Team:PKU Beijing/Project/AND Gate 1/Design
From 2009.igem.org
(Difference between revisions)
| Line 4: | Line 4: | ||
==='''Mechanism'''=== | ==='''Mechanism'''=== | ||
| + | |||
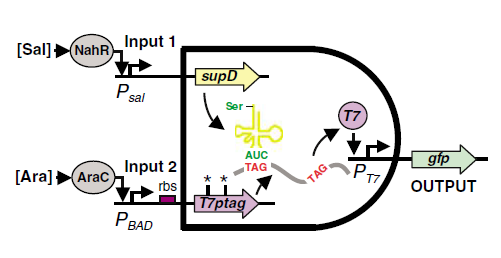
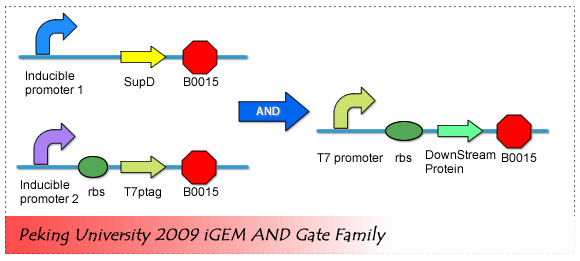
Many AND Gate have been designed and constructed in the past years. In our project, We adopted the design of Christopher Voigt, Which is published on ''Molecular Systems Biology''. | Many AND Gate have been designed and constructed in the past years. In our project, We adopted the design of Christopher Voigt, Which is published on ''Molecular Systems Biology''. | ||
| Line 16: | Line 17: | ||
==='''Sensor System (The input interface)'''=== | ==='''Sensor System (The input interface)'''=== | ||
| + | |||
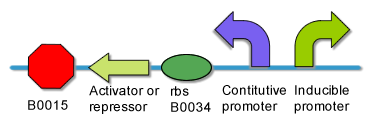
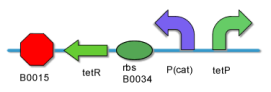
The input interface includes two inducible promoter which can be induced to expression by two different environmental small molecules. | The input interface includes two inducible promoter which can be induced to expression by two different environmental small molecules. | ||
| Line 43: | Line 45: | ||
And because the LuxR-LuxP sensor has already been constructed by iGEM2006_PennState. So we didn’t reverse the LuxR coding sequence but just use their construct. | And because the LuxR-LuxP sensor has already been constructed by iGEM2006_PennState. So we didn’t reverse the LuxR coding sequence but just use their construct. | ||
<br><br><br><br><br> | <br><br><br><br><br> | ||
| - | The salicylate | + | The salicylate Sensor: |
| - | The salicylate | + | The salicylate Sensor is not made up of standard parts, it is originally on the pSal-SupD plasmid that Voigt Lab mailed us. We got the sequence from NCIBI there is no standard enzymes inside of its sequence. We designed primers and PCR it from the plasmid. Then the promter is placed upstream of part E0840 to characterize its induction curve. |
| + | |||
| + | ==='''Core of the AND Gate'''=== | ||
| + | |||
| + | SupD tRNA and T7ptag mRNA makes the core of this AND Gate. | ||
| + | T7ptag gene is a T7 polymerase coding gene with two amber mutation at it Serine codon. The mutated T7 polymerase gene can only be transcribed but cannot be successfully expressed for the failure of translation. | ||
| + | The SupD tRNA is an amber mutation suppressor, this tRNA recognizes the TAG stop codon and mandatorily translate it into a Serine. Thus with supD, the function of the T7 polymerase gene is recovered. According to the Paper ''"Environmental signal integration by a modular AND Gate"'', SupD tRNA has little side effect on E.coli, although it can suppress normal termination of translation. | ||
| + | |||
| + | We Use PCR to make T7ptag and SupD into standard biobrick. The primer is designed according to the paper, we keep the complementary sequence in the original primer, and replace the enzyme cut site in the original primer with standard prefix and suffix. The original plasmid where we get SupD sequence and T7ptag sequence is shown below: | ||
| + | |||
| + | [[Image:PKU_Original_Plasmid.PNG|200px|thumb|fig8. The Original Plasmid Where We Got the SupD and T7ptag Gene]] | ||
| + | |||
{{PKU_Beijing/Foot}} | {{PKU_Beijing/Foot}} | ||
Revision as of 14:54, 10 October 2009
 "
"