Team:PKU Beijing/Project/AND Gate 1/Design
From 2009.igem.org
(→Mechanism) |
|||
| (23 intermediate revisions not shown) | |||
| Line 3: | Line 3: | ||
{{PKU_Beijing/Header2}} | {{PKU_Beijing/Header2}} | ||
[[Team:PKU_Beijing/Project|Project]] > [[Team:PKU_Beijing/Project/AND_Gate_1|AND Gate 1]] > [[Team:PKU_Beijing/Project/AND_Gate_1/Design|Design]] | [[Team:PKU_Beijing/Project|Project]] > [[Team:PKU_Beijing/Project/AND_Gate_1|AND Gate 1]] > [[Team:PKU_Beijing/Project/AND_Gate_1/Design|Design]] | ||
| - | |||
| - | + | ==='''Mechanism'''=== | |
| - | [[Image:PKU_AND_Gate_Mechanism.png|600px|center|thumb|fig1. '''The mechanism of the AND Gate''' | + | Many AND Gates have been designed and constructed recently. In our project, We adopted the design of an existing AND gate(Anderson et al. Molecular Systems Biology) and improved its efficiency |
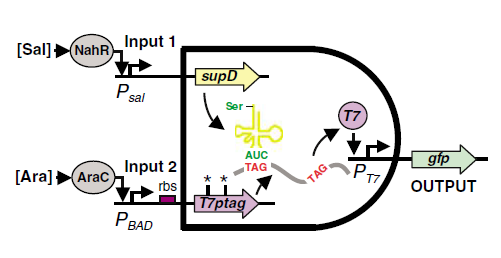
| + | [[Image:PKU_AND_Gate_Mechanism.png|600px|center|thumb|fig1. '''The mechanism of the AND Gate''' Anderson, Christopher JC Voigt CA and Arkin AP.''Environmental signal integration by a modular AND gate''. 2007,'''Molecular Systems Biology 3:133''']] | ||
| - | Fig.1 illustrates the mechanism of the design. One of the inputs of the | + | Fig.1 illustrates the mechanism of the design. One of the inputs of the AND gate leads to the expression of T7ptag mRNA which is a T7 polymerase coding sequence with amber mutations inside. The other input of the AND gate leads to the expression of supD tRNA, which acts as an amber mutation suppressor. The concurrence of the mRNA and the tRNA can make functional T7 polymerase, which can activate the downstream T7 promoter. |
| - | The AND Gate can be further decoupled into | + | The AND Gate can be further decoupled into an input sensor, a core, and an output. |
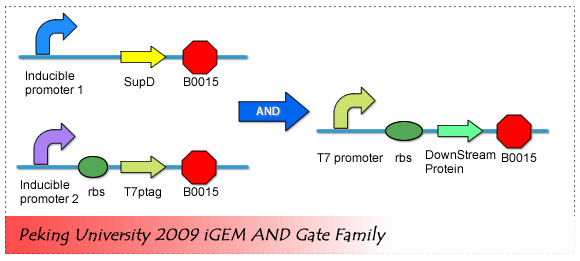
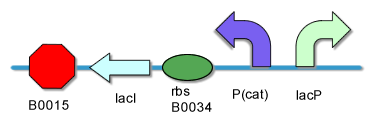
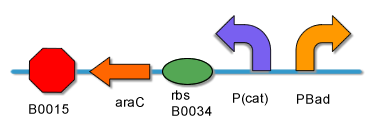
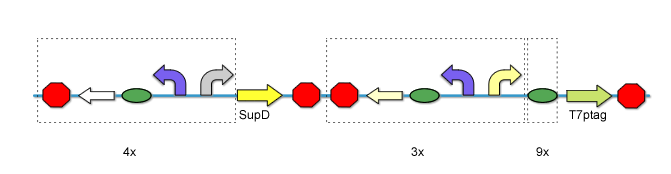
[[Image:PKU_AND_Gate.png|center|500px|frame|fig2. '''AND Gate Biological Components''']] | [[Image:PKU_AND_Gate.png|center|500px|frame|fig2. '''AND Gate Biological Components''']] | ||
| Line 52: | Line 52: | ||
The salicylate Sensor is not made up of standard parts, it is originally on the pSal-SupD plasmid that Voigt Lab mailed us. We got the sequence from NCIBI there is no standard enzymes inside of its sequence. We designed primers and PCR it from the plasmid. Then the promter is placed upstream of part E0840 to characterize its induction curve. | The salicylate Sensor is not made up of standard parts, it is originally on the pSal-SupD plasmid that Voigt Lab mailed us. We got the sequence from NCIBI there is no standard enzymes inside of its sequence. We designed primers and PCR it from the plasmid. Then the promter is placed upstream of part E0840 to characterize its induction curve. | ||
| - | ===''' | + | ==='''AND Gate 1 Module'''=== |
SupD tRNA and T7ptag mRNA makes the core of this AND Gate. | SupD tRNA and T7ptag mRNA makes the core of this AND Gate. | ||
| Line 68: | Line 68: | ||
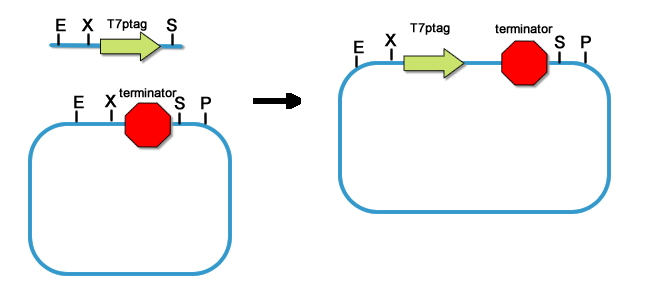
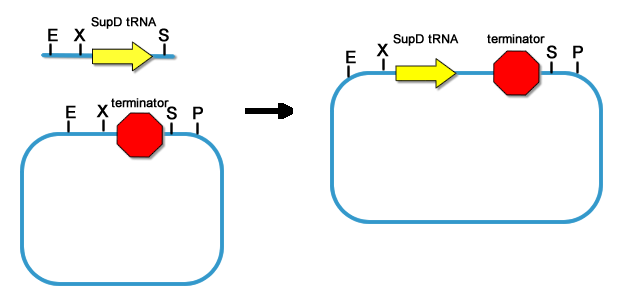
[[Image:PKU_T7ptag-term_into_rbs.png|300px|left|thumb|fig11. Placing rbs upstream of T7ptag-terminator]] | [[Image:PKU_T7ptag-term_into_rbs.png|300px|left|thumb|fig11. Placing rbs upstream of T7ptag-terminator]] | ||
<br><br><br><br> | <br><br><br><br> | ||
| - | For T7ptag, we clone it downstream of 9 different rbs. While SupD is just tRNA, we don’t need to add rbs. So far, the core of the AND Gate is constructed. | + | For T7ptag, we clone it downstream of 9 different rbs. While SupD is just tRNA, we don’t need to add rbs. So far, the core of the AND Gate is constructed. Before the AND Gate is functional, standard regulatory parts should be placed upstream of the T7ptag and SupD construct. |
