Team:Paris/Production testing delay
From 2009.igem.org
(Difference between revisions)
(→part2) |
(→Delay) |
||
| (14 intermediate revisions not shown) | |||
| Line 4: | Line 4: | ||
== WetLab == | == WetLab == | ||
| - | <center> [[team:Paris/WetLab#WetLab| Main]] - [[Team:Paris/Addressing_testing#top|Addressing]] - [[Team:Paris/Production_testing#top| Production]] - [[Team:Paris/ | + | <center> [[team:Paris/WetLab#WetLab| Main]] - [[Team:Paris/Addressing_testing#top|Addressing]] - [[Team:Paris/Production_testing#top| Production]] - [[Team:Paris/Transduction_testing#top | reception]] - </center> |
<center> '''Production'''</center> | <center> '''Production'''</center> | ||
<html> | <html> | ||
| Line 19: | Line 19: | ||
[[Team:Paris/Production_testing_delay#part1 | Part1 : testing delay phase 1]] | [[Team:Paris/Production_testing_delay#part1 | Part1 : testing delay phase 1]] | ||
| + | |||
| + | [[Team:Paris/Production_testing_delay#functional testing of the part 1 | Part1 : functional testing of the part 1]] | ||
| + | |||
[[Team:Paris/Production_testing_delay#part2 | Part2 : testing delay phase 2]] | [[Team:Paris/Production_testing_delay#part2 | Part2 : testing delay phase 2]] | ||
| + | [[Team:Paris/Production_testing_delay#functional testing of the part 2 | Part2 : functional testing of the part 2]] | ||
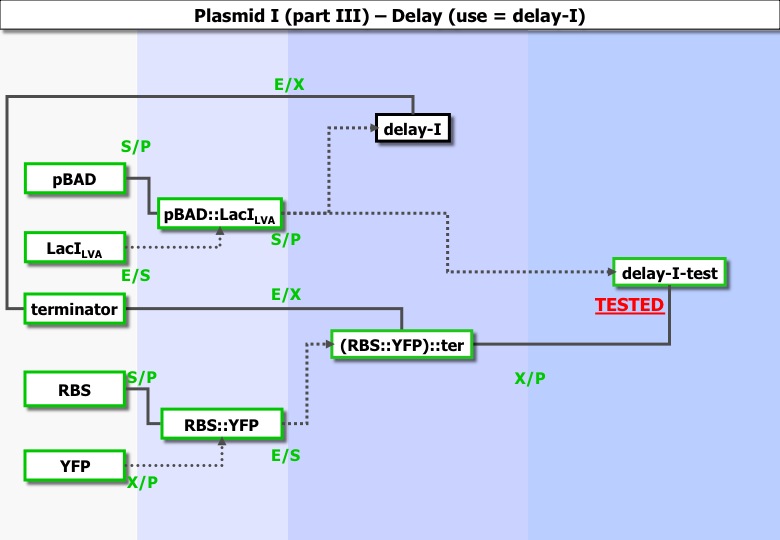
===part1 === | ===part1 === | ||
| - | [[Image:Paris_construc_3_delay1.jpg | | + | [[Image:Paris_construc_3_delay1.jpg |800px|center|plasmid = PSB3T5]] |
| Line 37: | Line 41: | ||
| + | {| | ||
| + | |- style="background: #0a3585; text-align: center; color:black; font-weight:bold; " | ||
| + | |width="300px" ; bgcolor="#F8F8F8"| column 1 | ||
| + | |width="300px" ; bgcolor="#E0E3FE"| column 2 | ||
| + | |width="300px" ; bgcolor="#CBD1FD"| column 3 | ||
| + | |width="300px" ; bgcolor="#BCCDFD"| column 4 | ||
| + | |||
| + | |||
| + | |- style="background: #bebebe; text-align: center;" | ||
| + | |'''PCR :''' | ||
| + | |||
| + | TatABCE matrix : | ||
| + | |||
| + | |'''PCR :''' | ||
| + | |'''PCR :''' | ||
| + | |'''PCR :''' | ||
| + | |||
| + | |- style="background: #d8d8d8; text-align: center;" | ||
| + | |'''Verification on gel :''' | ||
| + | |||
| + | ok | ||
| + | |||
| + | |'''Verification on gel :''' | ||
| + | |||
| + | - | ||
| + | |||
| + | |'''Verification on gel :''' | ||
| + | |||
| + | - | ||
| + | |'''Verification on gel :''' | ||
| + | |||
| + | - | ||
| + | |||
| + | |- style="background: #bebebe; text-align: center;" | ||
| + | |'''Purification on gel :''' | ||
| + | |||
| + | ok | ||
| + | |||
| + | |'''Purification on gel :''' | ||
| + | |||
| + | - | ||
| + | |||
| + | |'''Purification on gel :''' | ||
