Team:Groningen/Modelling/Arsenic
From 2009.igem.org
The raw model
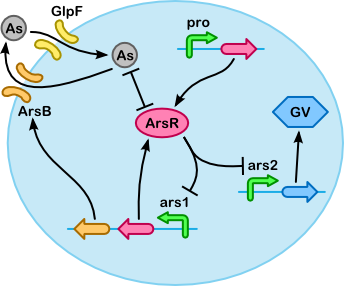
The following variables play an important role in our system (these can be concentrations of substances, the density of the cell, etc.):
- Extracellular:
- As(III)ex
- Membrane (all naturally occurring, but we plan to bring GlpF to overexpression):
- GlpF (concentration w.r.t. the exterior of the cell)
- GlpFAs (concentration w.r.t. the exterior of the cell)
- ArsB (concentration w.r.t. the interior of the cell)
- ArsBAs (concentration w.r.t. the interior of the cell)
- Intracellular (ars2, pro and GV are introduced):
- As(III)
- ars1 (concentration of unbound promoters naturally occurring in E. coli)
- ars2 (concentration of unbound promoters in front of gas vesicle genes)
- pro (concentration of constitutive promoters in front of arsR)
- ArsR ArsR binds to ars to repress production of the genes they regulate, and binds to As(III) to make it less of a problem for the cell.i
- ArsRAs (bound to As(III))
- ArsRars1 (bound to ars1)
- ArsRars2 (bound to ars2)
- GV (concentration of gas vesicles)
The variables above can be related to each other through the following "reactions" (color coding is continued below to show which parts of the differential equations refer to which groups of reactions):
- Transport (based on Rosen1996, Meng2004 and Rosen2009)
- As(III)ex + GlpF ↔ GlpFAs
- GlpFAs → GlpF + As(III)
- As(III) + ArsB ↔ ArsBAs
- ArsBAs → ArsB + As(III)ex
- ArsB → null (degradation)
- Accumulation (based on Chen1997)
- As(III) + ArsR ↔ ArsRAs
- ars1 + 2 ArsR ↔ ArsRars1
- ars2 + 2 ArsR ↔ ArsRars2
- ars1 → ars1 + ArsR + ArsB (transcription + translation)
- ars2 → ars2 + GV (transcription + translation)
- pro → pro + ArsR (transcription + translation)
- ArsR → null (degradation)
- GV → null (degradation)
Resulting in the following differential equations (please note that some can be formed by linear combinations of the others), using color coding to show the correspondence to the reactions above:
- (d/dt) As(III)ex = -k5on As(III)ex GlpF + k5off GlpFAs + (Vc/Vs) k8 ArsBAs
- (d/dt) GlpF = - (d/dt) GlpFAs
- (d/dt) GlpFAs = k5on As(III)ex GlpF - (k5off+k6) GlpFAs
- (d/dt) ArsB = - (d/dt) ArsBAs + β4 ars1 - ln(2)/τB ArsB
- (d/dt) ArsBAs = k7on As(III) ArsB - (k7off+k8) ArsBAs
- (d/dt) As(III) = - (d/dt) ArsRAs - k7on As(III) ArsB + k7off ArsBAs + (Vs/Vc) k6 GlpFAs
- (d/dt) ars1 = - (d/dt) ArsRars1
- (d/dt) ars2 = - (d/dt) ArsRars2
- (d/dt) ArsR = β1 ars1 + β3 pro - (ln(2)/τR) ArsR - (d/dt) ArsRAs - 2 (d/dt) ArsRars1 - 2 (d/dt) ArsRars2
- (d/dt) ArsRAs = k1on ArsR As(III) - k1off ArsRAs
- (d/dt) ArsRars1 = k3on ArsR² ars1 - k3off ArsRars1
- (d/dt) ArsRars2 = k3on ArsR² ars2 - k3off ArsRars2
- (d/dt) GV = β5 ars2 - ln(2)/τG GV
Using the following constants/definitions:
| Name | Units | Description |
|---|---|---|
| k1on, k2on, etc. | 1/(M·s) | Reaction rate constants for reactions to a complex. |
| k1off, k2off, etc. | 1/s | Reaction rate constants for reactions from a complex. |
| k6, k8 | 1/s | Reaction rate constants representing how fast transporters transport their cargo to "the other side". |
| K1d - K4d | M | Dissociation constants. |
| τR, τG | s | Half-lifes (of ArsR and GV, respectively). Degradation rate = ln(2)/τ If you take just the degradation into account you will have the equation dC/dt = -k*C, which leads to C(t) = C(0) e-k t. So if k = ln(2)/τ we get C(t) = C(0) e-ln(2)/τ t = C(0) 2-t/τ. In other words τ is the time it takes for the concentration to half. i |
| β1, β2, etc. | 1/s | Production rates.
