Team:Groningen/Notebook/3 September 2009
From 2009.igem.org
m (→GVP Cluster) |
m (→GVP Cluster) |
||
| Line 265: | Line 265: | ||
Restriction was kept at 37C for 40 min. and put on ice until used for gel purification. | Restriction was kept at 37C for 40 min. and put on ice until used for gel purification. | ||
| + | |||
| + | |||
| + | [[Image:Zymoclean Gel DNA Recovery Kit (D4001) 2.jpg|thumb|300px| www.zymoresearch.com]] | ||
'''Purification''' | '''Purification''' | ||
| Line 273: | Line 276: | ||
| - | A " | + | A "Zymoclean(TM) Gel DNA Recovery Kit" standard protocol was used. |
| - | + | * In step 7 an amount of 10μL MQ was added to elute the DNA fragments. | |
Revision as of 09:11, 4 September 2009
Wet
LB-medium is badly needed!!!!!!!!!!!!!!!!!!!!!!!!!!!!!!!!!!!!!
GVP Cluster
EM pictures
- → TODO Today we will try to get some Electon microscopy pictures of the gas vesicle producing bacteria from the ON cultures.
Orinigal pNL29 iptg-GVP and the biobricks ars-GVP and
- → TODO Choose colonies from plates for growth of E.coli TOP10 with GVP behind LAC/pBAD (+RBS) promoter in pSB1AC3
- → TODO Make glycerol stocks of pBad/araC, pBad/araC+RBS+GVP, and pLacI+RBS+GVP (both in pSB1A2 and pSB1AC3 plasmid)
- → TODO Test control of bouyancy in Saline solution (grow plates with GVP constructs)
- → TODO Order synthetic DNA for GVP
- → TODO Order primer for PstI site removal
- → TODO Test promoter strenght compared to BBa_J23101 promoter (Sven)
- → TODO Enter sequences of constructs to Sandbox
Colonies on Plates
| Name | Plasmid Used | Antibiotics on Plasmid | No. of Colonies | Date |
| pLacI-GVP | http://partsregistry.org/Part:pSB1A2 pSB1A2] | Ampicillin | 26 | 3/9 |
| pLacI-GVP (concentrated) | http://partsregistry.org/Part:pSB1A2 pSB1A2] | Ampicillin | ~100 | 3/9 |
| pBad/araC-GVP | http://partsregistry.org/Part:pSB1A2 pSB1A2] | Ampicillin | 1 | 3/9 |
| pBad/araC-GVP (concentrated) | http://partsregistry.org/Part:pSB1A2 pSB1A2] | Ampicillin | ~40 | 3/9 |
| Negative Control | MQ | None | 0 | 3/9 |
| Positive Control | J61002-J23101 | Ampicillin | ~2000 | 3/9 |
- → The plates showed normal colony growth.
O.n. precultures
- → All twelve o.n. precultures showed bacterial growth, and could be used to isolate plasmid. The first four are registry vector pSB2K3 with RFP as reporter (plate 1, 7C) which were transformed on tuesday. The isolated plasmid is used to house all GVP constructs to combine with the vector pSB1AC3 with transporter and accumulation genes.
- → The next eight tubes contain our own designed biobricks and isolated plasmid will be sent to the registry on friday!
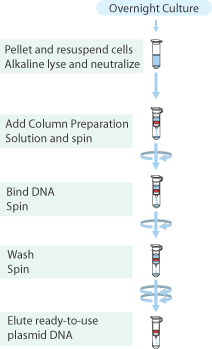
Plasmid Purification
Plasmid isolation was performed on the cultures of E.coli TOP10 containing the above mentioned plasmids with the "Sygma-Aldrich™ GenElute™ Plasmid Miniprep Kit".
- From each tube 4mL of culture was collected in a 2.0mL cup, and the cells were pelleted by centrifugation for 1 min. at max. speed and the supernatant discarded.