| + | |||
| + | - | ||
| + | |'''Purification on gel :''' | ||
| + | |||
| + | - | ||
| + | |||
| + | |||
| + | |||
| + | |- style="background: #d8d8d8; text-align: center;" | ||
| + | |'''digestion:''' | ||
| + | |||
| + | TatABCE | ||
| + | |||
| + | |'''digestion:''' | ||
| + | |'''digestion:''' | ||
| + | |'''digestion:''' | ||
| + | |||
| + | |- style="background: #bebebe; text-align: center;" | ||
| + | |'''verification digestion:''' | ||
| + | |||
| + | ok | ||
| + | |||
| + | |'''verification digestion:''' | ||
| + | |||
| + | - | ||
| + | |||
| + | |'''verification digestion:''' | ||
| + | |||
| + | - | ||
| + | |'''verification digestion:''' | ||
| + | |||
| + | - | ||
| + | |||
| + | |||
| + | |- style="background: #d8d8d8; text-align: center;" | ||
| + | |'''ligation:''' | ||
| + | |||
| + | TatABCE | ||
| + | |||
| + | |'''ligation:''' | ||
| + | |'''ligation:''' | ||
| + | |'''ligation:''' | ||
| + | |||
| + | |- style="background: #bebebe; text-align: center;" | ||
| + | |'''PCR colo :''' | ||
| + | |||
| + | ok | ||
| + | |||
| + | |'''PCR colo :''' | ||
| + | |||
| + | - | ||
| + | |||
| + | |'''PCR colo :''' | ||
| + | |||
| + | - | ||
| + | |'''PCR colo :''' | ||
| + | |||
| + | - | ||
| + | |||
| + | |- style="background: #d8d8d8; text-align: center;" | ||
| + | |'''miniprep:''' | ||
| + | |||
| + | STOPPED | ||
| + | |||
| + | |'''miniprep:''' | ||
| + | |||
| + | - | ||
| + | |||
| + | |'''miniprep:''' | ||
| + | |||
| + | - | ||
| + | |||
| + | |'''miniprep:''' | ||
| + | |||
| + | - | ||
| + | |||
| + | |- style="background: #bebebe; text-align: center;" | ||
| + | |'''sequencing :''' | ||
| + | |||
| + | - | ||
| + | |||
| + | |'''sequencing :''' | ||
| + | |||
| + | - | ||
| + | |||
| + | |'''sequencing :''' | ||
| + | |||
| + | - | ||
| + | |'''sequencing :''' | ||
| + | |||
| + | - | ||
| + | |||
| + | |} | ||
| + | |||
| + | ===functional testing of the part 1=== | ||
| + | |||
| + | PBAD LacI LVA- YFP was transforme into Top ten Bacteria and grew on medium : | ||
| + | |||
| + | |||
| + | 1% arabinose | ||
| + | |||
| + | 1% glucose | ||
| + | |||
| + | LB pure | ||
| + | |||
| + | |||
| + | <html> | ||
| + | </div> | ||
| + | <div id="paris_content_boxtop"> | ||
| + | </div> | ||
| + | <div id="paris_content"> | ||
| + | </html> | ||
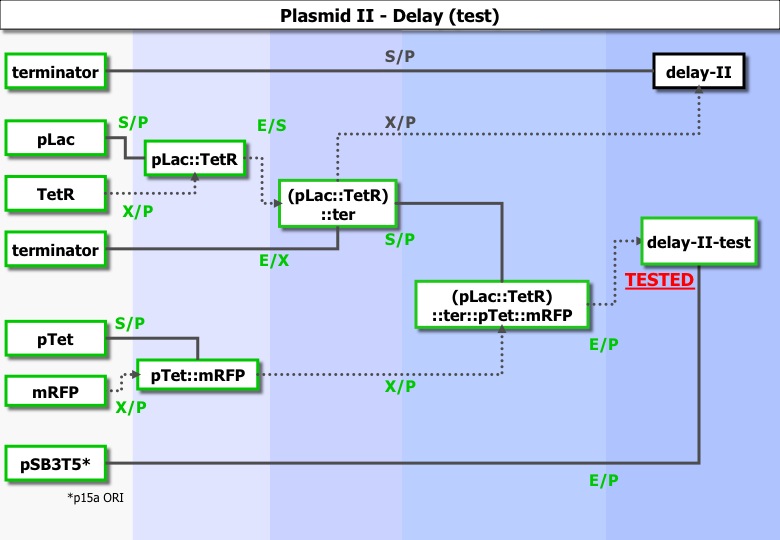
===part2 === | ===part2 === | ||
| - | [[Image:Paris_construc_5_delay2.jpg | | + | [[Image:Paris_construc_5_delay2.jpg |800px|center|plasmid = ]] |
| Line 48: | Line 207: | ||
Experiments ran : | Experiments ran : | ||
| + | {| | ||
| + | |- style="background: #0a3585; text-align: center; color:black; font-weight:bold; " | ||