|
| Vs | L | Volume of solution (excluding cells). |
| Vc | L | Total volume of cells (in solution) (so Vs+Vc is the total volume). |
| v5, v7 | mol/(s·L) | Maximum reaction rates per liter of cells (see Michaelis-Menten equation). Note that we have purposefully chosen to write the units as mol/(s·L) instead of M/s, to emphasize the fact that it the rates are per liter of cells.
|
| v7'·ars | Reaction rate when As(III) = K7
| |
| K5, K7 | M | Concentration at which the reaction reaches half its maximum reaction rate (see Michaelis-Menten equation).
|
See Chen1997 for the interplay between ArsR and ArsD (the latter has a role similar to ArsR, but we do not treat it, as it is not present in our system).
Quasi-steady-state
When there are many molecules "waiting" to be transported and/or the concentrations in the cell and outside the cell are relatively slow changing compared to it is not unreasonable to assume that the amount of bound transporters is constant. Together with the assumption that the total amount of transporters is constant this leads to the well-known Michaelis-Menten equation (focussing just on import):
0 = (d/dt) GlpFAs = k5on As(III)ex GlpF - (k5off+k6) GlpFAs
k5on As(III)ex GlpF = (k5off+k6) GlpFAs
GlpF = (k5off+k6)/k5on GlpFAs / As(III)ex
GlpF = K5 GlpFAs / As(III)ex
GlpFT = GlpF + GlpFAs
GlpFT = GlpFAs (K5 / As(III)ex + 1)
GlpFAs = GlpFT / (K5 / As(III)ex + 1)
GlpFAs = GlpFT As(III)ex / (K5 + As(III)ex)
(d/dt) As(III) = (Vs/Vc) k6 GlpFAs
= (Vs/Vc) k6 GlpFT As(III)ex / (K5 + As(III)ex)
= v5 As(III)ex / (K5 + As(III)ex)
For export the situation is complicated by the fact that ArsB is also produced by the fact that ArsB is produced by the ars operon. However, by looking at the total amount of ArsB and assuming that ArsBAs reaches an equilibrium quickly, we can derive (focussing just on export)
0 = (d/dt) ArsBAs = k7on As(III) ArsB - (k7off+k8) ArsBAs
k7on As(III) ArsB = (k7off+k8) ArsBAs
ArsB = (k7off+k8) ArsBAs / (k7on As(III))
ArsB = K7 ArsBAs / As(III)
ArsBT = ArsB + ArsBAs
ArsBT = K7 ArsBAs / As(III) + ArsBAs
ArsBT = ArsBAs (1 + K7 / As(III))
ArsBAs = ArsBT / (1 + K7 / As(III))
ArsBAs = ArsBT As(III) / (As(III) + K7)
ArsB = ArsBT - ArsBAs
= ArsBT (1 - As(III)/(As(III) + K7))
= ArsBT ((As(III) + K7)/(As(III) + K7) - As(III)/(As(III) + K7))
= ArsBT K7/(As(III) + K7)
(d/dt) As(III)ex = (Vc/Vs) k8 ArsBAs
= (Vc/Vs) k8 ArsBT As(III) / (As(III) + K7)
ArsBT = ArsB + ArsBAs
(d/dt) ArsBT = (d/dt) ArsB + (d/dt) ArsBAs
= - (d/dt) ArsBAs + β4 ars1 - ln(2)/τB ArsB + (d/dt) ArsBAs
= β4 ars1 - ln(2)/τB ArsB