- Plasmids were eluted with 30μL MQ and stored in the fridge
Concentrations
| Plasmid | Conc. ng/μL | 260/280 | 260/230 | -20 box (michael | Restriction Control |
| pSB2K3 (7C) no.1 | 178.1 | 1.87 | 2.32 | D-1 | Yes (EcoRI/PstI) |
| pSB2K3 (7C) no.2 | 206.8 | 1.89 | 2.34 | D-2 | Yes (EcoRI/PstI) |
| pSB2K3 (7C) no.3 | 147.1 | 1.87 | 2.31 | D-3 | Yes (EcoRI/PstI) |
| pSB2K3 (7C) no.4 | 186.2 | 1.85 | 2.15 | D-4 | Yes (EcoRI/PstI) |
| BBa_K190015 | 155.3 | 1.88 | 2.14 | B-1 | Yes (EcoRI/PstI) |
| BBa_K190016 | 69.4 | 1.85 | 2.28 | B-2 | Yes (EcoRI/PstI) |
| BBa_K190017 | 85.8 | 1.90 | 2.22 | B-3 | Yes (EcoRI/PstI) |
| BBa_K190022 | 94.9 | 1.82 | 1.70 | B-4 | Yes (EcoRI/PstI) |
| BBa_K190023 | 90.4 | 1.87 | 2.03 | B-5 | Yes (EcoRI/PstI) |
| BBa_K190024 | 82.2 | 1.84 | 2.06 | B-6 | Yes (EcoRI/PstI) |
| BBa_K190025 | 297.1 | 1.85 | 2.37 | B-7 | Yes (EcoRI/PstI) |
| BBa_K190026 | 13.2 | 1.91 | 2.20 | B-8 | Yes (EcoRI/PstI) |
Restriction for Assembly
The vector BBa_J61035 containing the pArsR, pZntR, and pCueO with GVP composite parts were cut with PstI and EcoRI to create correct ends for insert into pSB2K3, which was also cut with EcoRI and PstI (4x).
| Plasmid | Amount μL | MQ μL | Fast digest buffer | EcoRI fast digest enzyme | XbaI fast digest enzyme | SpeI fast digest enzyme | PstI fast digest enzyme |
| pSB2K3 (7C) no.1 | 6.0 | 10.0 | 3.0 | 1.0 | x | x | 1.0 |
| pSB2K3 (7C) no.2 | 6.0 | 10.0 | 3.0 | 1.0 | x | x | 1.0 |
| pSB2K3 (7C) no.3 | 6.0 | 10.0 | 3.0 | 1.0 | x | x | 1.0 |
| pSB2K3 (7C) no.4 | 6.0 | 10.0 | 3.0 | 1.0 | x | x | 1.0 |
| J61035-pArsR-GVP | 16.0 | x | 3.0 | 1.0 | x | x | 1.0 |
| J61035-pZntR-GVP | 10.0 | 6.0 | 3.0 | 1.0 | x | x | 1.0 |
| J61035-pCueO-GVP | 10.0 | 6.0 | 3.0 | 1.0 | x | x | 1.0 |
Restriction was kept at 37C for 40 min. and put on ice until used for gel purification.
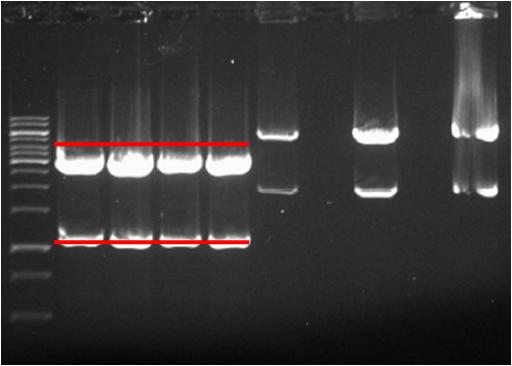
Purification
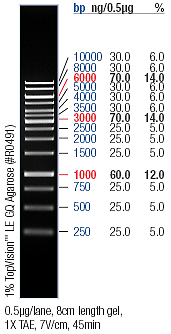
- → From left to right: 1kB ladder, pSB2K3 (no.1, no.2, no.3, and no.4), J61035-pArsR-GVP , Empty Slot, J61035-pZntR-GVP, Empty Slot, J61035-pCueO-GVP
A "Zymoclean(TM) Gel DNA Recovery Kit" standard protocol was used.
- In step 7 an amount of 10μL MQ was added to elute the DNA fragments.
Concentrations
| Plasmid | Conc. ng/μL | 260/280 | 260/230 | -20 box (michael | Restriction Control |
| pSB2K3 (7C) no.1 (EcoRI,PstI) | 37.7 | 1.84 | 0.52 | fridge | See Gel |
| pSB2K3 (7C) no.2 (EcoRI,PstI) | 42.9 | 1.87 | 1.90 | fridge | See Gel |
| pSB2K3 (7C) no.3 (EcoRI,PstI) | 32.9 | 1.84 | 1.52 | fridge | See Gel |
| pSB2K3 (7C) no.4 (EcoRI,PstI) | 51.2 | 2.07 | 0.07 | fridge | See Gel |
| pArsR-GVP (EcoRI,PstI) | 8.9 | 2.06 | 0.03 | fridge | See Gel |
| pZntR-GVP (EcoRI,PstI) | 22.6 | 1.71 | 1.01 | fridge | See Gel |
| pCueO-GVP (EcoRI,PstI) | 17.3 | 1.83 | 0.55 | fridge | See Gel |
Transporters
Metal Accumulation
- Send MBP-ArsR for sequencing again
- Send pLow-fMT for sequencing
- Send pLac-fMT for sequencing
Vectors
Dry
|
|
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
|
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
 "
"