| + | |width="300px" ; bgcolor="#F8F8F8"| column 1 | ||
| + | |width="300px" ; bgcolor="#E0E3FE"| column 2 | ||
| + | |width="300px" ; bgcolor="#CBD1FD"| column 3 | ||
| + | |width="300px" ; bgcolor="#BCCDFD"| column 4 | ||
| + | |||
| + | |||
| + | |- style="background: #bebebe; text-align: center;" | ||
| + | |'''PCR :''' | ||
| + | |||
| + | TatABCE matrix : | ||
| + | |||
| + | |'''PCR :''' | ||
| + | |'''PCR :''' | ||
| + | |'''PCR :''' | ||
| + | |||
| + | |- style="background: #d8d8d8; text-align: center;" | ||
| + | |'''Verification on gel :''' | ||
| + | |||
| + | ok | ||
| + | |||
| + | |'''Verification on gel :''' | ||
| + | |||
| + | - | ||
| + | |||
| + | |'''Verification on gel :''' | ||
| + | |||
| + | - | ||
| + | |'''Verification on gel :''' | ||
| + | |||
| + | - | ||
| + | |||
| + | |- style="background: #bebebe; text-align: center;" | ||
| + | |'''Purification on gel :''' | ||
| + | |||
| + | ok | ||
| + | |||
| + | |'''Purification on gel :''' | ||
| + | |||
| + | - | ||
| + | |||
| + | |'''Purification on gel :''' | ||
| + | |||
| + | - | ||
| + | |'''Purification on gel :''' | ||
| + | |||
| + | - | ||
| + | |||
| + | |||
| + | |||
| + | |- style="background: #d8d8d8; text-align: center;" | ||
| + | |'''digestion:''' | ||
| + | |||
| + | TatABCE | ||
| + | |||
| + | |'''digestion:''' | ||
| + | |'''digestion:''' | ||
| + | |'''digestion:''' | ||
| + | |||
| + | |- style="background: #bebebe; text-align: center;" | ||
| + | |'''verification digestion:''' | ||
| + | |||
| + | ok | ||
| + | |||
| + | |'''verification digestion:''' | ||
| + | |||
| + | - | ||
| + | |||
| + | |'''verification digestion:''' | ||
| + | |||
| + | - | ||
| + | |'''verification digestion:''' | ||
| + | |||
| + | - | ||
| + | |||
| + | |||
| + | |- style="background: #d8d8d8; text-align: center;" | ||
| + | |'''ligation:''' | ||
| + | |||
| + | TatABCE | ||
| + | |||
| + | |'''ligation:''' | ||
| + | |'''ligation:''' | ||
| + | |'''ligation:''' | ||
| + | |||
| + | |- style="background: #bebebe; text-align: center;" | ||
| + | |'''PCR colo :''' | ||
| + | |||
| + | ok | ||
| + | |||
| + | |'''PCR colo :''' | ||
| + | |||
| + | - | ||
| + | |||
| + | |'''PCR colo :''' | ||
| + | |||
| + | - | ||
| + | |'''PCR colo :''' | ||
| + | |||
| + | - | ||
| + | |||
| + | |- style="background: #d8d8d8; text-align: center;" | ||
| + | |'''miniprep:''' | ||
| + | |||
| + | STOPPED | ||
| + | |||
| + | |'''miniprep:''' | ||
| + | |||
| + | - | ||
| + | |||
| + | |'''miniprep:''' | ||
| + | |||
| + | - | ||
| + | |||
| + | |'''miniprep:''' | ||
| + | |||
| + | - | ||
| + | |||
| + | |- style="background: #bebebe; text-align: center;" | ||
| + | |'''sequencing :''' | ||
| + | |||
| + | - | ||
