Team:Groningen/Notebook/15 September 2009
From 2009.igem.org
Wet
GVP Cluster
Planning
- → TODO work out the wiki page for GVP
- → DONE make a doodle for presentation planning (1-19 oct.)
- → TODO media attention
- → DONE place an ethics survey link on twitter
- → TODO
Plates
| Name | Plasmid Used | Antibiotics on Plasmid | No. of Colonies | Date |
| GlpF no.1 (low) | pSB1AC3 | Ampicillin | 0 | 14 sept. |
| GlpF no.1 (high) | pSB1AC3 | Ampicillin | 2 | 14 sept. |
| GlpF no.2 (low) | pSB1AC3 | Ampicillin | 1 | 14 sept. |
| GlpF no.2 (high) | pSB1AC3 | Ampicillin | 1 | 14 sept. |
| pLacI (low) | pSB1AC3 | Ampicillin | ~30 | 14 sept. |
| pLacI (high) | pSB1AC3 | Ampicillin | ~80 | 14 sept. |
| pLacI-GVP no.1 (low) | pSB1A2 | Ampicillin | 0 | 14 sept. |
| pLacI-GVP no.1 (high) | pSB1A2 | Ampicillin | 4 | 14 sept. |
| pLacI-GVP no.2 (low) | pSB1A2 | Ampicillin | 0 | 14 sept. |
| pLacI-GVP no.2 (high) | pSB1A2 | Ampicillin | 9 | 14 sept. |
| Positive (J23101) (high) | J61002 | Ampicillin | ~1500 | 14 sept. |
| Negative (MQ) (high) | MQ | Ampicillin | 0 | 14 sept. |
- → The plates showed a low amount of colonies, and also red in the case of the positive plate.
- → The two plates with stripes of pZntR-GVP (pSB2K3) and pCueO-GVP (pSB2K3) culture for glycerol stocks showed growth and some single colonies.
- → All plates were stored in the fridge for further use.
Cultures
The overnight cultures with LB-amp100 medium of colonies E.coli TOP10 with pArsR-GVP (4x), GVP (2x), pNL29 (2x), and HmtA (1-3) all showed expected growth of bacteria.
- Two of the pArsR-GVP tubes were handed over to Steven-Jelle for use of inoculating bigger cultures, the remaining tubes were used for plasmid isolation.
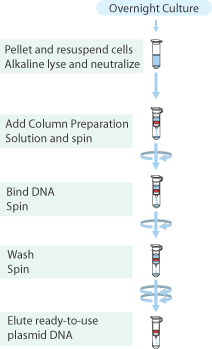
Plasmid isolation
Plasmid isolation was performed on the cultures of E.coli TOP10 containing the above mentioned plasmids with the "Sygma-Aldrich™ GenElute™ Plasmid Miniprep Kit".
- From each tube 5 to 10mL of culture was collected in a 2.0mL cup (tubes from pArsR-GVP, GVP and pNL29 were combined), and the cells were pelleted by centrifugation for 1 min. at max. speed and the supernatant discarded.
- Plasmids were eluted with 30μL MQ and stored in the fridge
Concentrations
| Plasmid | Conc. ng/μL | 260/280 | 260/230 | -20 box (michael | Restriction Control |
| pArsR-GVP (J61035) | 108.9 | 1.83 | 1.91 | x | Yes (EcoRI/PstI) |
| GVP (J61035) | 469.9 | 1.83 | 2.35 | x | x |
| pNL29 | 18.1 | 1.83 | 1.83 | x | x |
| HmtA no.1 | 52.1 | 1.85 | 1.87 | x | Yes (PstI) |
| HmtA no.2 | 49.4 | 1.83 | 1.61 | x | Yes (PstI) |
| HmtA no.3 | 42.6 | 1.91 | 2.06 | x | Yes (PstI) |
Restriction for Assembly, and Control (HmtA)
The plasmids from the o.n. precultures of pArsR-GVP and pSB2K3 (earlier last week) were cut with PstI and EcoRI to cut out the entire part between the pre- and suffix. The plasmids with PstI were cut with PstI for control.
