Team:Groningen/Notebook/24 July 2009
From 2009.igem.org
(→Transporters) |
(→Vectors) |
||
| (7 intermediate revisions not shown) | |||
| Line 6: | Line 6: | ||
===Transporters=== | ===Transporters=== | ||
| - | Below the PCR1.1 and PCR2 of 24/07, | + | Below the PCR1.1, with New designed forward primer with mutation for EcoRI restricion site, and PCR2 of 24/07, PCR 2 shows a fragment of desired size (~261pb) and is excised for DNA extraction for PCR3. Our previously excised PCR1 fragment will be used as template(still containing EcoRI)and see if we can reproduce products PCR1 and make PCR1.1. |
| - | [[Image:F102471_2009-07- | + | [[Image:F102471_2009-07-24_12hr_49minPCR1.1PCR2.jpg]] |
{| | {| | ||
| Line 16: | Line 16: | ||
{| border="1" | {| border="1" | ||
| - | |+ '''PCR1''' | + | |+ '''PCR1.1''' |
! Component !! amount | ! Component !! amount | ||
|- | |- | ||
| Line 39: | Line 39: | ||
<!--Tabel 2 hier--> | <!--Tabel 2 hier--> | ||
{| | {| | ||
| - | ! PCR1 program | + | ! PCR1.1/PCR2 program |
! Temperature | ! Temperature | ||
! Time | ! Time | ||
| Line 84: | Line 84: | ||
|Final elongation | |Final elongation | ||
|72° | |72° | ||
| - | | | + | |5 min |
|- | |- | ||
|Hold | |Hold | ||
| Line 116: | Line 116: | ||
<!--Tabel 3 hier--> | <!--Tabel 3 hier--> | ||
|} | |} | ||
| + | |||
| + | PCR from PCR1 gelproduc failed. Now we try to do a gelextraction first for PCR. | ||
| + | |||
| + | |||
| + | Today we have been scavenging the departments for genomic DNA to clone our genes out of since the Colony PCR's did not work. | ||
===Metal Accumulation=== | ===Metal Accumulation=== | ||
===Vectors=== | ===Vectors=== | ||
| + | |||
| + | *'''Run the PCR products and restriction digest from Paul (GVP) on gel''' | ||
| + | |||
| + | {| | ||
| + | | | ||
| + | <!--Tabel 1 hier--> | ||
| + | |width="10"| | ||
| + | |||
| + | |||
| + | [[Image:F102471 2009-07-24 10hr 32min KACPaulGlpF.JPG]] | ||
| + | |||
| + | <!--tabel 2 hier--> | ||
| + | |width="10"| | ||
| + | | | ||
| + | |||
| + | '''Legenda:''' | ||
| + | {| | ||
| + | |'''Lane''' | ||
| + | |'''Upper''' | ||
| + | |'''Lower''' | ||
| + | |- | ||
| + | |1 | ||
| + | |1kb marker | ||
| + | |1kb marker | ||
| + | |- | ||
| + | |2 | ||
| + | |PCR K-H | ||
| + | |Jolanda nr 1 (GlpF) | ||
| + | |- | ||
| + | |3 | ||
| + | |PCR K-M | ||
| + | |Jolanda nr 2 (GlpF) | ||
| + | |- | ||
| + | |3 | ||
| + | |PCR K-L | ||
| + | | | ||
| + | |- | ||
| + | |4 | ||
| + | |K | ||
| + | |1kb marker | ||
| + | |- | ||
| + | |5 | ||
| + | |AC-H | ||
| + | |L1 | ||
| + | |- | ||
| + | |6 | ||
| + | |AC-M | ||
| + | |L2 | ||
| + | |- | ||
| + | |7 | ||
| + | |AC-L | ||
| + | | | ||
| + | |- | ||
| + | |8 | ||
| + | |AC | ||
| + | | | ||
| + | |- | ||
| + | |9 | ||
| + | | Paul 100R | ||
| + | | | ||
| + | |- | ||
| + | |10 | ||
| + | |1kb marker | ||
| + | | | ||
| + | |} | ||
| + | |||
| + | |||
| + | |} | ||
| + | |||
| + | The gel shows that there is no product for K-H, K-M and AC, there is no difference seen for K-promoters / AC-promoters and the empty vectors (like K). The size of the Lac promoters seems to be okay. For all product a restriction digest should be done. | ||
| + | |||
| + | * '''Glycerol stocks:''' | ||
| + | Made for: | ||
| + | Ecoli + pSB1AC3- | ||
| + | **J23100 (High) | ||
| + | **J23106 (Med) | ||
| + | **J23109 (Low) | ||
| + | Ecoli + pSBK3K- | ||
| + | **J23100 (High) | ||
| + | **J23106 (Med) | ||
| + | **J23109 (Low) | ||
| + | Ecoli + pSB1A2- | ||
| + | **R0010 (pLac) nr 1 | ||
| + | **R0010 (pLac) nr 2 | ||
| + | They were put in [https://2009.igem.org/Team:Groningen/Parts -80 freezer] | ||
| + | |||
| + | *'''Transformation of E. coli Top10 with two pBAD inducible promoters''' | ||
| + | Use normal protocol. | ||
| + | The following plasmids were transformed: | ||
| + | **pSB2K3-I0500 (pBAD1) --> with extra IPTG added to the LBA | ||
| + | **pSB1A2-K113009 (pBAD2) --> from plate 3, location 1M of the registry | ||