= β4 ars1 - ln(2)/τB K7/(As(III) + K7) ArsBT
Similarly, for the protein production related equations we assume that the reactions that bind/unbind the promoters are very fast compared to protein production. To simplify the equations we first group the two ars promoters (we assume the two promoters are bound equally often) and introduce ArsRT (the total amount of ArsR):
arsT = ars + ArsRars
ars1 / ars1T = ars2 / ars2T
ars = ars1 + ars2
ars = ars1 (1 + ars2T / ars1T)
ars1 = ars / (1 + ars2T / ars1T)
ars1 = ars ars1T / arsT
ars2 = ars ars2T / arsT
(d/dt) ars = - (d/dt) ArsRars
(d/dt) ArsRars = k3on ArsR² ars - k3off ArsRars
(d/dt) GV = β5 ars ars2T/arsT - ln(2)/τG GV
ArsRT = ArsR + ArsRAs + 2 ArsRars
(d/dt) ArsRT = (d/dt) ArsR + (d/dt) ArsRAs + 2 (d/dt) ArsRars
= β1 ars ars1T/arsT + β3 pro - (ln(2)/τR) ArsR
Now, by setting the derivatives of ars and ArsRars to zero we can derive the following equations for the derivatives of ArsRT and GV:
0 = (d/dt) ArsRars = k3on ArsR² ars - k3off ArsRars
k3on ArsR² ars = k3off ArsRars
ArsRars = ArsR² ars / K3d²
arsT = ars + ArsRars
arsT = ars + ArsR² ars / K3d²
arsT = ars (1 + ArsR² / K3d²)
ars = arsT / (1 + ArsR² / K3d²)
(d/dt) ArsRT = β1 ars ars1T/arsT + β3 pro - (ln(2)/τR) ArsR
= β1 ars1T / (1 + ArsR² / K3d²) + β3 pro - (ln(2)/τR) ArsR - (d/dt) ArsRAs
(d/dt) GV = β5 ars ars2T/arsT - ln(2)/τG GV
= β5 ars2T / (1 + ArsR² / K3d²) - ln(2)/τG GV
By looking at the total arsenic content in the cell these equations can be further simplified:
As(III)T = As(III) + ArsRAs
(d/dt) As(III)T = (
0 = (d/dt) ArsRAs = k1on ArsR As(III) - k1off ArsRAs
k1on ArsR As(III) = k1off ArsRAs
ArsRAs = ArsR As(III) / K1d
As(III)T = As(III) + ArsRAs
= As(III) + ArsR As(III) / K1d
= As(III) (1 + ArsR / K1d)
Obviously any increase in concentration (of arsenic) in the cell is accompanied by a proportional decrease in concentration outside the cell (and vice versa). So:
(d/dt) As(III)ex = -(Vc/Vs) v5 As(III)ex / (K5 + As(III)ex)
Transport
By looking at the system in equilibrium we can more easily assess the impact of parameters and derive formulas for obtaining them. To this end we consider the system when all derivatives are zero (just taking the equations relevant for transport):
0 = (d/dt) As(III)ex = -k5on As(III)ex GlpF + k5off GlpFAs + (Vc/Vs) k8 ArsBAs 0 = (d/dt) GlpF = -k5on As(III)ex GlpF + (k5off+k6) GlpFAs 0 = (d/dt) GlpFAs = k5on As(III)ex GlpF - (k5off+k6) GlpFAs 0 = (d/dt) ArsB = -k7on As(III) ArsB + (k7off+k8) ArsBAs + β4 ars - ln(2)/τB ArsB 0 = (d/dt) ArsBAs = k7on As(III) ArsB - (k7off+k8) ArsBAs 0 = (d/dt) As(III) = -k7on As(III) ArsB + k7off ArsBAs + (Vs/Vc) k6 GlpFAs