| + | |||
| + | |'''sequencing :''' | ||
| + | |||
| + | - | ||
| + | |||
| + | |'''sequencing :''' | ||
| + | |||
| + | - | ||
| + | |'''sequencing :''' | ||
| + | |||
| + | - | ||
| + | |||
| + | |} | ||
| + | |||
| + | ===functional testing of the part 2=== | ||
| + | |||
| + | Ptet RFP Plac TetR was tranforme into Top10 bacteria and grew on media : | ||
| + | |||
| + | |||
| + | 0,1% IPTG . | ||
| + | |||
| + | 1% glucose. | ||
| + | |||
| + | LB pure. | ||
[[Team:Paris/Production_testing_delay#top | return to the top]] | [[Team:Paris/Production_testing_delay#top | return to the top]] | ||
Latest revision as of 14:43, 18 October 2009
iGEM > Paris > Production > Testing > delay
Delay
We separate the construction on two plasmids :
Part1 : functional testing of the part 1
Part2 : functional testing of the part 2
part1
Time required : A lOT !!!!!
Experiments ran :
| column 1 | column 2 | column 3 | column 4
|
| PCR :
TatABCE matrix : | PCR : | PCR : | PCR : |
| Verification on gel :
ok | Verification on gel :
- | Verification on gel :
- | Verification on gel :
- |
| Purification on gel :
ok | Purification on gel :
- | Purification on gel :
- | Purification on gel :
-
|
| digestion:
TatABCE | digestion: | digestion: | digestion: |
| verification digestion:
ok | verification digestion:
- | verification digestion:
- | verification digestion:
-
|
| ligation:
TatABCE | ligation: | ligation: | ligation: |
| PCR colo :
ok | PCR colo :
- | PCR colo :
- | PCR colo :
- |
| miniprep:
STOPPED | miniprep:
- | miniprep:
- | miniprep:
- |
| sequencing :
- | sequencing :
- | sequencing :
- | sequencing :
- |
functional testing of the part 1
PBAD LacI LVA- YFP was transforme into Top ten Bacteria and grew on medium :
1% arabinose
1% glucose
LB pure
part2
Time required : A lOT !!!!!
Experiments ran :
| column 1 | column 2 | column 3 | column 4
|
| PCR :
TatABCE matrix : | PCR : | PCR : | PCR : |
| Verification on gel :
ok | Verification on gel :
- | Verification on gel :
- | Verification on gel :
- |
| Purification on gel :
ok | Purification on gel :
- | Purification on gel :
- | Purification on gel :
-
|
| digestion:
TatABCE | digestion: | digestion: | digestion: |
| verification digestion:
ok | verification digestion:
- | verification digestion:
- | verification digestion:
-
|
| ligation:
TatABCE | ligation: | ligation: | ligation: |
| PCR colo :
ok | PCR colo :
- | PCR colo :
- | PCR colo :
- |
| miniprep:
STOPPED | miniprep:
- | miniprep:
- | miniprep:
- |
| sequencing :
- | sequencing :
- | sequencing :
- | sequencing :
- |
functional testing of the part 2
Ptet RFP Plac TetR was tranforme into Top10 bacteria and grew on media :
0,1% IPTG .
1% glucose.
LB pure.
 "
"