| + | **Negative control, MQ. | ||
| + | Plates were put o/n at 37dg. | ||
| + | |||
| + | *'''O/n culture of pSB3K3 vectors''' | ||
| + | Inoculate 2x ~10ml LB-Kan with: | ||
| + | **pSB3K3-pHigh, pMed, pLow | ||
| + | **pSB3K3 | ||
| + | Put o/n @ 37dg. | ||
==Dry== | ==Dry== | ||
| + | The first part of the day was spent processing the data of yesterday's [[http://www.jbc.org/cgi/content/abstract/M400037200v1 paper]]. However when we calculated the drop in concentration of As(III) outside the membrane vesicle we found that it did not drop significantly enough to make a model of the efflux rate of As(III) depending on concentration of Arsenic in the cell. However we were able to find a paper which a more likely candidate. [[http://www.jbc.org/cgi/content/full/272/1/326?view=long&pmid=8995265 Alternate energy coupling of ArsB, the membrane subunit of the Ars anion-translocating ATPase.]] | ||
| + | |||
| + | Jasper continued working on the RPU computations, the results of which are shown at [[Team:Groningen/Promoters]]. It looks like some weird things happened to the cultures. For example, two cultures which should be identical followed roughly the same growth curve upto a point and then one of them started behaving more erratically. | ||
{{Team:Groningen/Notebook/Day/Footer}} | {{Team:Groningen/Notebook/Day/Footer}} | ||
Latest revision as of 12:04, 25 July 2009
Wet
GVP Cluster
Transporters
Below the PCR1.1, with New designed forward primer with mutation for EcoRI restricion site, and PCR2 of 24/07, PCR 2 shows a fragment of desired size (~261pb) and is excised for DNA extraction for PCR3. Our previously excised PCR1 fragment will be used as template(still containing EcoRI)and see if we can reproduce products PCR1 and make PCR1.1.
|
|
|
PCR from PCR1 gelproduc failed. Now we try to do a gelextraction first for PCR.
Today we have been scavenging the departments for genomic DNA to clone our genes out of since the Colony PCR's did not work.
Metal Accumulation
Vectors
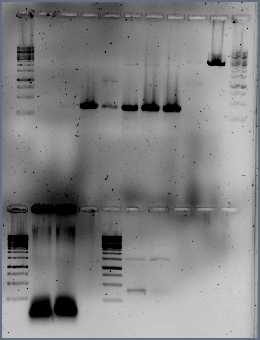
- Run the PCR products and restriction digest from Paul (GVP) on gel
|
Legenda:
|
The gel shows that there is no product for K-H, K-M and AC, there is no difference seen for K-promoters / AC-promoters and the empty vectors (like K). The size of the Lac promoters seems to be okay. For all product a restriction digest should be done.
- Glycerol stocks:
Made for: Ecoli + pSB1AC3-
- J23100 (High)
- J23106 (Med)
- J23109 (Low)
Ecoli + pSBK3K-
- J23100 (High)
- J23106 (Med)
- J23109 (Low)
Ecoli + pSB1A2-
- R0010 (pLac) nr 1
- R0010 (pLac) nr 2
They were put in -80 freezer
- Transformation of E. coli Top10 with two pBAD inducible promoters
Use normal protocol. The following plasmids were transformed:
- pSB2K3-I0500 (pBAD1) --> with extra IPTG added to the LBA
- pSB1A2-K113009 (pBAD2) --> from plate 3, location 1M of the registry
- Negative control, MQ.
Plates were put o/n at 37dg.
- O/n culture of pSB3K3 vectors
Inoculate 2x ~10ml LB-Kan with:
- pSB3K3-pHigh, pMed, pLow
- pSB3K3
Put o/n @ 37dg.
Dry
The first part of the day was spent processing the data of yesterday's [paper]. However when we calculated the drop in concentration of As(III) outside the membrane vesicle we found that it did not drop significantly enough to make a model of the efflux rate of As(III) depending on concentration of Arsenic in the cell. However we were able to find a paper which a more likely candidate. [Alternate energy coupling of ArsB, the membrane subunit of the Ars anion-translocating ATPase.]
Jasper continued working on the RPU computations, the results of which are shown at Team:Groningen/Promoters. It looks like some weird things happened to the cultures. For example, two cultures which should be identical followed roughly the same growth curve upto a point and then one of them started behaving more erratically.
|
|
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
|
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
 "
"