The third equation is equivalent to the second and the fifth can be eliminated from the fourth, leading to:
0 = -k5on As(III)ex GlpF + k5off GlpFAs + (Vc/Vs) k8 ArsBAs 0 = -k5on As(III)ex GlpF + (k5off+k6) GlpFAs 0 = β4 ars - ln(2)/τB ArsB 0 = k7on As(III) ArsB - (k7off+k8) ArsBAs 0 = -k7on As(III) ArsB + k7off ArsBAs + (Vs/Vc) k6 GlpFAs
By combining the last two equations a rather obvious equation can be derived that essentially expresses that the import rate equals the export rate:
0 - 0 = (-k7on As(III) ArsB + k7off ArsBAs + (Vs/Vc) k6 GlpFAs)
+ ( k7on As(III) ArsB - (k7off+k8) ArsBAs)
0 = (Vs/Vc) k6 GlpFAs - k8 ArsBAs
k8 ArsBAs = (Vs/Vc) k6 GlpFAs
Using the equations above (and the fact that the total amount of importers doesn't change in our model) GlpFAs can be expressed as follows (working towards a Michaelis-Menten equation):
k5on As(III)ex GlpF = (k5off+k6) GlpFAs
GlpF = GlpFAs (k5off+k6) / (k5on As(III)ex)
GlpFT = GlpF + GlpFAs
GlpFT = GlpFAs (1 + (k5off+k6) / (k5on As(III)ex))
GlpFAs = GlpFT / (1 + (k5off+k6) / (k5on As(III)ex))
GlpFAs = GlpFT As(III)ex / (As(III)ex + K5)
Without ArsB regulation
If we disregard ArsB regulation ArsBAs can be determined similarly to GlpFAs:
ArsBAs = ArsBT As(III) / (As(III) + K7)
By substituting the above equations for GlpFAs and ArsBAs we can now derive a relation between the extracellular and intracellular concentrations of arsenic (where we recognize the constants of the well-known Michaelis-Menten equation):
k8 ArsBAs = (Vs/Vc) k6 GlpFAs
k8 ArsBT As(III) / (As(III) + K7) = (Vs/Vc) k6 GlpFT As(III)ex / (As(III)ex + K5)
v7 As(III) / (As(III) + K7) = v5 As(III)ex / (As(III)ex + K5)
v7 As(III) (As(III)ex + K5) = v5 As(III)ex (As(III) + K7)
As(III) (v7 As(III)ex + v7 K5 - v5 As(III)ex) = v5 K7 As(III)ex
As(III) = v5 K7 As(III)ex / (v7 As(III)ex + v7 K5 - v5 As(III)ex)
As(III) = v5 K7 As(III)ex / ((v7 - v5) As(III)ex + v7 K5)
An important flaw in this model is that the production of ArsB is dependent on the concentration of arsenic in the cell (via regulation by ArsR, see our transport page). This could be one of the reasons that this model is unable to fit the curve shown in figure 3A in Kostal2004 (if we try a least squares fit with the equation above, filling in v5 and K5 from what we computed for figure 1B in Meng2004, we get negative values for v7 and K7). (And something similar is true of figure 1 from Singh2008.) This led us to consider the regulation of ArsB, as is described in the following section.