| Plasmid | Amount μL | MQ μL | Fast digest buffer | EcoRI fast digest enzyme | XbaI fast digest enzyme | SpeI fast digest enzyme | PstI fast digest enzyme |
| pArsR-GVP | 13.0 | 3.0 | 3.0 | 1.0 | x | x | 1.0 |
| pSB2K3 | 4.0 | 11.0 | 3.0 | 1.0 | x | x | 1.0 |
| HmtA no.1 | 16.0 | x | 3.0 | x | x | x | 1.0 |
| HmtA no.2 | 16.0 | x | 3.0 | x | x | x | 1.0 |
| HmtA no.3 | 16.0 | x | 3.0 | x | x | x | 1.0 |
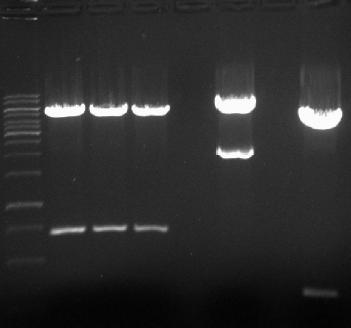
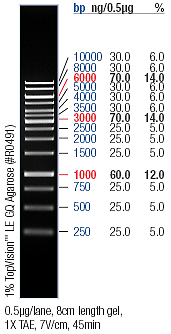
- → From left to right: 1kB ladder, HmtA no.1, HmtA no.2, HmtA no.3, Empty Slot, pArsR-GVP, Empty Slot, pSB2K3
Purification
- → In step 7 the fragments were eluted in 12μL MQ and was stored on ice until use.
Concentrations
| Plasmid | Conc. ng/μL | 260/280 | 260/230 | -20 box (michael | Restriction Control |
| pArsR-GVP (E/P) | 36.1 | 1.91 | 0.17 | x | Yes (EcoRI/PstI) |
| pSB2K3 (E/P) | 58.0 | 1.93 | 0.13 | x | Yes (EcoRI/PstI) |
Ligation
A total amount of vector of 100ng was used in a 1:3 ratio with insert.
- 3 uL Ligase buffer
- 1 uL T4 Ligase
- 9 uL MQ
- 4 uL plasmid pSB2K3 EcoRI/PstI
- 3 uL GVP restricted with EcoRI/PstI
Incubate:
- 25°C 50min.
- kept on ice for 10min.
Tranformation
- add 10uL of the ligation product to 50uL competent E.coli TOP10 cells.
Incubate:
- 30 min @ ice
- 90 sec 42°C
- 2 min @ ice
- add 800uL LB-medium
- incubate for 1 h at 37°C
- plate on LB-kan50 plates
- → Negative control was MQ.
Saline Test
The o.n. plates for Saline test (5 in total) were taken from the stove and 4mL Saline solution was added to dissolve the colonies. The volume of dissolved cells was adjusted to 8mL, and a ten times dilution was made for OD600 measurements.
| Sample | OD600 (dilution) | OD600 |
| Control (J23101) | 0.324 | 3.24 |
| pLacI-GVP | 0.342 | 3.42 |
| pNL29 | 0.316 | 3.16 |
| L-GVP | 0.408 | 4.08 |
| M-GVP | 0.186 | 1.86 |
The tubes were diluted to adjust the OD600 to the lowest sample (1.86) of M-GVP. After adjustment the volumes were reduced to 8mL and an additional 3mL of Saline was added. The OD600 of all tubes was 1.3.
For the test a number of 5 tubes was filled with 10mL Saline and chilled in the fridge for 4 hours. From each solution of cells 1mL was carefully loaded on top of the column. With a bit of luck, a difference will appear between the cells with and without gas vesicles.
Transporters
Metal Accumulation
Vectors
Dry
|
|
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
|
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
 "
"