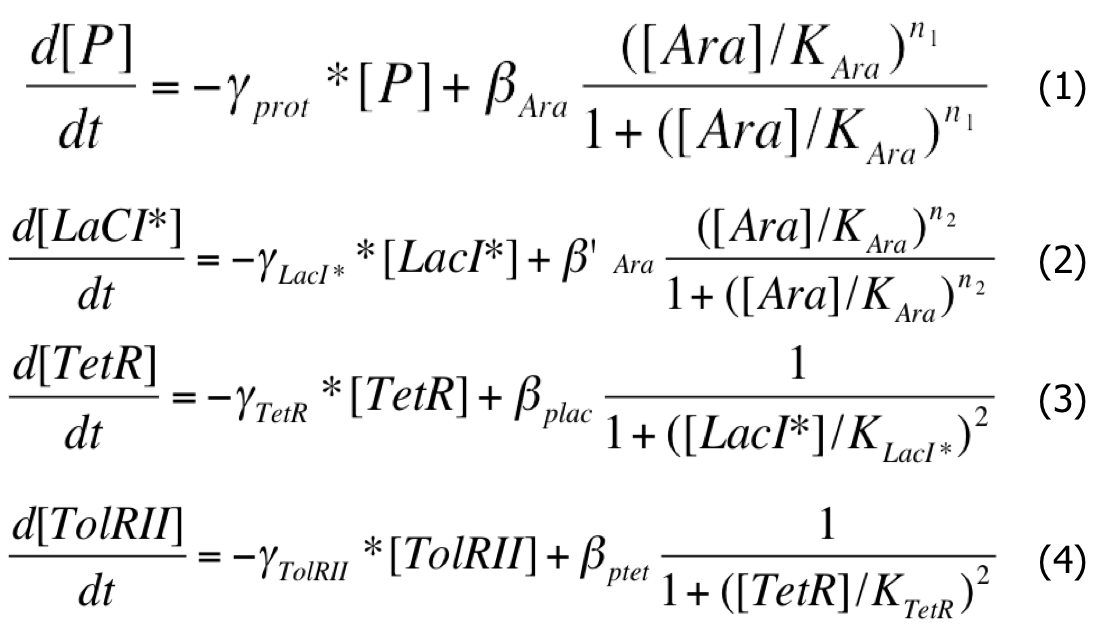Team:Paris/Production modeling
From 2009.igem.org
(Difference between revisions)
(→A. Genetic Network Regulation) |
(→A. Genetic Network Regulation) |
||
| Line 20: | Line 20: | ||
When the system is activated, there is Arabinose in the medium and the pBad promoters are activated. And the system can be described by this system of differential equations : | When the system is activated, there is Arabinose in the medium and the pBad promoters are activated. And the system can be described by this system of differential equations : | ||
| + | {| | ||
| + | |- style="background: #0d3e99; text-align: center; color:black;" | ||
| + | |width=100%| Differential System | ||
| + | |- style="background: #0d3e99; text-align: center" | ||
| + | |[[Image:Genetic network equations.png|350px|right| Modelling Overview]] | ||
| + | |} | ||
| - | |||
| - | + | *As the Plac promoter is leaking, we tried to model this behaviour by introducing a constant rate K in the activation coefficient of PLac. | |
| - | [[Image:Genetic | + | [[Image:Modelling Genetic Network.png|350px|left| Modelling Overview]] |
Revision as of 09:45, 2 September 2009
iGEM > Paris > Production > Modeling
Modeling
A. Genetic Network Regulation
The first thing that we wanted to study in modeling was the efficiency of the construction chosen to create a delay. Our first approach was a deterministic analysis of the system using differential equations. The regulation of promoters was described using the Hill functions and the methods described by U. Alon in his book An Introduction to Systems Biology.
When the system is activated, there is Arabinose in the medium and the pBad promoters are activated. And the system can be described by this system of differential equations :
| Differential System |
- As the Plac promoter is leaking, we tried to model this behaviour by introducing a constant rate K in the activation coefficient of PLac.
 "
"