Team:Groningen/Notebook/20 July 2009
From 2009.igem.org
Wet
GVP Cluster
Precipitation of the restriction fragments from GVP, J23109, J23100 and J23106
To concentrate BBa_J23109, BBa_J23100, BBa_J23106 and GVP it is first precipitated and then MQ is added.
Procedure:
- 100 μl absolute ethanol is added to ~50 μl Restriction fragment of GVP (12.1 ng/μL),J23109 (13.1 ng/μl), J23100 (11.9 ng/μl) and J23106 (11.1 ng/μl).
- Incubation -80°C for 1 h
- Centrifugation 30 min 0°C (14000rpm)
- Supernatant is removed
- Wash with 1000 μl 96% Ethanol (added ethanol and inverted a couple of times)
- Centrifugation 10 min 4°C (14000rpm)
- o/n air dried on bench top
Preparation of 1-14N and 1-1D for glycerol stocks, restiction analysis and 3A assembly
1-14N () and 1-1D () are inducible promoters, induced by L-arabinose and lactose (or IPTG) respectively.
- DNA from iGEM plates resuspended in 15 μL MQ and stored at -20°C
Transporters
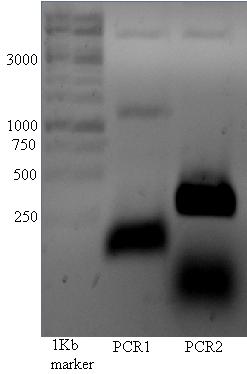
Below the PCR of 17/07, There seems to be a vague band at ~1150 in lane 1.
The band at ~1150 kb was cut out of the gel and used for pcr described below.
|
|
Metal Accumulation
Vectors
Positive control for the psB1AC3 vector done with both new and last years primers.
|
|
|
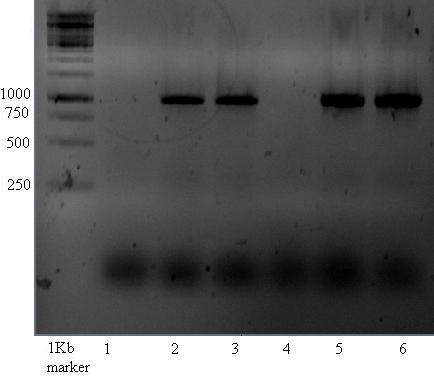
The gel shown below shows bands at about 1000 which is not what would be expected. Band of 315 were expected for lanes 1-6. --> contains ccdB gene, which would increase the expected fragment size to 991bp! This is about the size which is seen on gel....
vectors
Colony PCR on vectors for checking if promotors were inserted
- expected lengths (primer VF2 & VR) on:
psb1AC3 316 bp
Dry
Jasper looked at GlpF import, trying to create a prototype model in Simbiology by playing with the parameters. However, it turns out that Simbiology doesn't allow reactions using reactants from different compartments, so using a Michaelis-Menten equation (instead of using more fine-grained reactions) is pretty much the only option for transporters. Also, after discovering that it is quite hard to directly derive the required constants from the given graph manually (due to the different capacities of the cell and the solution) a table was made of the uptake graph (Fig. 1B) in Meng2004 by importing the graph in Inkscape and aligning the axes with its rulers.
|
|
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
|
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
 "
"