Team:Groningen/Notebook/11 August 2009
From 2009.igem.org
(→GVP Cluster) |
(→Vectors) |
||
| (7 intermediate revisions not shown) | |||
| Line 5: | Line 5: | ||
===GVP Cluster=== | ===GVP Cluster=== | ||
| - | :→ {{ | + | :→ {{done}} isolate plasmids from overnight precultures |
| - | :→ {{ | + | :→ {{done}} plate on cultures for storage in fridge (amp) |
:→ {{todo}} cut plasmid and GVP containing plasmid for ligation (ligation tomorrow) | :→ {{todo}} cut plasmid and GVP containing plasmid for ligation (ligation tomorrow) | ||
:→ {{todo}} purify wanted fragments (tomorrow) | :→ {{todo}} purify wanted fragments (tomorrow) | ||
| - | :→ {{ | + | :→ {{done}} make o.n. precultures from plate colonies |
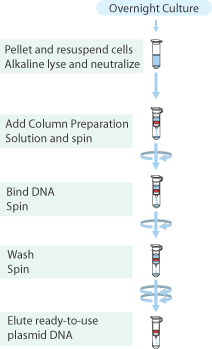
[[Image:Plasmid Miniprep Kit Flow Protocol.gif|thumb|150px| www.sigmaaldrich.com]] | [[Image:Plasmid Miniprep Kit Flow Protocol.gif|thumb|150px| www.sigmaaldrich.com]] | ||
| Line 51: | Line 51: | ||
|No | |No | ||
|} | |} | ||
| + | |||
| + | :→ These plasmids come from the glycerol stocks contaning E.coli TOP10 cells with iGEM plasmid [http://partsregistry.org/wiki/index.php?title=Part:BBa_J61002 J61002] and the [http://partsregistry.org/wiki/index.php?title=Part:BBa_J23100 high], [http://partsregistry.org/wiki/index.php?title=Part:BBa_J23100 medium] and [http://partsregistry.org/wiki/index.php?title=Part:BBa_J23100 low] constitutive promoters with RFP as reporter gene behind them. | ||
| + | |||
| + | :→ The GVP cluster can be ligated behind the promoters and self ligation of RFP will turn unwanted colonies red! | ||
| + | |||
| + | '''Plates''' | ||
| + | |||
| + | LB-agar plates with 50 μg/mL ampicillin were freshly prepared, and one drop of each o.n. preculture was spotted and subsequently swiped three times with sterile needle. The plates were stored at 37°C overnight. | ||
| + | |||
| + | '''Precultures''' | ||
| + | |||
| + | Over night cultures in 5mL LB-amp<sub>50</sub> medium were prepared from the following colonies: | ||
| + | |||
| + | → E.coli TOP10 pSB1AC3-med. const. promoter - GVP (amp.) (4x) (1 to 4) | ||
| + | |||
| + | → E.coli TOP10 pSB1AC3-low const. promoter - GVP (amp.) (4x) (5 to 8) | ||
| + | |||
| + | and put in the 37°C waterbath at 200 rpm. | ||
===Transporters=== | ===Transporters=== | ||
| Line 57: | Line 75: | ||
===Vectors=== | ===Vectors=== | ||
| + | *'''Check MymT construct and amplify mymT''' | ||
| + | **Isolate plasmid from o/n culture by Sigma Miniprep kit. | ||
| + | **Check pGB68 (mymT) by PstI restriction --> expected size 960bp+6000bp | ||
| + | **Check pGB68 (mymT) by PCR | ||
==Dry== | ==Dry== | ||
| + | |||
| + | Update is the last week from Annelies: | ||
| + | I look at figure 2 from Dey1995 but we don’t used is because the graph is in time and there are no exact data about the concentration of cells. | ||
| + | I look at figure 1B from Sing2008, about the concentrations As in the cell. | ||
| + | Update literature list from accumulation. | ||
| + | ArsB is regulated by the same promoter from ArsR. They are lie in the same frame in the chromosomal DNA in E. coli K12. | ||
| + | Looking for other plasmids in the E. coli strain what we used. Looking of the Ars operon on the plasmid the same is on the chromosomal DNA in DHB10. I have used BLAST and have positive results. | ||
| + | On the moment I check the density of the E. coli. | ||
| + | |||
| + | Jasper worked on the (equilibrium) equations for ArsB related export taking regulation by ArsR into account. Together with Klaas-Bernd he looked at the impact this had on our ability to fit our model to the data from [[Team:Groningen/Literature#Kostal2004|Kostal2004]]. Using the revised model we did get a meaningful fit (no negative values), however the K5 value was about 5-10 times lower than computed on our [[Team:Groningen/Project/Transport|transport page]]. We now plan to see if we can also apply this new model to the data from [[Team:Groningen/Literature#Singh2008|Singh2008]], which is slightly more difficult (as it requires a time curve), and also to compare Kostal2004 and Singh2008. Jasper also made some modifications based on Wilfred's comments (see [[Team:Groningen/Notebook/10_August_2009|yesterday]]). | ||
