User:DavidC/14 September 2009
From 2009.igem.org
(New page: {{Template:Supbiotechcss6.css}} {{Template:SupbiotechparisEn}} <center> <div style="color: black; margin-bottom: 30px; width: 500px;"> {|align="center" ||{{#calendar: title=User:DavidC |...) |
|||
| (One intermediate revision not shown) | |||
| Line 12: | Line 12: | ||
</div> | </div> | ||
</center> | </center> | ||
| + | |||
| + | |||
| + | === Monday the 14th === | ||
| + | |||
| + | ==== Ligation: samples digested on the 13th of September ==== | ||
| + | |||
| + | Ligation pairs: plasmid / insert : <br> | ||
| + | |||
| + | BBa_B0014 / BBa_C0012 <br> | ||
| + | |||
| + | First report: <br> | ||
| + | Plasmid = 2µL <br> | ||
| + | Insert = 5µL <br> | ||
| + | |||
| + | Second report: <br> | ||
| + | Plasmid = 1,5µL <br> | ||
| + | Insert = 5µL <br> | ||
| + | |||
| + | Third report: <br> | ||
| + | Plasmide = 1µL <br> | ||
| + | Insert = 7µL <br> | ||
| + | |||
| + | ==== NEB Enzymes ==== | ||
| + | |||
| + | For each samples add sterilized water to obtain a maximum volume equals to 8µL. <br> | ||
| + | Ligation mix (NEB): <br> | ||
| + | 1,5µL T4 ligase + 1,5µL T4 ligase buffer. <br> | ||
| + | Add 2µL / tube. <br> | ||
| + | Incubation 1h at RT. <br> | ||
| + | |||
| + | ==== TAKARA DNA ligation kit === | ||
| + | |||
| + | First report: | ||
| + | Solution A (Buffer) = 28µL <br> | ||
| + | Solution B (Enzyme) = 7µL <br> | ||
| + | Incubation 1h at 16°C. <br> | ||
| + | |||
| + | ==== Electroporation ==== | ||
| + | |||
| + | Electroporation cuvettes = 2mm ; inoculums of electrocompetent <i>E.coli</i> DH5alpha= 40µL; pulse = 2,5KVolt ; 1h of incubation. <br> | ||
| + | |||
| + | Spread 1mL of inoculums into a petri dish with LB + ampicillin (50mg/mL) (20/0,02mL). <br> | ||
| + | |||
| + | ==== PCR ==== | ||
| + | |||
| + | ==== D protein ==== | ||
| + | |||
| + | Primers: <br> | ||
| + | Forward : Préfixe ATG BBa <br> | ||
| + | Reverse : Prt D_Rv_Bal1 <br><br> | ||
| + | |||
| + | ADN : <br> | ||
| + | 1. Lambda bacteriophage; <br> | ||
| + | 2. Purified and amplified D protein. <br><br> | ||
| + | |||
| + | Cycles: <br> | ||
| + | 94°C 2minutes; <br> | ||
| + | (94°C 1 minute; <br> | ||
| + | 46°C 1 minutes; <br> | ||
| + | 72°C 1 minute(s)) X 35; <br> | ||
| + | 72°C 4 minutes; <br> | ||
| + | |||
| + | ==== Penton base from the adenovirus 5 ==== | ||
| + | |||
| + | Primers: <br> | ||
| + | Forward : ADV5Fw_Bal1 <br> | ||
| + | Reverse : ADV5Rv_Bal1 <br><br> | ||
| + | |||
| + | ADN : <br> | ||
| + | Purified and amplified penton base from plasmid coding for adenovirus 5. <br><br> | ||
| + | |||
| + | Cycles: <br> | ||
| + | 94°C 2minutes; <br> | ||
| + | (94°C 1 minute; <br> | ||
| + | 57,3°C 1 minutes; <br> | ||
| + | 72°C 2 minute(s)) X 35; <br> | ||
| + | 72°C 5 minutes; <br> | ||
| + | |||
| + | ==== DNA electrophoresis ==== | ||
| + | |||
| + | 85 Volt, 15 minutes. <br> | ||
| + | 105 Volt, 40 minutes. <br> | ||
| + | Ladder fermentas 1 Kb. <br><br> | ||
| + | |||
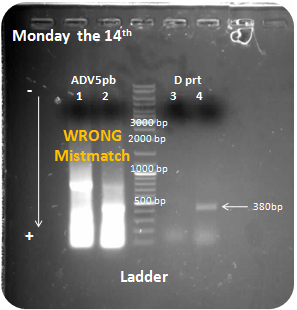
| + | Samples: Adenovirus penton base, D protein <br> | ||
| + | |||
| + | [[image:M1409.png|center]] | ||
| + | |||
| + | |||
| + | Interpretation: <br> | ||
| + | |||
| + | We obtain a small amplification for D protein on the sample 4. But other samples have bad results, we obtain to much DNA for samples 1 and 2 PCR, we also amplified the wrong DNA, maybe we use a bad temperature for primers annealing, finally sample 3 is negative. <br> | ||
Latest revision as of 14:39, 21 October 2009
Contents |
Monday the 14th
Ligation: samples digested on the 13th of September
Ligation pairs: plasmid / insert :
BBa_B0014 / BBa_C0012
First report:
Plasmid = 2µL
Insert = 5µL
Second report:
Plasmid = 1,5µL
Insert = 5µL
Third report:
Plasmide = 1µL
Insert = 7µL
NEB Enzymes
For each samples add sterilized water to obtain a maximum volume equals to 8µL.
Ligation mix (NEB):
1,5µL T4 ligase + 1,5µL T4 ligase buffer.
Add 2µL / tube.
Incubation 1h at RT.
= TAKARA DNA ligation kit
First report:
Solution A (Buffer) = 28µL
Solution B (Enzyme) = 7µL
Incubation 1h at 16°C.
Electroporation
Electroporation cuvettes = 2mm ; inoculums of electrocompetent E.coli DH5alpha= 40µL; pulse = 2,5KVolt ; 1h of incubation.
Spread 1mL of inoculums into a petri dish with LB + ampicillin (50mg/mL) (20/0,02mL).
PCR
D protein
Primers:
Forward : Préfixe ATG BBa
Reverse : Prt D_Rv_Bal1
ADN :
1. Lambda bacteriophage;
2. Purified and amplified D protein.
Cycles:
94°C 2minutes;
(94°C 1 minute;
46°C 1 minutes;
72°C 1 minute(s)) X 35;
72°C 4 minutes;
Penton base from the adenovirus 5
Primers:
Forward : ADV5Fw_Bal1
Reverse : ADV5Rv_Bal1
ADN :
Purified and amplified penton base from plasmid coding for adenovirus 5.
Cycles:
94°C 2minutes;
(94°C 1 minute;
57,3°C 1 minutes;
72°C 2 minute(s)) X 35;
72°C 5 minutes;
DNA electrophoresis
85 Volt, 15 minutes.
105 Volt, 40 minutes.
Ladder fermentas 1 Kb.
Samples: Adenovirus penton base, D protein
Interpretation:
We obtain a small amplification for D protein on the sample 4. But other samples have bad results, we obtain to much DNA for samples 1 and 2 PCR, we also amplified the wrong DNA, maybe we use a bad temperature for primers annealing, finally sample 3 is negative.
 "
"