From 2009.igem.org
Wet
GVP Cluster
Discussion:
All colonies (transformants vector + promoter ligated with gvp) and o/n culture became red, so probably RFP is still in there...
What might be the problem? The vector with promoter self-ligated due to uncomplete digestion (done 15 July 2009) leading to the following fragments:
- vector+RFP linear
- vector
- RFP
When the upper fragment (~2kb) is contaminated with linear vector this would lead high efficiency self-ligation instead of ligating gvp with the emtpy vector.
- →TODO Therefore the restriction/purification/ligation will be redone and selection of non-red colonies will be made!
- →DONE Also plasmid from the o/n cultures will be purified and analysed by restriction analysis.
- For plasmid isolation the Sigma Plasmid isolation kit was used (eluted in 50ul MQ).
- DNA concentrations:
| Sample
| ng/ul
| 260/280
| 260/230
|
| gpv-promoter test (colony 9)
| 82.5
| 2.02
| 2.21
|
| gpv-promoter test (colony 10)
| 71.9
| 1.96
| 2.21
|
- Restriction analysis with SpeI and PstI to find insert length.
- Expected length of with / with RFP insert: 2096bp(promoter + plasmid backbone), 887bp(RFP)
- Expected length of with gvp insert: 2096bp(promoter + plasmid backbone),6096bp(gvp)
- Expected length of with gpv insert: 3513(plasmid backbone), 6096 (GVP cluster)
Restriction mixture for J23100+gvp and GVP
- 10 μL promoter plasmid (61.4 ng/μL, 96.1 ng/μL)
- 6μL MQ
- 2μL Fast digest buffer
- 1μL PstI fast digest enzyme
- 1μL SpeI fast digest enzyme
10ul was loaded on a 1%TBE agarose gel and ran for 30 @ 100V (V const)

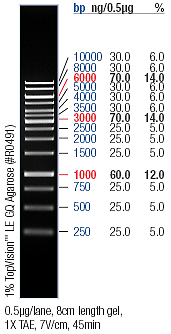
- →From left to right: 1kb marker, gvp-promoter () construct nr 9, nr 10, gvp in , promoter ()
So the assumption that the vector with the promoter self-ligated was right, as the gvp-promoter construct has the same bands as the promoter has and the size of the bands are as expected for a RFP insert.
Transporters
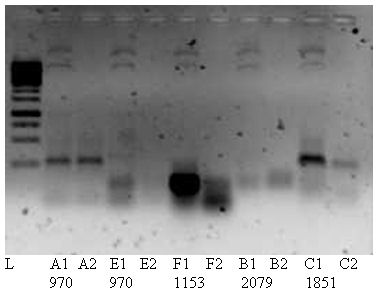
Because of the negative results we will try to get positive results with an other polymerase enzyme; Phusion. And try the cloning again as planned. PCR1 Fw,mut1RC should give product of 1153bp and PCR2 Rev,mut2RC a 261bp size product. Further more we will test a PCR with pBAD 24 Fw,rev to get smaller templates (2222 bp).
PCR1
| Component | amount
|
| 2x Phusion MM
| 12.5 uL
|
| MQ
| 9.5 uL
|
| Fw
| 1 uL
|
| mut1RC
| 1 uL
|
| DNA
| 1 uL
|
|
|
| PCR1 program
| Temperature
| Time
|
| Denaturing
| 98°
| 1.30 min
|
|
| Start Cycles 30X
|
| Denaturing
| 98°
| 10 sec
|
| Annealing
| 55°
| 20 sec
|
| Elongation
| 72°
| 2:10 min
|
|
| End cycles
|
| Final elongation
| 72°
| 10 min
|
| Hold
| 4°
| Forever
|
|
|
|
PCR2
| Component | amount
|
| 2x Phusion MM
| 12.5 uL
|
| MQ
| 9.5 uL
|
| Rev
| 1 uL
|
| mut2RC
| 1 uL
|
| DNA
| 1 uL
|
|
|
| PCR2 program
| Temperature
| Time
|
| Denaturing
| 98°
| 1.30 min
|
|
| Start Cycles 30X
|
| Denaturing
| 98°
| 10 sec
|
| Annealing
| 55°
| 20 sec
|
| Elongation
| 72°
| 20 sec
|
|
| End cycles
|
| Final elongation
| 72°
| 10 min
|
| Hold
| 4°
| Forever
|
|
|
|
pBAD24
| Component | amount
|
| mm NHSO
| 21 uL
|
| 24_F
| 1 uL
|
| 24_R
| 1 uL
|
| DNA
| 1 uL
|
| taq polymerase
| 1 uL
|
|
|
| PCR pBAD-24 program
| Temperature
| Time
|
| Denaturing
| 95°
| 2.00 min
|
|
| Start Cycles 25X
|
| Denaturing
| 95°
| 30 sec
|
| Annealing
| 55°
| 20 sec
|
| Elongation
| 72°
| 2.10 min
|
|
| End cycles
|
| Final elongation
| 72°
| 10 min
|
| Hold
| 4°
| Forever
|
|
|
|
Metal Accumulation
Vectors
Dry
 "
"