With ArsB regulation
As explained above ArsBAs needs a slightly different approach, as ArsB can be produced in response to arsenic ArsBT is not a (known) constant and we have a formula dependent on ars instead:
β4 ars = ln(2)/τB ArsB
ArsB = (β4 τB/ln(2)) ars
(k7off+k8) ArsBAs = k7on As(III) ArsB
(k7off+k8) ArsBAs = (β4 τB/ln(2)) ars k7on As(III)
ArsBAs = (β4 τB/ln(2)) ars (k7on/(k7off+k8)) As(III)
ArsBAs = (β4 τB/ln(2)) ars As(III) / K7
This now leads to the following relation between intra- and extracellular As(III), note that v7' (in contrast to v7) is not the maximum export rate (it has units of 1/second):
k8 ArsBAs = (Vs/Vc) k6 GlpFAs
k8 (β4 τB/ln(2)) ars As(III) / K7 = (Vs/Vc) k6 GlpFT As(III)ex / (As(III)ex + K5)
v7'/K7 ars As(III) = v5 As(III)ex / (As(III)ex + K5)
As(III) = v5/(v7' ars) K7 As(III)ex / (As(III)ex + K5)
To get rid of the unknown ars in this equation we can use two equations that are derived below for accumulation:
0 = β1 ars + β3 pro - (ln(2)/τR) ArsR
(ln(2)/τR) ArsR = β1 ars + β3 pro
ArsR = (β1 ars + β3 pro) (τR/ln(2))
ars = arsT/(1 + ArsR²/K3d²)
arsT = (1 + ArsR²/K3d²) ars
arsT = ars + ArsR² ars / K3d²
0 = K3d² ars + ArsR² ars - K3d² arsT
0 = K3d² ars + (β1 ars + β3 pro)² (τR/ln(2))² ars - K3d² arsT
0 = K3d² ars + (β1 ars + β3 pro) (β1 ars² + β3 pro ars) (τR/ln(2))² - K3d² arsT
0 = K3d² ars + β1 ars β1 ars² (τR/ln(2))² + β1 ars β3 pro ars (τR/ln(2))² + β3 pro β1 ars² (τR/ln(2))² + β3 pro β3 pro ars (τR/ln(2))² - K3d² arsT
0 = β1² (τR/ln(2))² ars³ + 2 β1 β3 (τR/ln(2))² pro ars² + (β3² (τR/ln(2))² pro² + K3d²) ars - K3d² arsT
According to Mathematica's solution of Reduce[eq && K3d > 0 && arsT >= 0 && pro >= 0 && β1 > 0 && β3 > 0 && τR > 0, ars, Reals] (where eq is the equation above) there is only one real root, so we solve the equation safely using Newton's (or Halley's) method.
Accumulation
For many purposes, like determining the total amount of accumulated arsenic, it can be quite useful to consider the system in equilibrium. That is, when the derivatives of all variables to time are zero (just taking the equations relevant for accumulation here, assuming the As(III) concentration to be constant):
0 = (d/dt) ArsR = β1 ars + β3 pro - (ln(2)/τR) ArsR - (d/dt) ArsRAs - 2 (d/dt) ArsRars 0 = (d/dt) ArsRAs = k1on ArsR As(III) - k1off ArsRAs 0 = (d/dt) ArsRars = k3on ArsR² ars - k3off ArsRars
By eliminating the last two equations from the first and dividing the last two by k?on we are left with:
0 = β1 ars + β3 pro - (ln(2)/τR) ArsR 0 = ArsR As(III) - K1d ArsRAs 0 = ArsR² ars - K3d² ArsRars
Using the fact that the "concentration" of operators remains constant the last equation can be used to derive an equation for ars:
0 = ArsR² ars - K3d² ArsRars
ArsRars = ars ArsR² / K3d²
arsT = ars + ArsRars
arsT = ars + ars ArsR² / K3d²
arsT = ars (1 + ArsR²/K3d²)
ars = arsT/(1 + ArsR²/K3d²)
This leads to the following:
0 = β1 ars + β3 pro - (ln(2)/τR) ArsR 0 = β1 arsT + (β3 pro - (ln(2)/τR) ArsR) (ArsR²/K3d² + 1) 0 = β1 arsT + β3 pro ArsR²/K3d² + β3 pro - (ln(2)/τR) ArsR³/K3d² - (ln(2)/τR) ArsR 0 = K3d² (τR/ln(2)) β1 arsT + (τR/ln(2)) β3 pro ArsR² + K3d² (τR/ln(2)) β3 pro - ArsR³ - K3d² ArsR 0 = ArsR³ - (τR/ln(2)) β3 pro ArsR² + K3d² ArsR - K3d² (τR/ln(2)) (β1 arsT + β3 pro)
According to Mathematica's solution of Reduce[eq && K3d > 0 && arsT >= 0 && pro >= 0 && β1 > 0 && β3 > 0 && τR > 0, ArsR, Reals] (where eq is the equation shown above) there is only one real solution, so we can find it safely using Newton's (or Halley's) method.
Analogously to the derivation of the equation for ars an equation can be derived for the fraction of unbound arsenic in the cell:
As(III)T = As(III) + ArsRAs As(III)/As(III)T = 1/(ArsR/K1d + 1)
 "
"
