From 2009.igem.org
Wet
GVP Cluster
Discussion:
All colonies (transformants vector + promoter ligated with gvp) and o/n culture became red, so probably RFP is still in there...
What might be the problem? The vector with promoter self-ligated due to uncomplete digestion (done 15 July 2009) leading to the following fragments:
- vector+RFP linear
- vector
- RFP
When the upper fragment (~2kb) is contaminated with linear vector this would lead high efficiency self-ligation instead of ligating gvp with the emtpy vector.
- →TODO Therefore the restriction/purification/ligation will be redone and selection of non-red colonies will be made!
- →DONE Also plasmid from the o/n cultures will be purified and analysed by restriction analysis.
- For plasmid isolation the Sigma Plasmid isolation kit was used (eluted in 50ul MQ).
- DNA concentrations:
| Sample
| ng/ul
| 260/280
| 260/230
|
| gpv-promoter test (colony 9)
| 82.5
| 2.02
| 2.21
|
| gpv-promoter test (colony 10)
| 71.9
| 1.96
| 2.21
|
- Restriction analysis with SpeI and PstI to find insert length.
- Expected length of with / with RFP insert: 2096bp(promoter + plasmid backbone), 887bp(RFP)
- Expected length of with gvp insert: 2096bp(promoter + plasmid backbone),6096bp(gvp)
- Expected length of with gpv insert: 3513(plasmid backbone), 6096 (GVP cluster)
Restriction mixture for J23100+gvp and GVP
- 10 μL promoter plasmid (61.4 ng/μL, 96.1 ng/μL)
- 6μL MQ
- 2μL Fast digest buffer
- 1μL PstI fast digest enzyme
- 1μL SpeI fast digest enzyme
10ul was loaded on a 1%TBE agarose gel and ran for 30 @ 100V (V const)

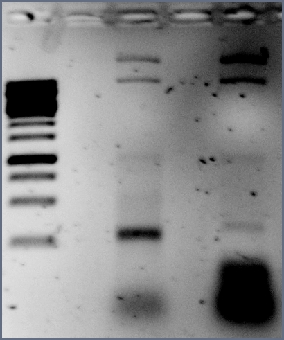
- →From left to right: 1kb marker, gvp-promoter () construct nr 9, nr 10, gvp in , promoter ()
So the assumption that the vector with the promoter self-ligated was right, as the gvp-promoter construct has the same bands as the promoter has and the size of the bands are as expected for a RFP insert.
Transporters
Below yesterdays PCRs, There seems to be a vague band at ~1000 in A1 and E1. therefore loaded the last 20 uL PCR Product on gel as well. showing an expected product op 970bp.
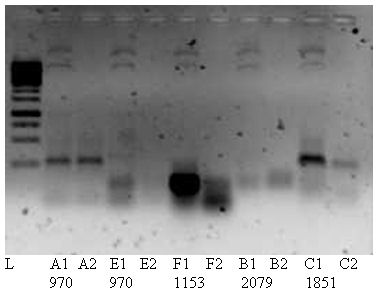
Because of the negative results we will try to get positive results with an other polymerase enzyme; Phusion. And try the cloning again as planned. PCR1 Fw,mut1RC should give product of 1153bp and PCR2 Rev,mut2RC a 261bp size product. Further more we will test a PCR with pBAD 24 Fw,rev to test the template and look at options to get smaller templates (2222 bp).
PCR1
| Component | amount
|
| 2x Phusion MM
| 12.5 uL
|
| MQ
| 9.5 uL
|
| Fw
| 1 uL
|
| mut1RC
| 1 uL
|
| DNA
| 1 uL
|
|
|
| PCR1 program
| Temperature
| Time
|
| Denaturing
| 98°
| 1.30 min
|
|
| Start Cycles 30X
|
| Denaturing
| 98°
| 10 sec
|
| Annealing
| 55°
| 20 sec
|
| Elongation
| 72°
| 30 sec
|
|
| End cycles
|
| Final elongation
| 72°
| 10 min
|
| Hold
| 4°
| Forever
|
|
|
|
PCR2
| Component | amount
|
| 2x Phusion MM
| 12.5 uL
|
| MQ
| 9.5 uL
|
| Rev
| 1 uL
|
| mut2RC
| 1 uL
|
| DNA
| 1 uL
|
|
|
| PCR2 program
| Temperature
| Time
|
| Denaturing
| 98°
| 1.30 min
|
|
| Start Cycles 30X
|
| Denaturing
| 98°
| 10 sec
|
| Annealing
| 55°
| 20 sec
|
| Elongation
| 72°
| 20 sec
|
|
| End cycles
|
| Final elongation
| 72°
| 10 min
|
| Hold
| 4°
| Forever
|
|
|
|
pBAD24
| Component | amount
|
| mm NHSO
| 21 uL
|
| 24_F
| 1 uL
|
| 24_R
| 1 uL
|
| DNA
| 1 uL
|
| taq polymerase
| 1 uL
|
|
|
| PCR pBAD-24 program
| Temperature
| Time
|
| Denaturing
| 95°
| 2.00 min
|
|
| Start Cycles 30X
|
| Denaturing
| 95°
| 30 sec
|
| Annealing
| 55°
| 20 sec
|
| Elongation
| 72°
| 2.10 min
|
|
| End cycles
|
| Final elongation
| 72°
| 10 min
|
| Hold
| 4°
| Forever
|
|
|
|
Metal Accumulation
Vectors
Dry
We mostly tried to find other/better software to use instead of Matlab. In simbiology you are able to export a model using SBML. At the SBML (Systems Biology Markup Language) website if you click on SBML Software Guide and then choose SBML Software Matrix you get a table of all sorts of modelingprograms. It specifies what each program does and in what manner it is compatible with SBML. Specifically we looked at: VCell, Tinkercell and Cytoscape. Although they all claim to be compatible with SBML we had trouble loading our model in VCell and Tinkercell. Cytoscape was able to load our model, but it is in its core stronger in analyzing networks then running simulations.
Furthermore we tried to get SEMPPR to work. We failed under Windows, but succeeded after recompiling it under Linux without -static. We still have to figure out exactly how to interpret the results though (in any case, the production rates only apply to the translation step), to which end I (Jasper) have started a list of concepts on the modelling page.
 "
"




