Team:Paris/Addressing testing
From 2009.igem.org
(→Export system) |
(→Functional Testing:) |
||
| (2 intermediate revisions not shown) | |||
| Line 300: | Line 300: | ||
| - | PBAD ClyA RFP on PSB3T5 was transformed into Delta Tol bacteria , in this case we are supposed to see fluorescent vesicles | + | PBAD ClyA RFP on PSB3T5 was transformed into Delta Tol bacteria , in this case we are supposed to see fluorescent vesicles when the medium contains 1 % arabinose , and to have no fluorescent on 1% glucose. |
| + | |||
| + | |||
| + | To learn more, see the [[Team:Paris/Conclusion | Conclusion page]] | ||
| Line 357: | Line 360: | ||
ok | ok | ||
| - | |bgcolor="#E0E3FE"| | + | |bgcolor="#E0E3FE"| |
- | - | ||
| Line 383: | Line 386: | ||
|- style="background: #bebebe; text-align: center;" | |- style="background: #bebebe; text-align: center;" | ||
| - | |bgcolor="#F8F8F8"|''' | + | |bgcolor="#F8F8F8"|'''Digestion checking:''' |
none | none | ||
| - | |bgcolor="#E0E3FE"|''' | + | |bgcolor="#E0E3FE"|'''Digestion checking:''' |
none | none | ||
| - | |bgcolor="#CBD1FD"|''' | + | |bgcolor="#CBD1FD"|'''Digestion checking:''' |
none | none | ||
| - | |bgcolor="#BCCDFD"|''' | + | |bgcolor="#BCCDFD"|'''Digestion checking:''' |
none | none | ||
Latest revision as of 03:53, 22 October 2009
iGEM > Paris > WetLab > Addressing
WetLab - Addressing the message
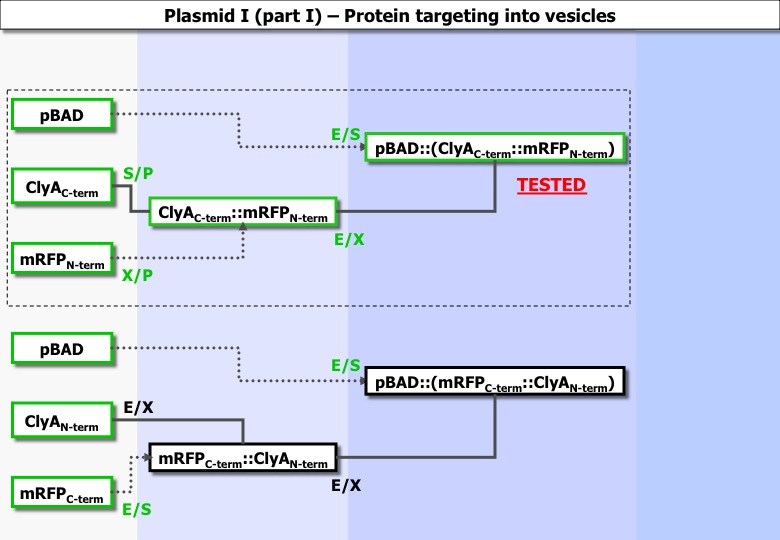
Global constructions :
Experiments ran :
| Column 1 | Column 2 | Column 3 | Column 4 |
| PCR :
pBAD: plate 2008
Oligo :O57 and O58 TM :
Oligo :O59 and O60 TM :
|
- |
- |
- |
| Verification on gel :
ok |
- |
- |
-
|
| Purification on gel :
ok |
-
|
- |
-
|
| Digestion:
ClyA Cterm S/P
| Digestion:
ClyA Cter-RFP Nter vector PSB1A3 E/X |
- |
- |
| Verification digestion:
ok | Verification digestion:
ok | - |
- |
| Ligation:
ClyA Cter-RFP Nter vector PSB1A3 | Ligation:
PBAD E/S ClyA Cter-RFP Nter PSB1A3 E/X (x2)
| Ligation:
- |
|
| Colony PCR :
ok | Colony PCR :
ok | Colony PCR :
ok |
|
| Miniprep:
clone : | Miniprep:
clone : | Miniprep:
clone 3 clone 5 |
|
| Sequencing :
ok | Sequencing :
ok | Sequencing :
ok |
-
|
| Stock glycerol:
ClyA CTer: S47(clone 3) and S48(clone 7) ClyA Nter: S72 RFP Cter: S55 (Clone 3) RFP Nter: S56 (Clone 3)
| Stock glycerol
Cly A(Cter)-(Nter)RFP: S72(clone 8) | Stock glycerol
pBAD ClyA RFP : S89 (clone 3) pBAD ClyA RFP: S90 (clone 5) |
|
Functional Testing:
PBAD ClyA RFP was transformed into Top10 bacteria in order to localize the fluorescence, we are supposed to have a superior fluorescence in the membrane.
PBAD ClyA RFP on PSB3T5 was transformed into Delta Tol bacteria , in this case we are supposed to see fluorescent vesicles when the medium contains 1 % arabinose , and to have no fluorescent on 1% glucose.
To learn more, see the Conclusion page
Export system
We finally thought that it won't be neccesary to overexpress the Tat system, nevertheless we have run a few experiments before starting to focus on others parts of the project.
Experiments ran :
| Column 1 | Column 2 | Column 3 | Column 4 |
| PCR :
TatABCE matrix : | - | - | -
|
| Verification on gel :
ok | - |
- |
- |
| Purification on gel :
ok |
- |
- |
-
|
| Digestion:
none | Digestion:
none | Digestion:
none | Digestion:
none |
| Digestion checking:
none | Digestion checking:
none
| Digestion checking:
none | Digestion checking:
none
|
| Ligation:
none | Ligation:
none | Ligation:
none | Ligation:
none |
| Colony PCR :
none | Colony PCR :
none | Colony PCR :
none | Colony PCR :
none |
| Miniprep:
none | Miniprep:
none | Miniprep:
none | Miniprep:
none |
| Sequencing :
none | Sequencing :
none | Sequencing :
none | Sequencing :
none |
| Stock glycerol:
none | Stock glycerol
none | Stock glycerol
none | Stock glycerol
none |

 "
"