Team:Newcastle/Labwork/28 September 2009
From 2009.igem.org
Formal Lab Session - 28th September 2009
Previously Metal Sensing Team tried to ligate cotC biobrick with pmutin4 and transform E.coli DH5 cells. Although the transformations have worked, the control plates wothout the insert had also colonies.
We selected 10 colonies from the plate with the insert + the backbone and inoculated to LB+Amp for ON cultures for tomorrow's miniprep.
We prepared some agar gel.
Chassis team
Introduction
- We had cultured 4 potential right strains for kinA transformation on sunday, today after we double check the mini prep result, we can carry out Midi prep with the right strain.
Experiment procedure
Digestion of mini prep result from last friday
- The digestion reaction for No.4,5,6,10
dd H2O 7ul
10X fast digest buffer 2ul
Fast HindIII 1ul
Mini Prep DNA 10ul
------------------------------
20ul
- 37 degree for 1 hour.
- Run the sample on 0.8% agarose gel.
- Scan the result under UV light.
Midi Prep
- It seems the No.6 has the high probability to be the right clone.
- We carried out a Midi Prep for No.6 Midi culture cells.
Conclusion
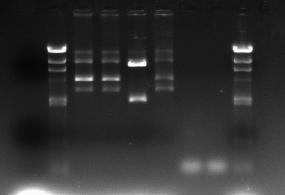
- After the ligation, pGFP-rrnB:kinA has 2 HindIII restriction sites, one is inside the kinA sequence, another was located in pGFP-rrnB backbone. When we cut the pGFP-rrnB with HindIII, two fragments would be generated. One is about 8000bp sequence, another is 2000bp sequence .
- The gel picture showed that the No.6 seems like the right one.
lane 1: ladder lane 2: No.4 lane 3: No.5 lane 4: No.6 lane 5: No.10 lane 8: ladder
Futher plan
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
News
Events
- 20 – 21 June 2009 - Europe workshop (London)
- 23 – 24 June 2009 - UK iGEM meetup (Edinburgh)
- 23 October Practice Presentation (Newcastle)
- 23 October T-shirts are ready
- 27 October Practice Presentation (Sunderland)
- 27 October Poster is ready
- 30 October – 2 November 2009 - Jamboree (Boston)
Social Net
 "
"