Team:Groningen/Notebook/22 July 2009
From 2009.igem.org
(→Vectors) |
(→Dry) |
||
| Line 117: | Line 117: | ||
==Dry== | ==Dry== | ||
| + | Jasper had a look at ArsD and Annelies at ArsAB, but at some point we realized that ArsA and ArsD might actually only be on some plasmids and may not be part of the genome of our E. coli (DH10B). To verify this we searched for a database which we could use for BLASTing ([http://blast.hgsc.bcm.tmc.edu/bcm/blast/microbialblast.cgi?organism=EcoliDH10B which we found]) and BLASTed the sequences of several ars genes. The results (which can be viewed at [[Team:Groningen/BLAST]]) indicate that our E. coli most likely indeed does NOT have ArsA and/or ArsD, it most likely only has ArsB, ArsC and ArsR. This should save us quite a bit of work! | ||
Today we worked on the efflux of arsenic by ArsA ArsB and ArsD. KB looked at ArsB first we looked at diffrent papers and we found some very promising ones In the afternoon we looked especially at As(III) and Sb(III) Uptake by GlpF and Efflux by ArsB in Escherichia coli by Yu-Ling Meng, Zijuan Liu, and Barry P. Rosen‡ [[http://www.jbc.org/cgi/content/abstract/M400037200v1 link]] | Today we worked on the efflux of arsenic by ArsA ArsB and ArsD. KB looked at ArsB first we looked at diffrent papers and we found some very promising ones In the afternoon we looked especially at As(III) and Sb(III) Uptake by GlpF and Efflux by ArsB in Escherichia coli by Yu-Ling Meng, Zijuan Liu, and Barry P. Rosen‡ [[http://www.jbc.org/cgi/content/abstract/M400037200v1 link]] | ||
Revision as of 15:36, 22 July 2009
Wet
GVP Cluster
Transporters
Metal Accumulation
Vectors
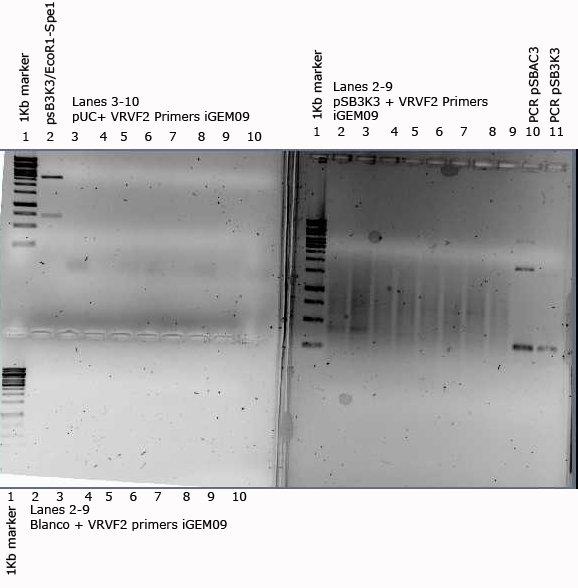
In order to check the VR VF2 primers that were ordered this year a gradient PCR was done with a vector that is expected to work , a pUC vector and a negative control with no vector.
|
|
As a check-up we also did a restrition analysis for the vector with EcoR1 and Spe1
| 10x fast digest buffer | 2 uL |
| pSB3K3 (22,5 ngr/uL) | 8 uL (=180ngr up to 1ugr is recommended at fermentas) |
| EcoR1 | 1 uL |
| Spe1 | 1 uL |
| MilliQ | 8 uL |
| Total | 20 uL |
- Mix gently and spin down
- incubate at 37° for 30 min.
Results of the PCR and restiction shown below. Bands were expected in the pSB3K3 lanes with VR-VF2 primers at 316bps. A band somewhat larger than 316 can be seen in lanes 2,3 (50°, 51.5°).
The expected band for the digestion is 2727 bps, which is about the size of the band shown in the figure above.
Dry
Jasper had a look at ArsD and Annelies at ArsAB, but at some point we realized that ArsA and ArsD might actually only be on some plasmids and may not be part of the genome of our E. coli (DH10B). To verify this we searched for a database which we could use for BLASTing (which we found) and BLASTed the sequences of several ars genes. The results (which can be viewed at Team:Groningen/BLAST) indicate that our E. coli most likely indeed does NOT have ArsA and/or ArsD, it most likely only has ArsB, ArsC and ArsR. This should save us quite a bit of work!
Today we worked on the efflux of arsenic by ArsA ArsB and ArsD. KB looked at ArsB first we looked at diffrent papers and we found some very promising ones In the afternoon we looked especially at As(III) and Sb(III) Uptake by GlpF and Efflux by ArsB in Escherichia coli by Yu-Ling Meng, Zijuan Liu, and Barry P. Rosen‡ [link] Escherichia coli* what it's influence is on the efflux of As and Sb, the influence of NADH, the influence of FCCP and the influences As and Sb on each other.
|
|
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
|
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
 "
"