{{Team:Groningen/Notebook/Day/Footer}} | {{Team:Groningen/Notebook/Day/Footer}} | ||
Latest revision as of 09:41, 12 August 2009
Wet
GVP Cluster
- → DONE isolate plasmids from overnight precultures
- → DONE plate on cultures for storage in fridge (amp)
- → TODO cut plasmid and GVP containing plasmid for ligation (ligation tomorrow)
- → TODO purify wanted fragments (tomorrow)
- → DONE make o.n. precultures from plate colonies
Plasmid Purification
Plasmid isolation was performed on the cultures of E.coli TOP10 containing plasmids J61002 with high, medium and low constitutive promoters with the "Sygma-Aldrich™ GenElute™ Plasmid Miniprep Kit".
- From each tube 4mL of culture was collected in a 2.0mL cup, and the cells were pelleted by centrifugation for 1 min. at max. speed and the supernatant discarded.
- Plasmids were eluted with 20μL MQ and stored in the fridge
Concentrations
| Plasmid | Conc. ng/μL | 260/280 | 260/230 | -20 box (michael | Restriction Control |
| pJ61002-BBa_J23100 (high) | 174.6 | 1.94 | 2.40 | E-8 | No |
| pJ61002-BBa_J23106 (medium) | 149.2 | 1.95 | 2.41 | E-9 | No |
| pJ61002-BBa_J23109 (low) | 152.8 | 1.94 | 2.38 | E-10 | No |
- → These plasmids come from the glycerol stocks contaning E.coli TOP10 cells with iGEM plasmid J61002 and the high, medium and low constitutive promoters with RFP as reporter gene behind them.
- → The GVP cluster can be ligated behind the promoters and self ligation of RFP will turn unwanted colonies red!
Plates
LB-agar plates with 50 μg/mL ampicillin were freshly prepared, and one drop of each o.n. preculture was spotted and subsequently swiped three times with sterile needle. The plates were stored at 37°C overnight.
Precultures
Over night cultures in 5mL LB-amp50 medium were prepared from the following colonies:
→ E.coli TOP10 pSB1AC3-med. const. promoter - GVP (amp.) (4x) (1 to 4)
→ E.coli TOP10 pSB1AC3-low const. promoter - GVP (amp.) (4x) (5 to 8)
and put in the 37°C waterbath at 200 rpm.
Transporters
Metal Accumulation
Vectors
- Check MymT construct and amplify mymT
- Isolate plasmid from o/n culture by Sigma Miniprep kit.
- Check pGB68 (mymT) by PstI restriction --> expected size 960bp+6000bp
- Check pGB68 (mymT) by PCR
Dry
Update is the last week from Annelies: I look at figure 2 from Dey1995 but we don’t used is because the graph is in time and there are no exact data about the concentration of cells. I look at figure 1B from Sing2008, about the concentrations As in the cell. Update literature list from accumulation. ArsB is regulated by the same promoter from ArsR. They are lie in the same frame in the chromosomal DNA in E. coli K12. Looking for other plasmids in the E. coli strain what we used. Looking of the Ars operon on the plasmid the same is on the chromosomal DNA in DHB10. I have used BLAST and have positive results. On the moment I check the density of the E. coli.
Jasper worked on the (equilibrium) equations for ArsB related export taking regulation by ArsR into account. Together with Klaas-Bernd he looked at the impact this had on our ability to fit our model to the data from Kostal2004. Using the revised model we did get a meaningful fit (no negative values), however the K5 value was about 5-10 times lower than computed on our transport page. We now plan to see if we can also apply this new model to the data from Singh2008, which is slightly more difficult (as it requires a time curve), and also to compare Kostal2004 and Singh2008. Jasper also made some modifications based on Wilfred's comments (see yesterday).
|
|
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
|
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
 "
"