| - | + | <br><br><br><br> | |
| + | In order to pass signal from this AND gate to downstream modules, a T7 promoter is needed as an output interface. So the downstream protein can be regulated by the T7 promoter. Which is described at the end of this page. | ||
<br><br><br><br><br><br> | <br><br><br><br><br><br> | ||
| - | ==='''Integration of | + | |
| + | ==='''Integration of the Sensor to the input interface of AND Gate 1'''=== | ||
Actually, the sensors we constructed is not all successful, for example, the aTc, IPTG sensor (tetP and LacP with its repressor constitutively expressed) is leaky and shows little induction. Later, we got four inducible promoter that shows induction in the presence of the inducer. The arabinose inducible promoter is successfully constructed, and the salicylate inducible promoter is also PCRed and well characterized, the LuxR system, although leaky, show a maximum induction of 8 fold. The lac promoter can only work in a E.coli Strain that has lacIq mutation, we chose JM109 strain. | Actually, the sensors we constructed is not all successful, for example, the aTc, IPTG sensor (tetP and LacP with its repressor constitutively expressed) is leaky and shows little induction. Later, we got four inducible promoter that shows induction in the presence of the inducer. The arabinose inducible promoter is successfully constructed, and the salicylate inducible promoter is also PCRed and well characterized, the LuxR system, although leaky, show a maximum induction of 8 fold. The lac promoter can only work in a E.coli Strain that has lacIq mutation, we chose JM109 strain. | ||
| Line 77: | Line 79: | ||
For those successfully constructed, either two of the inputs are combined to make an AND Gate. Theorectically, there are 12 combinations in total. | For those successfully constructed, either two of the inputs are combined to make an AND Gate. Theorectically, there are 12 combinations in total. | ||
| - | {|cellpadding= | + | {|cellpadding=10 style="background:yellow" |
| ||LacP-SupD||LuxP-SupD||PBad-SupD||SalP-SupD | | ||LacP-SupD||LuxP-SupD||PBad-SupD||SalP-SupD | ||
|- | |- | ||
| - | |LacP-T7ptag|||||| | + | |LacP-T7ptag||N/A||Shan Shen ||Shan Shen||Siheng He |
|- | |- | ||
| - | |LuxP-T7ptag||||||Rencheng Gao| | + | |LuxP-T7ptag||Not Made||N/A||Rencheng Gao||Not Made |
|- | |- | ||
| - | |PBad-T7ptag|| | + | |PBad-T7ptag||Shuke Wu ||Not Made||N/A||Haoqian Zhang |
|- | |- | ||
| - | |SalP-T7ptag|| | + | |SalP-T7ptag||Not Made||Not Made||Guosheng Zhang||N/A |
|} | |} | ||
| Line 93: | Line 95: | ||
We constructed and tested 8x9 of them, and finally got two AND Gates that best fits our need. | We constructed and tested 8x9 of them, and finally got two AND Gates that best fits our need. | ||
| - | ===''' | + | ==='''Directing the Output Interface'''=== |
| - | + | ||
| - | + | ||
| - | + | In our project, we need to pass signal from AND Gate 1 to CI repressor protein to trigger the bistable module. We also constructed T7 promoter-rbs-CI-terminator library for later system assembly. At the same time, in order to test our AND Gate, we constructed a T7 promoter-GFP reporter output. The AND Gate module is moved to the pSB4K5 plasmid, and we constructed a T7 promoter-GFP reporter on a Amp resistant and colE1 origin plasmid. By transforming both plasmids, and measured the fluorescence by plate reader after induction, we successfully selected two AND Gates that best fits our need. | |
{{PKU_Beijing/Foot}} | {{PKU_Beijing/Foot}} | ||
Latest revision as of 20:47, 21 October 2009
|
||||||||||||||||||||||||||||
|
||||||||||||||||||||||||||||
 "
"