous we can concider the polar system of equation:
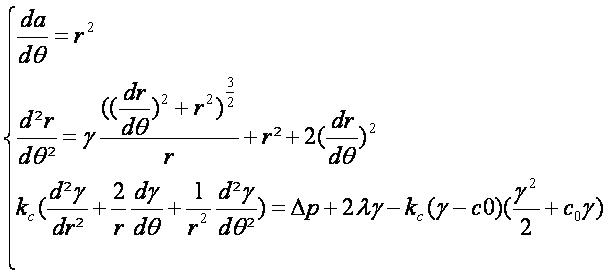
|
- to have a unique solution we must use some boundary conditions to compute our model (Cauchy Lipshitz thorem for the mathematician ;)) those conditions are simple without the peptidoglycan attachement lypoproteins (Pal). As the model is dependent of the angle we must impose the fact that r(0)=r(2π). In addition the Δp must be concidered as dependent of the quantity of periplasmic species.
- Indeed the osmotic pressure can be written in this way:
- And by concidering that we have weak concentrations we can concider a simpliest formula analogue to the perfect gases law:
- Finaly we can write:
- And Thous the whole parameters are given. In addition we can translate the role of the Tol/Pal System has boundary condition on the whole system: a cluster of tol-Pal can be translate as a point of radius equal to the peptidoglycan ones. Finally we can have this type of results:
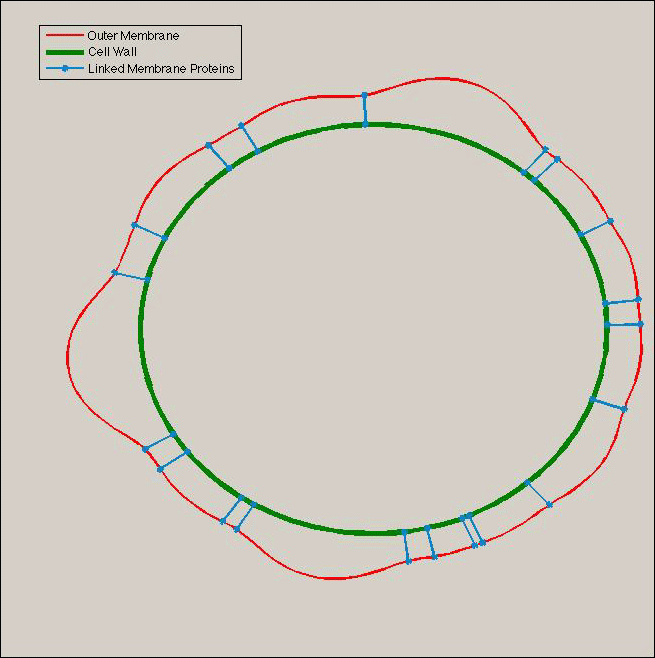
|
- Brownian motion:
- The Brownian motion links the concentration of proteins with the shape of the membrane they are includes in:
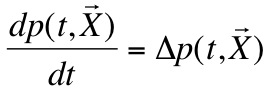
|
- P is the density probability of presence from proteins.
- The Laplacian of P explains the fact that the concentration of proteins like Pal increased in regions of negative curvature. Such a model can explain some last observations [Kumaran & Losick] “negative membrane curvatures as a cue for sub-cellular localization of a bacterial protein”. So we can assume that the Pal will be confined to region with negative curvature and so the probability of linking between Pal and Tol increased in those regions enhancing the stability and so enlarging those regions.
- So Brownian motion is a good explanation for the proteins migration toward the septa during division.
- But this Brownian motion can explain the vesiculation too. When there are too few proteins we can assume the creation of a vesicle at germinal states (called bleb). At the border of the blebs an amount of Pal protein prevent the whole membrane to swallow and keep the vesicle in form. But at the border between the blebs and the protein rich area curvature is highly negative and so the quantity of proteins in those regions increases enlarging the region toward the centre of the blebs.
- Then two different options are expectable: the blebs is too small and then the proteins are “zippering” back the blebs. But if the bleb is big enough the zippering became a separation between the vesicle and the bacteria. The good vision of this blebs forming can be understood in this way: the proteins rich region on the border pass from an annular form toward a circular one.
 "
"